|
|
| (122 intermediate revisions by the same user not shown) |
| Line 1: |
Line 1: |
| 1. ''''''A chain''' of arylformamidase''' | | =='''Methods'''== |
| | |
| | The structure of arylfromamidase was obtained from RCSB Protein Data Bank (PDB ID: 2PBL). |
| | http://www.rcsb.org/pdb/home/home.do |
| | |
| | The predicted interaction arylformamidase with other proteins was determined using the STRING database (STRING: Search Tool for the Retrieval of Interacting Genes/Proteins). http://string.embl.de// |
| | |
| | The DALI database was used for the structural comparison of arylformamidase with other proteins. http://ekhidna.biocenter.helsinki.fi/dali_server/ |
| | |
| | PDBsum database was used to determine the secondary structure of arylformamidase. |
| | http://www.ebi.ac.uk/pdbsum/ |
| | |
| | |
| | |
| | == '''Results''' == |
| | |
| | |
| | == Structure of Arylformamidase == |
| | |
| | Structure was determined using X-ray diffraction by the Joint Center for Structural Genomics (JCSG). The organism is Silicibacter SP. TM1040 and the protein expression system is Escherichia Coli (vector type: plasmid). The resolution is 1.79 A with R-value of 0.224 and R-free value of 0.270. The closer the R values are to each other, the better the quality of the structure. |
| | |
| | |
| | |
| | |
| | '''Figure: Arylformamidase (All Chains)''' |
| | |
| | |
| | [[Image:Arylformamidase the whole protein.PNG]] |
| | |
| | |
| | ''The image above shows the chains A (upper right), B (upper left), C (lower right) & D (lower left) interacting. The molecules in the middle of chains A & B and chains C & D is phosphate ion (PO4). The green molecule between chain B & D is a magnesium ion (Mg). These ions aren't biologically significant and could only be an artefact. When crystallizing proteins they often form complexes (dimer, tetramers etc) but that doesn't mean that the functional structure is the same. They could be functional monomers. The chains in the protein of interest exist as individual functional units because in the PDB file it assumes the functional biological molecule as a monomer.'' |
| | |
| | ''Image from PDB ProteinWorkshop 1.5'' |
| | |
| | |
| | |
| | '''Figure:''Chain A'' of arylformamidase''' |
| | |
| | |
| | [[Image:ChainA1.PNG]] |
|
| |
|
| [[Image:chain_A.jpg]]
| |
|
| |
|
| ''The red molecule in the middle is an unknown ligand containing a ring composed of 9 oxygen molecules. The green sphere is a chloride ion.'' | | ''The red molecule in the middle is an unknown ligand containing a ring composed of 9 oxygen molecules. The green sphere is a chloride ion.'' |
|
| |
|
| '''2. Arylformamidase''' | | ''Image from PDB ProteinWorkshop 1.5'' |
| | |
| | '''''The protein backbone is coloured by conformation type:''''' |
| | |
| | '''''Turn - blue''''' |
| | |
| | '''''Coil- pink''''' |
| | |
| | '''''Helix- green''''' |
| | |
| | '''''strand- purple''''' |
| | |
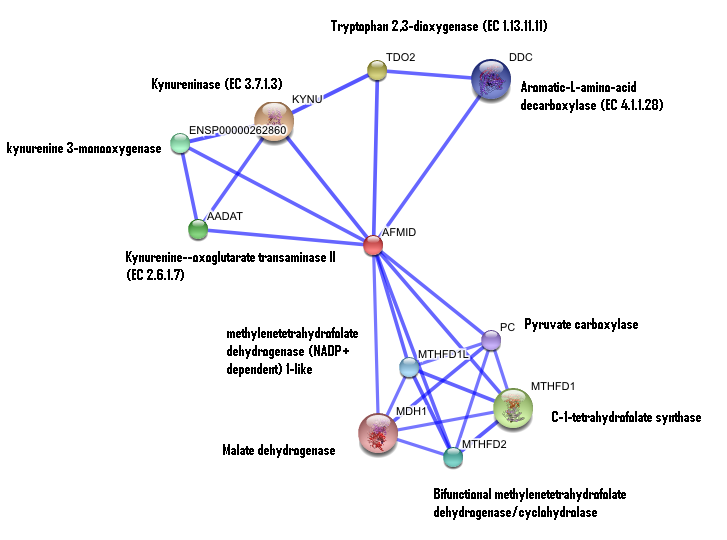
| | == '''Interaction of human arylformamidase (AFMID) with other proteins''' == |
| | |
| | |
| | |
| | [[Image:Confidence_interaction_with_names.png]] |
| | |
| | |
| | ''The interaction between the proteins have been determined from curated STRING database (significant score). However there is no significant evidence for:'' |
| | |
| | ''1- Neighborhood in the genome'' |
| | |
| | ''2- Gene fusions '' |
| | |
| | ''3- Cooccurence across genomes '' |
| | |
| | ''4- Co-Expression '' |
| | |
| | ''5- Experimental/Biochemical data'' |
| | |
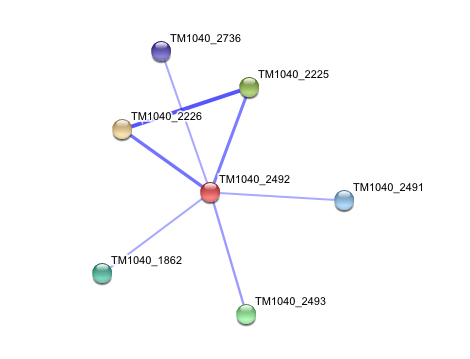
| | == '''Interaction of Silicibacter Sp. arylformamidase (AFMID) with other proteins''' == |
| | |
| | |
| | |
| | [[Image:Examplec.jpg]] |
| | |
| | |
| | TM1040_2226 ''Tryptophan 2,3-dioxygenase (279 aa)'' |
| | |
| | TM1040_2225 ''Kynureninase (396 aa)'' |
| | |
| | TM1040_2493 ''Succinic semialdehyde dehydrogenase (490 aa)'' |
| | |
| | TM1040_1862 ''Hypothetical protein (212 aa)'' |
| | |
| | TM1040_2491 ''Creatinase (402 aa'') |
| | |
| | TM1040_2736 ''Transketolase, putative (794 aa)'' |
| | |
| | |
| | ''There is no significant evidence for these interactions (score= ~0.5)'' |
| | |
| | == '''DALI OUTPUT''' == |
| | |
| | |
| | |
| | The DALI tool produces proteins that are structurally similar to the protein of interest. |
| | |
| | The search result showed similarities to mostly carboxylesterases/hydrolases. Hence there is strong evidence that our protein might also be a carboxylesterase. |
| | |
| | [[Image:DALI RESULT.txt]] |
| | |
| | |
| | |
| | '''Figure: Metagenomic Archea Carboxylesterase (Chain A ONLY)''' |
| | |
| | [[Image:ChainA 2c7b.PNG]] |
| | |
| | [[Image:Carboxylase.txt ]] |
| | |
| | '''PDB''' [http://www.rcsb.org/pdb/explore/explore.do?structureId=2C7B link title] |
| | |
| | ''Note: Chain B not shown'' |
| | |
| | ''From PDB ProteinWorkshop 1.5'' |
| | |
| | |
| | '''Figure: Archaeoglobus fulgidus Carboxylesterase (Chain A ONLY)''' |
| | |
| | [[Image:ChainA 1jji.PNG]] |
| | |
| | [[Image:Carboxylesterase (archaeon).txt]] |
| | |
| | '''PDB''' [http://www.rcsb.org/pdb/explore/explore.do?structureId=1JJI link title] |
| | |
| | ''Note: Chains B, C & D not shown'' |
| | |
| | ''From PDB ProteinWorkshop 1.5'' |
| | |
| | |
| | ''Both of the above Archaeal carboxylesterases' chains exist as monomers (from literature). Hence it is expected that our protein exists as a monomer but during crystalization it interacts with its chains.'' |
| | |
| | == '''Secondary structure analysis''' == |
| | |
| | |
| | |
| | '''PDBSum output for arylformamidase''' |
| | |
| | [[Image:PDBSum pblA.PNG]] |
|
| |
|
| [[Image:Whole protein.png]] | | PDBSUM [http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/GetPage.pl?pdbcode=2pbl&template=main.html] |
|
| |
|
| ''The image above shows the chains A (upper right), B (upper left), C (lower right) & D (lower left) interacting. The molecules in the middle of chains A & B and chains C & D is phosphate ion (PO4). The green molecule between chain B & D is a magnesium ion (Mg).''
