Arylformamidase Function Slide 3: Difference between revisions
From MDWiki
Jump to navigationJump to search
Thomasparker (talk | contribs) No edit summary |
Thomasparker (talk | contribs) |
||
| (6 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
== Evidence from Similar | == Evidence from Similar Sequences == | ||
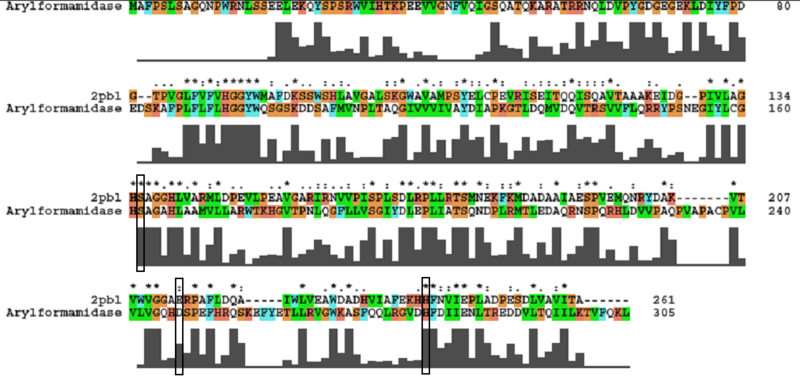
- Catalytic triad identified in paper - Ser162, Asp247, and His279. | |||
- To assess functional similarity, conservation of the catalytic triad was analysed. | |||
[[Image:arylformamidase_alignment.png|centre|framed|'''ClustalW alignment showing conservation of the catalytic triad between Arylformamidase and 2pbl.''']] | |||
- Aspartic acid --> Glutamic acid - Semi-conservative: both polar, acidic. | |||
[[Arylformamidase Function Slide 2| ...Previous slide ]]|[[Arylformamidase| Return to the main page ]]|[[Arylformamidase Function Slide 4| Next slide... ]] | [[Arylformamidase Function Slide 2| ...Previous slide ]]|[[Arylformamidase| Return to the main page ]]|[[Arylformamidase Function Slide 4| Next slide... ]] | ||
Latest revision as of 23:34, 9 June 2008
Evidence from Similar Sequences
- Catalytic triad identified in paper - Ser162, Asp247, and His279.
- To assess functional similarity, conservation of the catalytic triad was analysed.
- Aspartic acid --> Glutamic acid - Semi-conservative: both polar, acidic.