| |
|
| |
|
| | '''Figure: Archeon Carboxylesterase secondary structure''' |
|
| |
|
| | [[Image:Pdbsums archeal.PNG]] |
|
| |
|
|
| |
|
| '''Thioesterase: Splits ester into acid and alcohol, in the presence of water, specifically at a thiol group''' | | ''The secondary structure shows the conservation of the order of different conformation types between the protein of interest and the archaeal carboxylesterases.'' |
|
| |
|
| | ''Images from PDBsum'' |
|
| |
|
| | == '''The conservation of the ser/his/asp catalytic triad''' == |
|
| |
|
| | [[Image:Catalytic triad conversation.PNG]] |
|
| |
|
| | ''Yellow indicates conservation'' |
|
| |
|
| | ''Blue indicates semi-conservation'' |
|
| |
|
|
| |
|
| '''Interaction of human arylformamidase with other proteins'''
| |
|
| |
|
| [[Image:Confidence_interaction_with_names.png]]
| |
|
| |
|
| | '''Figure: The catalytic triad''' |
| | |
| | |
| | [[Image:Untitled2.PNG]] |
| | |
| | |
| | ''The above image shows the conserved residues of the catalytic triad in arylformamidase, with the unknown ligand (Blue) protruding from a surface groove. The residues are serine 136, Histidine 241 and Glutamate 214. '''Note:''' The actual residue numbers are n+1'' |
| | |
| | ''Image generated using Pymol'' |
| | |
| | |
| | '''Figure: The conserved residues of arylformamidase''' |
| | |
| | |
| | [[Image:Cat triad red.PNG]] |
| | |
| | |
| | ''The blue region shows the residues conserved among species. It is mostly around the unknown ligand. The conserved residues were obtained from observing the clustal alignment.'' |
| | |
| | ''Image generated using Pymol'' |
| | |
| | |
| | '''Figure: The catalytic triad''' |
| | |
| | |
| | [[Image:CATALYTIC TRIAD 1.PNG]] |
| | |
| | ''The above image shows the distance between the catalytic triad conserved residues and how each amino acid is linked to a turn region. This catalytic triad is also conserved in the Metagenomic Archea Carboxylesterase (PDB ID 2C7B) and the Archaeoglobus fulgidus Carboxylesterase (PDB ID 1JJI)'' |
| | |
| | '' From PDB ProteinWorkshop 1.5'' |
| | |
| | |
| | |
| | '''Figure: The conserved catalytic triad in Archaeoglobus fulgidus Carboxylesterase (PDB ID 1JJI)''' |
|
| |
|
| [[Arylformamidase-associated FASTA sequences | FASTA sequences]]
| |
|
| |
|
|
| |
|
| | [[Image:Cat triad 1jji.PNG]] |
|
| |
|
| DALI:
| |
|
| |
|
| | ''The catalytic triad in Archaeoglobus fulgidus Carboxylesterase is very close to the ligand which is also present in aryformamidase.'' |
|
| |
|
|
| |
|
| Query: mol1A MOLECULE: PUTATIVE ESTERASE/LIPASE/THIOESTERASE;
| |
| Matches are sorted by Z-score. Similarities with a Z-score lower than 2 are spurious.
| |
| Summary
| |
|
| |
|
| No: Chain Z rmsd lali nres %id Description
| | '''Sequence alignment:''' |
| 1: 2pbl-A 52.4 0.0 262 262 100 MOLECULE: PUTATIVE ESTERASE/LIPASE/THIOESTERASE;
| |
| 2: 2pbl-B 50.7 0.1 262 262 100 MOLECULE: PUTATIVE ESTERASE/LIPASE/THIOESTERASE;
| |
| 3: 2pbl-D 50.0 0.3 262 262 100 MOLECULE: PUTATIVE ESTERASE/LIPASE/THIOESTERASE;
| |
| 4: 2pbl-C 50.0 0.3 262 262 100 MOLECULE: PUTATIVE ESTERASE/LIPASE/THIOESTERASE;
| |
| 5: 2c7b-A 23.4 3.0 231 294 16 MOLECULE: CARBOXYLESTERASE;
| |
| 6: 1jji-B 23.1 3.1 233 311 15 MOLECULE: CARBOXYLESTERASE;
| |
| 7: 1jji-D 23.0 3.1 233 311 15 MOLECULE: CARBOXYLESTERASE;
| |
| 8: 1jji-C 23.0 3.1 233 311 15 MOLECULE: CARBOXYLESTERASE;
| |
| 9: 1jji-A 23.0 3.1 233 311 15 MOLECULE: CARBOXYLESTERASE;
| |
| 10: 1lzl-A 22.0 2.9 233 317 18 MOLECULE: HEROIN ESTERASE;
| |
| 11: 1u4n-A 21.8 2.7 215 308 18 MOLECULE: CARBOXYLESTERASE EST2;
| |
| 12: 1qz3-A 21.8 3.0 227 309 17 MOLECULE: CARBOXYLESTERASE EST2;
| |
| 13: 1lzk-A 21.8 2.8 232 317 19 MOLECULE: HEROIN ESTERASE;
| |
| 14: 1evq-A 21.4 2.9 225 308 18 MOLECULE: SERINE HYDROLASE;
| |
| 15: 1jkm-A 20.7 3.0 229 358 13 MOLECULE: BREFELDIN A ESTERASE;
| |
| 16: 1jkm-B 20.6 3.0 226 361 13 MOLECULE: BREFELDIN A ESTERASE;
| |
| 17: 2hu8-A 19.5 2.9 218 575 16 MOLECULE: ACYLAMINO-ACID-RELEASING ENZYME;
| |
| 18: 2qru-A 19.4 2.8 213 272 15 MOLECULE: UNCHARACTERIZED PROTEIN;
| |
| 19: 2o7r-A 19.3 3.2 224 307 18 MOLECULE: CXE CARBOXYLESTERASE;
| |
| 20: 1ve6-A 19.2 3.0 218 574 16 MOLECULE: ACYLAMINO-ACID-RELEASING ENZYME;
| |
| 21: 2o7v-A 19.1 3.2 222 307 18 MOLECULE: CXE CARBOXYLESTERASE;
| |
| 22: 1vkh-A 18.9 2.6 207 261 16 MOLECULE: PUTATIVE SERINE HYDROLASE;
| |
| 23: 3bjr-A 18.8 2.7 206 244 15 MOLECULE: PUTATIVE CARBOXYLESTERASE;
| |
| 24: 2hu5-B 18.8 2.9 217 575 16 MOLECULE: ACYLAMINO-ACID-RELEASING ENZYME;
| |
| 25: 1ve6-B 18.8 2.9 217 574 17 MOLECULE: ACYLAMINO-ACID-RELEASING ENZYME;
| |
| 26: 2oae-A 18.3 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 27: 2ajb-D 18.3 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 28: 1ufo-D 18.3 2.7 197 238 17 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 29: 2o2g-A 18.2 2.5 187 216 17 MOLECULE: DIENELACTONE HYDROLASE;
| |
| 30: 2hrq-A 18.2 2.9 213 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 31: 2ajc-C 18.2 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 32: 1ufo-B 18.2 2.6 196 238 17 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 33: 1ufo-A 18.2 2.6 196 238 16 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 34: 1nu6-B 18.2 2.9 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 35: 1mx5-F 18.2 2.8 212 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 36: 2j4c-A 18.1 2.7 211 524 17 MOLECULE: CHOLINESTERASE;
| |
| 37: 2h7c-F 18.1 2.9 213 531 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 38: 2bgn-A 18.1 2.9 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 39: 1ufo-F 18.1 2.6 196 238 17 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 40: 1ufo-E 18.1 2.6 196 238 17 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 41: 1ufo-C 18.1 2.6 196 238 17 MOLECULE: HYPOTHETICAL PROTEIN TT1662;
| |
| 42: 1n1m-A 18.1 2.9 212 726 14 MOLECULE: DIPEPTIDYL PEPTIDASE IV SOLUBLE FORM;
| |
| 43: 1mx1-A 18.1 2.7 211 532 14 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 44: 2hrq-E 18.0 2.7 211 531 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 45: 2hrq-D 18.0 2.8 212 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 46: 2h7c-A 18.0 3.0 211 532 12 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 47: 2bua-A 18.0 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 48: 1n1m-B 18.0 2.8 210 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV SOLUBLE FORM;
| |
| 49: 1mx5-B 18.0 2.9 213 531 14 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 50: 1f6w-A 18.0 2.8 215 533 13 MOLECULE: BILE SALT ACTIVATED LIPASE;
| |
| 51: 2jf0-B 17.9 3.3 219 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 52: 2hrr-A 17.9 2.8 211 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 53: 2ecf-A 17.9 2.8 211 700 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 54: 2ack 17.9 2.7 212 527 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 55: 1z68-A 17.9 2.9 210 719 11 MOLECULE: FIBROBLAST ACTIVATION PROTEIN, ALPHA SUBUNIT;
| |
| 56: 1xlu-A 17.9 2.7 212 524 17 MOLECULE: BUTYRYLCHOLINESTERASE;
| |
| 57: 1mx5-D 17.9 2.8 210 532 12 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 58: 1mx5-C 17.9 2.9 213 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 59: 1mx5-A 17.9 2.8 210 532 12 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 60: 1aql-A 17.9 2.8 212 532 13 MOLECULE: BILE-SALT ACTIVATED LIPASE;
| |
| 61: 3bjm-A 17.8 3.0 213 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 62: 2h7c-D 17.8 2.9 211 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 63: 2ajc-A 17.8 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 64: 2ace 17.8 2.7 212 527 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 65: 1p0m-A 17.8 2.7 212 523 16 MOLECULE: CHOLINESTERASE;
| |
| 66: 1mx5-E 17.8 2.7 211 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 67: 1dx6-A 17.8 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 68: 3biw-D 17.7 2.7 214 533 14 MOLECULE: NEUROLIGIN-1;
| |
| 69: 3biw-C 17.7 2.7 214 533 14 MOLECULE: NEUROLIGIN-1;
| |
| 70: 3biw-B 17.7 2.7 214 533 14 MOLECULE: NEUROLIGIN-1;
| |
| 71: 3biw-A 17.7 2.7 214 533 14 MOLECULE: NEUROLIGIN-1;
| |
| 72: 2pm8-A 17.7 2.7 212 530 17 MOLECULE: CHOLINESTERASE;
| |
| 73: 2h7c-E 17.7 2.8 212 531 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 74: 2h7c-C 17.7 2.8 212 531 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 75: 2ckm-A 17.7 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 76: 2buc-B 17.7 2.7 209 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 77: 2ajd-C 17.7 2.8 210 729 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 78: 2ajd-A 17.7 2.8 211 729 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 79: 2aj8-B 17.7 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 80: 2aj8-A 17.7 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 81: 1xlw-A 17.7 2.7 210 524 16 MOLECULE: BUTYRYLCHOLINESTERASE;
| |
| 82: 1xlv-A 17.7 2.7 213 524 16 MOLECULE: BUTYRYLCHOLINESTERASE;
| |
| 83: 1pfq-A 17.7 3.0 213 728 14 MOLECULE: DIPEPTIDYL PEPTIDASE IV SOLUBLE FORM;
| |
| 84: 1p0i-A 17.7 2.7 213 523 15 MOLECULE: CHOLINESTERASE;
| |
| 85: 1odc-A 17.7 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 86: 1nu6-A 17.7 3.0 213 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 87: 1mx1-E 17.7 2.8 211 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 88: 2v96-A 17.6 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 89: 2hrq-C 17.6 2.8 211 531 14 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 90: 2ajd-D 17.6 2.9 213 729 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 91: 2ajc-D 17.6 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 92: 2ajb-A 17.6 2.7 209 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 93: 1w1i-A 17.6 2.9 212 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 94: 1r9m-B 17.6 2.9 212 733 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 95: 1orw-B 17.6 2.8 210 729 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 96: 1orw-A 17.6 2.9 211 729 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 97: 1orv-D 17.6 2.8 212 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 98: 1orv-C 17.6 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 99: 1orv-B 17.6 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 100: 1orv-A 17.6 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 101: 1mx1-C 17.6 2.7 210 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 102: 1h22-A 17.6 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 103: 1eve 17.6 2.7 212 534 13 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 104: 1aql-B 17.6 2.9 215 532 13 MOLECULE: BILE-SALT ACTIVATED LIPASE;
| |
| 105: 3bix-A 17.5 2.5 211 555 14 MOLECULE: NEUROLIGIN-1;
| |
| 106: 2pm8-B 17.5 2.8 212 530 16 MOLECULE: CHOLINESTERASE;
| |
| 107: 2hrr-C 17.5 2.8 211 531 14 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 108: 2gyu-B 17.5 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 109: 2gbf-A 17.5 2.9 210 730 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 110: 2gbc-A 17.5 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 111: 2buc-D 17.5 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 112: 2buc-C 17.5 2.7 209 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 113: 2buc-A 17.5 2.8 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 114: 2ajc-B 17.5 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 115: 2aj8-C 17.5 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 116: 1ya4-A 17.5 2.8 211 531 14 MOLECULE: CES1 PROTEIN;
| |
| 117: 1r1d-A 17.5 2.7 186 242 16 MOLECULE: CARBOXYLESTERASE;
| |
| 118: 1qon-A 17.5 2.9 209 540 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 119: 1qo9-A 17.5 2.9 212 540 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 120: 1mx9-A 17.5 2.9 212 532 14 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 121: 1c7i-A 17.5 2.8 213 483 15 MOLECULE: PARA-NITROBENZYL ESTERASE;
| |
| 122: 1acl 17.5 2.7 212 527 14 ACETYLCHOLINESTERASE (E.C.3.1.1.7) COMPLEXED WITH
| |
| 123: 1acj 17.5 2.7 212 528 14 ACETYLCHOLINESTERASE (E.C.3.1.1.7) COMPLEXED WITH TACRINE
| |
| 124: 2ocl-A 17.4 2.8 201 254 7 MOLECULE: VALACYCLOVIR HYDROLASE;
| |
| 125: 2jbw-B 17.4 3.7 220 360 15 MOLECULE: 2,6-DIHYDROXY-PSEUDO-OXYNICOTINE HYDROLASE;
| |
| 126: 2gbi-A 17.4 2.8 211 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 127: 2dqy-A 17.4 2.9 210 532 12 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 128: 2bua-D 17.4 2.8 210 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 129: 2bua-C 17.4 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 130: 2bua-B 17.4 2.8 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 131: 2ajb-C 17.4 2.8 209 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 132: 2ajb-B 17.4 2.8 211 728 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 133: 1yah-A 17.4 3.0 212 532 12 MOLECULE: CES1 PROTEIN;
| |
| 134: 1r1d-B 17.4 2.7 186 242 16 MOLECULE: CARBOXYLESTERASE;
| |
| 135: 1n5r-B 17.4 3.1 216 534 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 136: 1mx1-F 17.4 3.0 213 531 14 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 137: 1maa-B 17.4 3.1 217 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 138: 1h23-A 17.4 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 139: 1fss-A 17.4 2.7 212 532 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 140: 1c2o-B 17.4 3.1 217 539 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 141: 1akn 17.4 3.0 218 547 12 MOLECULE: BILE-SALT ACTIVATED LIPASE;
| |
| 142: 2va9-A 17.3 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 143: 2ock-A 17.3 2.8 201 254 8 MOLECULE: VALACYCLOVIR HYDROLASE;
| |
| 144: 2oci-A 17.3 2.8 201 255 8 MOLECULE: VALACYCLOVIR HYDROLASE;
| |
| 145: 2ocg-A 17.3 2.8 201 254 8 MOLECULE: VALACYCLOVIR HYDROLASE;
| |
| 146: 2hrq-F 17.3 2.9 212 531 14 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 147: 2ha6-B 17.3 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 148: 2ha5-B 17.3 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 149: 2gyw-A 17.3 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 150: 2dr0-A 17.3 2.9 211 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 151: 2ajd-B 17.3 2.8 209 729 12 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 152: 1zoi-B 17.3 2.6 187 274 17 MOLECULE: ESTERASE;
| |
| 153: 1r9n-A 17.3 2.9 213 728 14 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 154: 1q84-B 17.3 3.1 216 531 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 155: 1q83-B 17.3 3.1 216 531 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 156: 1nu8-A 17.3 3.0 213 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 157: 1n5m-B 17.3 3.1 216 531 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 158: 1n5m-A 17.3 3.1 216 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 159: 1mx9-F 17.3 2.9 213 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 160: 1mx9-E 17.3 2.8 209 531 12 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 161: 1mx1-B 17.3 2.7 209 531 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 162: 1k4y-A 17.3 2.8 210 501 14 MOLECULE: LIVER CARBOXYLESTERASE;
| |
| 163: 1j07-B 17.3 3.1 216 534 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 164: 1j06-A 17.3 3.1 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 165: 1gz7-C 17.3 2.8 212 534 13 MOLECULE: LIPASE 2;
| |
| 166: 1c2o-D 17.3 3.1 217 539 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 167: 1c2o-C 17.3 3.1 217 539 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 168: 1c2b-A 17.3 3.1 216 539 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 169: 2rgu-A 17.2 2.9 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 170: 2jf0-A 17.2 3.3 218 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 171: 2i3z-A 17.2 2.9 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4 (DIPEPTIDYL PEPTIDASE IV)
| |
| 172: 2dfp-A 17.2 2.7 213 534 13 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 173: 2cmf-A 17.2 2.6 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 174: 1zoi-C 17.2 2.7 189 275 17 MOLECULE: ESTERASE;
| |
| 175: 1rwq-A 17.2 2.9 211 728 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 176: 1qfs-A 17.2 2.9 220 710 11 MOLECULE: PROLYL OLIGOPEPTIDASE;
| |
| 177: 1q84-A 17.2 3.1 216 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 178: 1pfq-B 17.2 3.0 212 725 13 MOLECULE: DIPEPTIDYL PEPTIDASE IV SOLUBLE FORM;
| |
| 179: 1llf-A 17.2 2.8 211 534 13 MOLECULE: LIPASE 3;
| |
| 180: 1j07-A 17.2 3.1 216 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 181: 1gz7-A 17.2 2.8 212 534 13 MOLECULE: LIPASE 2;
| |
| 182: 2jey-B 17.1 3.4 221 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 183: 2jey-A 17.1 3.3 220 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 184: 2ha5-A 17.1 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 185: 2dqz-A 17.1 2.9 211 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 186: 1zoi-A 17.1 2.7 188 275 16 MOLECULE: ESTERASE;
| |
| 187: 1yaj-A 17.1 2.9 211 532 12 MOLECULE: CES1 PROTEIN;
| |
| 188: 1w75-A 17.1 2.7 212 528 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 189: 1vlq-I 17.1 3.0 205 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 190: 1vlq-F 17.1 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 191: 1q83-A 17.1 3.1 216 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 192: 1h2x-A 17.1 3.0 221 710 12 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 193: 1cle-A 17.1 2.8 212 534 13 MOLECULE: CHOLESTEROL ESTERASE;
| |
| 194: 1c7j-A 17.1 2.7 211 485 14 MOLECULE: PARA-NITROBENZYL ESTERASE;
| |
| 195: 1amn 17.1 2.7 212 526 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 196: 3bl8-A 17.0 2.6 212 556 14 MOLECULE: NEUROLIGIN-2;
| |
| 197: 2vh8-A 17.0 2.7 214 544 12 MOLECULE: NEUROLIGIN 4, X-LINKED;
| |
| 198: 2jgf-A 17.0 2.8 213 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 199: 2gyv-A 17.0 3.3 218 535 16 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 200: 2gbg-A 17.0 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 201: 2c0p-B 17.0 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 202: 1vlq-L 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 203: 1vlq-K 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 204: 1vlq-H 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 205: 1vlq-G 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 206: 1vlq-E 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 207: 1vlq-D 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 208: 1vlq-C 17.0 3.0 205 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 209: 1vlq-B 17.0 3.0 204 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 210: 1vlq-A 17.0 2.9 202 322 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 211: 1uoo-A 17.0 2.9 221 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 212: 1odt-C 17.0 2.9 206 316 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 213: 1ods-A 17.0 2.9 205 316 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 214: 1o6f-A 17.0 3.0 219 712 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 215: 1mx9-D 17.0 2.9 212 533 14 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 216: 1mx9-C 17.0 2.8 210 531 12 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 217: 1lpn 17.0 2.8 212 534 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL LIPASE) COMPLEXED WITH
| |
| 218: 1l7a-A 17.0 2.8 203 318 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 219: 1h2w-A 17.0 3.0 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 220: 2gyv-B 16.9 3.3 220 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 221: 2fj0-A 16.9 2.7 213 530 15 MOLECULE: JUVENILE HORMONE ESTERASE;
| |
| 222: 2c0q-B 16.9 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 223: 1vz3-A 16.9 2.9 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 224: 1vot 16.9 2.7 212 529 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 225: 1uoq-A 16.9 2.9 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 226: 1uop-A 16.9 2.9 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 227: 1tqh-A 16.9 2.6 184 242 16 MOLECULE: CARBOXYLESTERASE PRECURSOR;
| |
| 228: 1orw-C 16.9 2.8 211 729 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 229: 1odt-H 16.9 2.8 203 316 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 230: 1llf-B 16.9 2.9 212 534 13 MOLECULE: LIPASE 3;
| |
| 231: 1l7a-B 16.9 2.9 206 318 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 232: 1j2e-A 16.9 2.9 211 729 12 MOLECULE: DIPEPTIDYL PEPTIDASE IV;
| |
| 233: 1j06-B 16.9 3.1 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 234: 1h2y-A 16.9 3.0 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 235: 1e8m-A 16.9 2.9 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 236: 1e5t-A 16.9 2.8 217 710 12 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 237: 1ax9 16.9 2.7 212 527 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 238: 2ogs-A 16.8 2.9 212 479 16 MOLECULE: THERMOSTABLE CARBOXYLESTERASE EST50;
| |
| 239: 2jgm-A 16.8 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 240: 2gyw-B 16.8 3.2 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 241: 1vz2-A 16.8 2.9 220 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 242: 1lpp 16.8 2.8 212 534 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL LIPASE) COMPLEXED WITH
| |
| 243: 1lpm 16.8 2.8 212 534 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL LIPASE) COMPLEXED WITH
| |
| 244: 1o6g-A 16.7 3.0 221 712 12 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 245: 1n5r-A 16.7 3.1 216 536 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 246: 1maa-C 16.7 3.1 216 540 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 247: 1ku6-A 16.7 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 248: 1h2z-A 16.7 2.8 217 712 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 249: 3bl8-B 16.6 2.5 205 546 14 MOLECULE: NEUROLIGIN-2;
| |
| 250: 2gyu-A 16.6 3.1 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 251: 2c0p-A 16.6 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 252: 1yr2-A 16.6 2.7 205 680 12 MOLECULE: PROLYL OLIGOPEPTIDASE;
| |
| 253: 1qfm-A 16.6 3.0 217 710 12 MOLECULE: PROLYL OLIGOPEPTIDASE;
| |
| 254: 1gz7-D 16.6 3.0 213 534 13 MOLECULE: LIPASE 2;
| |
| 255: 2c0q-A 16.5 3.2 217 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 256: 2bce 16.5 2.8 208 532 13 MOLECULE: CHOLESTEROL ESTERASE;
| |
| 257: 1r88-B 16.5 3.1 205 267 10 MOLECULE: MPT51/MPB51 ANTIGEN;
| |
| 258: 1r88-A 16.5 3.0 205 267 10 MOLECULE: MPT51/MPB51 ANTIGEN;
| |
| 259: 1lpo 16.5 2.8 212 534 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL LIPASE) COMPLEXED WITH
| |
| 260: 1gz7-B 16.5 2.9 214 534 13 MOLECULE: LIPASE 2;
| |
| 261: 1e8n-A 16.5 3.0 219 710 11 MOLECULE: PROLYL ENDOPEPTIDASE;
| |
| 262: 1crl 16.5 2.8 212 534 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 263: 1cle-B 16.5 2.7 209 534 13 MOLECULE: CHOLESTEROL ESTERASE;
| |
| 264: 1c2o-A 16.5 3.1 217 539 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 265: 3bdi-A 16.4 2.9 182 207 14 MOLECULE: UNCHARACTERIZED PROTEIN TA0194;
| |
| 266: 1maa-A 16.4 3.0 216 540 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 267: 1lps 16.4 2.8 212 534 13 LIPASE (E.C.3.1.1.3) COMPLEXED WITH (1S)-MENTHYL HEXYL
| |
| 268: 1c4x-A 16.4 3.0 200 281 13 MOLECULE: 2-HYDROXY-6-OXO-6-PHENYLHEXA-2,4-DIENOATE
| |
| 269: 1dqy-A 16.3 2.9 208 283 12 MOLECULE: ANTIGEN 85-C;
| |
| 270: 1a88-A 16.3 2.6 189 275 20 MOLECULE: CHLOROPEROXIDASE L;
| |
| 271: 2oae-B 16.2 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 272: 1mtz-A 16.2 2.9 191 290 12 MOLECULE: PROLINE IMINOPEPTIDASE;
| |
| 273: 1mah-A 16.2 3.1 217 533 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 274: 1maa-D 16.2 3.1 215 541 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 275: 1f8u-A 16.2 2.8 212 531 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 276: 1a88-B 16.2 2.6 188 275 19 MOLECULE: CHLOROPEROXIDASE L;
| |
| 277: 2veo-A 16.1 3.0 218 430 13 MOLECULE: LIPASE A;
| |
| 278: 2gbg-B 16.1 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 279: 2gbf-B 16.1 2.8 211 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 280: 1k8q-A 16.1 3.1 211 377 14 MOLECULE: TRIACYLGLYCEROL LIPASE, GASTRIC;
| |
| 281: 1b41-A 16.1 2.8 211 531 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 282: 1a88-C 16.1 2.7 189 275 19 MOLECULE: CHLOROPEROXIDASE L;
| |
| 283: 2uz0-A 16.0 3.1 201 253 12 MOLECULE: TRIBUTYRIN ESTERASE;
| |
| 284: 2i3z-B 16.0 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4 (DIPEPTIDYL PEPTIDASE IV)
| |
| 285: 2i3d-B 16.0 2.8 186 220 13 MOLECULE: HYPOTHETICAL PROTEIN ATU1826;
| |
| 286: 2gbi-B 16.0 2.9 210 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 287: 2gbc-B 16.0 2.8 212 730 13 MOLECULE: DIPEPTIDYL PEPTIDASE 4;
| |
| 288: 1yaj-D 16.0 2.9 210 532 12 MOLECULE: CES1 PROTEIN;
| |
| 289: 1trh 16.0 2.8 209 534 12 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 290: 1mu0-A 16.0 2.9 193 294 12 MOLECULE: PROLINE IMINOPEPTIDASE;
| |
| 291: 1j1i-A 16.0 3.3 196 258 12 MOLECULE: META CLEAVAGE COMPOUND HYDROLASE;
| |
| 292: 1auo-A 16.0 2.3 172 218 15 MOLECULE: CARBOXYLESTERASE;
| |
| 293: 2fuk-A 15.9 2.7 186 218 14 MOLECULE: XC6422 PROTEIN;
| |
| 294: 1ukc-A 15.9 2.9 210 517 13 MOLECULE: ESTA;
| |
| 295: 1a8s 15.9 2.7 188 273 19 MOLECULE: CHLOROPEROXIDASE F;
| |
| 296: 1f0p-A 15.8 3.1 208 284 13 MOLECULE: ANTIGEN 85-B;
| |
| 297: 1aur-A 15.8 2.3 172 218 15 MOLECULE: CARBOXYLESTERASE;
| |
| 298: 1a8q 15.8 2.8 192 274 15 MOLECULE: BROMOPEROXIDASE A1;
| |
| 299: 1u2e-A 15.7 3.1 195 286 14 MOLECULE: 2-HYDROXY-6-KETONONA-2,4-DIENEDIOIC ACID
| |
| 300: 1iuo-A 15.7 3.0 187 272 12 MOLECULE: META-CLEAVAGE PRODUCT HYDROLASE;
| |
| 301: 1iun-B 15.7 3.3 200 276 12 MOLECULE: META-CLEAVAGE PRODUCT HYDROLASE;
| |
| 302: 2b4k-A 15.6 3.2 203 617 12 MOLECULE: ALPHA-AMINO ACID ESTER HYDROLASE;
| |
| 303: 1va5-A 15.6 3.0 206 283 11 MOLECULE: ANTIGEN 85-C;
| |
| 304: 1va4-A 15.6 2.8 189 271 16 MOLECULE: ARYLESTERASE;
| |
| 305: 1sfr-A 15.6 3.2 206 288 12 MOLECULE: ANTIGEN 85-A;
| |
| 306: 1iup-A 15.6 3.3 197 271 10 MOLECULE: META-CLEAVAGE PRODUCT HYDROLASE;
| |
| 307: 1dqz-B 15.6 2.9 203 280 11 MOLECULE: ANTIGEN 85-C;
| |
| 308: 1dqz-A 15.6 3.0 205 280 11 MOLECULE: ANTIGEN 85-C;
| |
| 309: 2pu5-A 15.5 3.1 198 285 14 MOLECULE: 2-HYDROXY-6-OXO-6-PHENYLHEXA-2,4-DIENOATE
| |
| 310: 1hkh-A 15.5 2.6 191 279 15 MOLECULE: GAMMA LACTAMASE;
| |
| 311: 1f0n-A 15.5 3.2 205 284 11 MOLECULE: ANTIGEN 85B;
| |
| 312: 1pv1-A 15.4 3.3 202 290 13 MOLECULE: HYPOTHETICAL 33.9 KDA ESTERASE IN SMC3-MRPL8
| |
| 313: 1iun-A 15.4 3.4 197 272 11 MOLECULE: META-CLEAVAGE PRODUCT HYDROLASE;
| |
| 314: 1hl7-A 15.4 2.6 188 279 15 MOLECULE: GAMMA LACTAMASE;
| |
| 315: 1bro-A 15.4 2.6 189 277 18 MOLECULE: BROMOPEROXIDASE A2;
| |
| 316: 1a8u-B 15.4 2.7 188 277 18 MOLECULE: CHLOROPEROXIDASE T;
| |
| 317: 1a7u-A 15.4 2.8 192 277 18 MOLECULE: CHLOROPEROXIDASE T;
| |
| 318: 1hlg-A 15.3 3.1 208 368 14 MOLECULE: LIPASE, GASTRIC;
| |
| 319: 1thg 15.2 3.0 210 544 11 LIPASE (E.C.3.1.1.3) TRIACYLGLYCEROL HYDROLASE
| |
| 320: 1a8u-A 15.2 2.7 185 277 17 MOLECULE: CHLOROPEROXIDASE T;
| |
| 321: 1a7u-B 15.2 2.7 188 277 18 MOLECULE: CHLOROPEROXIDASE T;
| |
| 322: 2d5l-A 15.1 3.0 216 665 13 MOLECULE: DIPEPTIDYL AMINOPEPTIDASE IV, PUTATIVE;
| |
| 323: 3b5e-B 15.0 2.6 176 213 15 MOLECULE: MLL8374 PROTEIN;
| |
| 324: 1r3d-A 15.0 3.0 180 257 17 MOLECULE: CONSERVED HYPOTHETICAL PROTEIN VC1974;
| |
| 325: 1fj2-A 15.0 2.7 177 229 15 MOLECULE: ACYL PROTEIN THIOESTERASE 1;
| |
| 326: 2yxp-X 14.9 3.0 199 310 11 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 327: 2vf2-A 14.9 3.3 202 284 14 MOLECULE: 2-HYDROXY-6-OXO-6-PHENYLHEXA-2,4-DIENOATE
| |
| 328: 2pky-X 14.9 3.0 199 308 13 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 329: 1brt 14.9 2.7 188 277 16 MOLECULE: BROMOPEROXIDASE A2;
| |
| 330: 1bro-B 14.9 2.8 186 277 17 MOLECULE: BROMOPEROXIDASE A2;
| |
| 331: 1q0z-A 14.8 2.8 190 297 11 MOLECULE: ACLACINOMYCIN METHYLESTERASE;
| |
| 332: 1q0r-A 14.8 2.7 190 297 13 MOLECULE: ACLACINOMYCIN METHYLESTERASE;
| |
| 333: 1jfr-A 14.8 2.6 175 260 14 MOLECULE: LIPASE;
| |
| 334: 1cij-A 14.8 3.0 200 310 13 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 335: 1b6g 14.8 3.0 198 310 12 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 336: 3bwx-A 14.7 3.2 192 285 13 MOLECULE: ALPHA/BETA HYDROLASE;
| |
| 337: 1zd3-A 14.7 3.1 201 547 11 MOLECULE: EPOXIDE HYDROLASE 2, CYTOPLASMIC;
| |
| 338: 1mpx-A 14.7 3.4 202 614 14 MOLECULE: ALPHA-AMINO ACID ESTER HYDROLASE;
| |
| 339: 1ede 14.7 2.9 197 310 12 HALOALKANE DEHALOGENASE (E.C.3.8.1.5) AT PH 8.2
| |
| 340: 1be0 14.7 3.1 201 310 11 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 341: 2r11-D 14.6 3.6 203 288 10 MOLECULE: CARBOXYLESTERASE NP;
| |
| 342: 1zd5-A 14.6 3.1 201 547 11 MOLECULE: EPOXIDE HYDROLASE 2, CYTOPLASMIC;
| |
| 343: 1tht-A 14.6 2.8 192 294 10 THIOESTERASE
| |
| 344: 1jt2-A 14.6 2.7 179 255 16 MOLECULE: ENDO-1,4-BETA-XYLANASE Z;
| |
| 345: 1ivy-A 14.6 3.0 205 452 10 MOLECULE: HUMAN PROTECTIVE PROTEIN;
| |
| 346: 1hde-A 14.6 2.9 197 310 12 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 347: 1edb 14.6 3.1 200 310 12 HALOALKANE DEHALOGENASE (E.C.3.8.1.5) COMPLEX WITH CHLORIDE
| |
| 348: 2fx5-A 14.5 2.4 171 258 16 MOLECULE: LIPASE;
| |
| 349: 1zic-A 14.5 2.6 177 233 14 MOLECULE: CARBOXYMETHYLENEBUTENOLIDASE;
| |
| 350: 1vj5-A 14.5 3.3 200 547 11 MOLECULE: EPOXIDE HYDROLASE 2, CYTOPLASMIC;
| |
| 351: 1aur-B 14.5 2.4 165 218 14 MOLECULE: CARBOXYLESTERASE;
| |
| 352: 1auo-B 14.5 2.4 165 218 15 MOLECULE: CARBOXYLESTERASE;
| |
| 353: 2yys-A 14.4 3.3 196 283 13 MOLECULE: PROLINE IMINOPEPTIDASE-RELATED PROTEIN;
| |
| 354: 1ggv-A 14.4 2.7 176 232 13 MOLECULE: DIENELACTONE HYDROLASE;
| |
| 355: 1azw-A 14.4 3.1 198 313 16 MOLECULE: PROLINE IMINOPEPTIDASE;
| |
| 356: 1azw-B 14.3 3.2 199 313 14 MOLECULE: PROLINE IMINOPEPTIDASE;
| |
| 357: 1qtr-A 14.2 3.2 199 314 15 MOLECULE: PROLYL AMINOPEPTIDASE;
| |
| 358: 1imj-A 14.2 2.8 176 208 13 MOLECULE: CCG1-INTERACTING FACTOR B;
| |
| 359: 1e3q-A 14.2 3.1 200 533 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 360: 2r8b-A 14.1 3.0 170 204 15 MOLECULE: UNCHARACTERIZED PROTEIN ATU2452;
| |
| 361: 1lns-A 14.1 3.1 203 763 11 MOLECULE: X-PROLYL DIPEPTIDYL AMINOPETIDASE;
| |
| 362: 1i6w-A 14.1 2.5 158 179 16 MOLECULE: LIPASE A;
| |
| 363: 2qxt-A 14.0 2.5 157 179 16 MOLECULE: LIPASE;
| |
| 364: 1y37-A 14.0 3.1 198 294 11 MOLECULE: FLUOROACETATE DEHALOGENASE;
| |
| 365: 1wom-A 13.9 3.0 186 271 13 MOLECULE: SIGMA FACTOR SIGB REGULATION PROTEIN RSBQ;
| |
| 366: 1vlq-J 13.9 3.3 195 321 15 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 367: 1iz8-A 13.9 3.3 201 294 9 MOLECULE: HALOALKANE DEHALOGENASE, LINB;
| |
| 368: 1ek2-B 13.9 3.2 199 541 10 MOLECULE: EPOXIDE HYDROLASE;
| |
| 369: 1ek1-B 13.9 3.0 198 541 11 MOLECULE: EPOXIDE HYDROLASE;
| |
| 370: 1din 13.9 2.8 175 232 13 MOLECULE: DIENELACTONE HYDROLASE;
| |
| 371: 1ycd-B 13.8 2.7 166 238 14 MOLECULE: HYPOTHETICAL 27.3 KDA PROTEIN IN AAP1-SMF2
| |
| 372: 1tca 13.8 2.8 183 317 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 373: 1mj5-A 13.8 3.2 197 297 9 MOLECULE: 1,3,4,6-TETRACHLORO-1,4-CYCLOHEXADIENE HYDROLASE;
| |
| 374: 1lbt 13.8 2.8 183 317 11 MOLECULE: LIPASE B;
| |
| 375: 1cqz-B 13.8 3.3 201 541 9 MOLECULE: EPOXIDE HYDROLASE;
| |
| 376: 2b20-A 13.7 3.2 199 391 12 MOLECULE: ENTEROCHELIN ESTERASE;
| |
| 377: 1tcc-B 13.7 2.9 183 317 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 378: 1qlw-A 13.7 2.7 184 318 15 MOLECULE: ESTERASE;
| |
| 379: 1l7r-A 13.7 2.9 196 573 17 MOLECULE: COCAINE ESTERASE;
| |
| 380: 2qm0-B 13.6 2.7 186 268 15 MOLECULE: IROE PROTEIN;
| |
| 381: 2h1i-A 13.6 3.0 174 212 16 MOLECULE: CARBOXYLESTERASE;
| |
| 382: 1tcc-A 13.6 2.9 183 317 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 383: 1tcb-B 13.6 3.0 183 317 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 384: 1tcb-A 13.6 2.9 184 317 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 385: 1ju3-A 13.6 3.0 195 570 17 MOLECULE: COCAINE ESTERASE;
| |
| 386: 1jjf-A 13.6 2.8 178 255 14 MOLECULE: ENDO-1,4-BETA-XYLANASE Z;
| |
| 387: 2qmq-A 13.5 3.4 190 278 9 MOLECULE: PROTEIN NDRG2;
| |
| 388: 1nx9-A 13.5 3.2 200 617 12 MOLECULE: ALPHA-AMINO ACID ESTER HYDROLASE;
| |
| 389: 1i6w-B 13.5 2.7 158 180 14 MOLECULE: LIPASE A;
| |
| 390: 3c5v-A 13.4 3.3 196 294 11 MOLECULE: PROTEIN PHOSPHATASE METHYLESTERASE 1;
| |
| 391: 1wb4-A 13.4 2.7 179 283 14 MOLECULE: ENDO-1,4-BETA-XYLANASE Y;
| |
| 392: 1m33-A 13.4 2.8 174 255 13 MOLECULE: BIOH PROTEIN;
| |
| 393: 1cr6-B 13.4 3.2 195 541 10 MOLECULE: EPOXIDE HYDROLASE;
| |
| 394: 1cqw-A 13.4 3.4 196 295 12 MOLECULE: HALOALKANE DEHALOGENASE; 1-CHLOROHEXANE
| |
| 395: 2zjf-A 13.3 3.0 198 346 13 MOLECULE: PROBABLE EPOXIDE HYDROLASE EPHB;
| |
| 396: 1mx1-D 13.3 3.1 193 531 11 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 397: 1gkk-A 13.3 2.8 180 283 14 MOLECULE: ENDO-1,4-BETA-XYLANASE Y;
| |
| 398: 3bf7-A 13.1 2.9 177 255 16 MOLECULE: ESTERASE YBFF;
| |
| 399: 1wpx-A 13.1 3.2 210 421 12 MOLECULE: CARBOXYPEPTIDASE Y;
| |
| 400: 1ods-H 13.1 3.4 194 316 13 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 401: 1ods-F 13.1 3.4 194 316 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 402: 1ods-C 13.1 3.4 194 316 14 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 403: 1ysc 13.0 3.1 206 421 13 SERINE CARBOXYPEPTIDASE (CPY, CPD-Y, OR PROTEINASE C)
| |
| 404: 1cpy 13.0 3.2 208 421 13 MOLECULE: SERINE CARBOXYPEPTIDASE;
| |
| 405: 2cjp-A 12.9 3.0 190 320 12 MOLECULE: EPOXIDE HYDROLASE;
| |
| 406: 1gkl-A 12.9 2.8 174 283 14 MOLECULE: ENDO-1,4-BETA-XYLANASE Y;
| |
| 407: 1ehy-D 12.9 3.3 194 282 11 MOLECULE: SOLUBLE EPOXIDE HYDROLASE;
| |
| 408: 1ehy-A 12.9 3.3 195 282 11 MOLECULE: SOLUBLE EPOXIDE HYDROLASE;
| |
| 409: 1ehy-C 12.8 3.2 191 282 12 MOLECULE: SOLUBLE EPOXIDE HYDROLASE;
| |
| 410: 1ehy-B 12.8 3.3 195 282 12 MOLECULE: SOLUBLE EPOXIDE HYDROLASE;
| |
| 411: 2b61-A 12.7 3.2 191 357 8 MOLECULE: HOMOSERINE O-ACETYLTRANSFERASE;
| |
| 412: 1pja-A 12.7 2.6 168 268 14 MOLECULE: PALMITOYL-PROTEIN THIOESTERASE 2 PRECURSOR;
| |
| 413: 1ods-G 12.7 3.5 198 316 13 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 414: 1ods-E 12.7 3.6 198 316 13 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 415: 1ods-B 12.7 3.6 200 316 13 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 416: 1sci-A 12.6 3.1 171 256 15 MOLECULE: (S)-ACETONE-CYANOHYDRIN LYASE;
| |
| 417: 1ods-D 12.6 3.6 198 316 13 MOLECULE: CEPHALOSPORIN C DEACETYLASE;
| |
| 418: 1fj2-B 12.6 2.8 165 229 15 MOLECULE: ACYL PROTEIN THIOESTERASE 1;
| |
| 419: 1y7i-B 12.4 3.2 174 262 16 MOLECULE: SALICYLIC ACID-BINDING PROTEIN 2;
| |
| 420: 1kez-A 12.4 3.4 184 267 16 MOLECULE: ERYTHRONOLIDE SYNTHASE;
| |
| 421: 1jmk-C 12.4 3.0 164 222 11 MOLECULE: SURFACTIN SYNTHETASE;
| |
| 422: 1xkt-A 12.3 3.7 178 260 13 MOLECULE: FATTY ACID SYNTHASE;
| |
| 423: 1jmk-O 12.2 3.3 173 230 12 MOLECULE: SURFACTIN SYNTHETASE;
| |
| 424: 1ac5 12.2 3.4 209 483 10 MOLECULE: KEX1(DELTA)P;
| |
| 425: 1exw-A 12.1 3.1 169 279 11 MOLECULE: PALMITOYL PROTEIN THIOESTERASE 1;
| |
| 426: 2dr0-C 12.0 3.6 195 532 14 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 427: 1mna-B 12.0 3.6 180 278 14 MOLECULE: POLYKETIDE SYNTHASE IV;
| |
| 428: 1ek1-A 11.9 3.4 197 490 8 MOLECULE: EPOXIDE HYDROLASE;
| |
| 429: 1ei9-A 11.9 2.7 164 279 10 MOLECULE: PALMITOYL PROTEIN THIOESTERASE 1;
| |
| 430: 1eh5-A 11.9 3.1 170 279 10 MOLECULE: PALMITOYL PROTEIN THIOESTERASE 1;
| |
| 431: 7yas-A 11.8 3.1 172 256 15 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 432: 5yas-A 11.8 3.2 174 256 16 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 433: 3yas-A 11.8 3.0 171 256 15 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 434: 2yas-A 11.8 3.1 172 256 15 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 435: 2cut 11.8 3.1 151 198 13 CUTINASE (E.C.3.1.1.-) COMPLEXED WITH THE INHIBITOR DIETHYL
| |
| 436: 1yb6-A 11.8 3.1 175 256 15 MOLECULE: (S)-ACETONE-CYANOHYDRIN LYASE;
| |
| 437: 1yas-A 11.8 3.1 172 256 15 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 438: 1ek2-A 11.8 3.2 194 487 11 MOLECULE: EPOXIDE HYDROLASE;
| |
| 439: 2dqy-C 11.7 3.6 195 532 14 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 440: 1qj4-A 11.7 3.0 169 256 15 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 441: 1mn6-B 11.7 3.4 178 278 15 MOLECULE: POLYKETIDE SYNTHASE IV;
| |
| 442: 1cex 11.7 3.0 147 197 14 MOLECULE: CUTINASE;
| |
| 443: 2dqz-C 11.6 3.7 195 532 13 MOLECULE: LIVER CARBOXYLESTERASE 1;
| |
| 444: 1yah-C 11.6 3.8 200 532 14 MOLECULE: CES1 PROTEIN;
| |
| 445: 1ya4-C 11.6 3.8 200 532 14 MOLECULE: CES1 PROTEIN;
| |
| 446: 1yaj-C 11.5 3.8 198 532 13 MOLECULE: CES1 PROTEIN;
| |
| 447: 1p0p-A 11.5 5.8 200 522 14 MOLECULE: CHOLINESTERASE;
| |
| 448: 1hqd-A 11.5 2.9 170 320 13 MOLECULE: LIPASE;
| |
| 449: 3lip 11.4 2.9 172 320 13 MOLECULE: TRIACYL-GLYCEROL-HYDROLASE;
| |
| 450: 2lip 11.4 3.0 174 320 14 MOLECULE: LIPASE;
| |
| 451: 1yaj-F 11.4 3.8 200 532 13 MOLECULE: CES1 PROTEIN;
| |
| 452: 1oil-A 11.4 2.9 172 320 13 MOLECULE: LIPASE;
| |
| 453: 1eb9-A 11.4 3.0 168 262 14 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 454: 1e89-A 11.4 3.1 171 262 13 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 455: 2jbw-D 11.3 3.8 207 359 16 MOLECULE: 2,6-DIHYDROXY-PSEUDO-OXYNICOTINE HYDROLASE;
| |
| 456: 1ya4-B 11.3 3.7 199 532 14 MOLECULE: CES1 PROTEIN;
| |
| 457: 1cqz-A 11.3 3.2 198 487 11 MOLECULE: EPOXIDE HYDROLASE;
| |
| 458: 2hih-A 11.2 3.1 185 387 15 MOLECULE: LIPASE 46 KDA FORM;
| |
| 459: 1cvl 11.2 3.1 168 316 13 MOLECULE: TRIACYLGLYCEROL HYDROLASE;
| |
| 460: 1cr6-A 11.2 3.4 202 487 11 MOLECULE: EPOXIDE HYDROLASE;
| |
| 461: 1qo7-A 11.1 3.1 185 385 9 MOLECULE: EPOXIDE HYDROLASE;
| |
| 462: 1mx9-G 11.1 3.7 196 532 13 MOLECULE: LIVER CARBOXYLESTERASE I;
| |
| 463: 1yaj-G 10.9 3.8 197 532 13 MOLECULE: CES1 PROTEIN;
| |
| 464: 1yaj-E 10.9 3.8 197 532 13 MOLECULE: CES1 PROTEIN;
| |
| 465: 1tah-C 10.9 3.2 174 318 12 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 466: 1tah-A 10.9 3.1 171 318 12 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 467: 1dwq-A 10.9 3.0 166 262 14 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 468: 1tah-D 10.8 3.3 176 318 13 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 469: 1tah-B 10.8 3.1 170 318 12 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 470: 2gzs-A 10.7 3.3 179 249 13 MOLECULE: IROE PROTEIN;
| |
| 471: 1qoz-A 10.7 3.3 154 207 11 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 472: 1p0q-A 10.6 5.8 200 522 13 MOLECULE: CHOLINESTERASE;
| |
| 473: 1w52-X 10.5 3.8 172 448 13 MOLECULE: PANCREATIC LIPASE RELATED PROTEIN 2;
| |
| 474: 2pvs-B 10.4 3.8 173 445 13 MOLECULE: PANCREATIC LIPASE-RELATED PROTEIN 2;
| |
| 475: 1g66-A 10.3 3.2 150 207 15 MOLECULE: ACETYL XYLAN ESTERASE II;
| |
| 476: 1cus 10.3 3.4 151 197 11 CUTINASE (E.C.3.1.1.-)
| |
| 477: 1ku0-A 9.9 3.3 179 388 11 MOLECULE: L1 LIPASE;
| |
| 478: 1bn6-A 9.9 3.4 161 291 14 MOLECULE: HALOALKANE DEHALOGENASE;
| |
| 479: 2axe 9.8 3.4 149 207 15 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 480: 1ku0-B 9.8 3.3 173 385 12 MOLECULE: L1 LIPASE;
| |
| 481: 1ji3-A 9.8 3.2 175 388 12 MOLECULE: LIPASE;
| |
| 482: 2ppl-A 9.7 3.3 166 449 13 MOLECULE: PANCREATIC LIPASE-RELATED PROTEIN 1;
| |
| 483: 1rp1 9.5 4.1 173 441 12 MOLECULE: PANCREATIC LIPASE RELATED PROTEIN 1;
| |
| 484: 1hpl-A 9.4 3.9 168 449 11 LIPASE (E.C.3.1.1.3) (TRIACYLGLYCEROL HYDROLASE)
| |
| 485: 1n8s-A 9.3 3.2 164 449 12 MOLECULE: TRIACYLGLYCEROL LIPASE, PANCREATIC;
| |
| 486: 1bu8-A 9.3 4.0 174 446 13 MOLECULE: PANCREATIC LIPASE RELATED PROTEIN 2;
| |
| 487: 2ha0-B 9.1 9.2 199 536 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 488: 1gpl 9.1 3.9 173 432 12 MOLECULE: RP2 LIPASE;
| |
| 489: 1bs9 8.8 3.1 135 207 16 MOLECULE: ACETYL XYLAN ESTERASE;
| |
| 490: 2jgg-B 8.7 12.1 186 533 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 491: 2jgj-B 8.6 12.2 185 533 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 492: 2jgh-B 8.6 12.2 186 532 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 493: 3bl8-D 8.5 5.5 185 541 13 MOLECULE: NEUROLIGIN-2;
| |
| 494: 2jgf-B 8.4 12.2 183 533 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 495: 2ha3-B 8.4 11.5 185 534 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 496: 2h9y-B 8.4 11.4 184 534 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 497: 2ha4-B 8.3 10.7 185 535 13 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 498: 2jgi-B 8.2 12.1 186 532 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 499: 1wht-A 8.2 3.9 143 256 10 SERINE CARBOXYPEPTIDASE II (E.C.3.4.16.1) COMPLEXED WITH
| |
| 500: 2jgk-B 8.0 12.1 185 530 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 501: 2jge-B 7.7 12.2 182 532 11 MOLECULE: ACETYLCHOLINESTERASE; SYNONYM:ACHE;
| |
| 502: 1jmy-A 7.1 12.2 185 515 11 MOLECULE: BILE-SALT-ACTIVATED LIPASE;
| |
| 503: 1gxs-A 7.1 3.9 138 267 9 MOLECULE: HYDROXYNITRILE LYASE;
| |
| 504: 2jez-B 6.6 12.0 182 533 12 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 505: 2ha7-B 6.5 11.5 185 534 10 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 506: 2ha2-B 6.5 11.5 185 534 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 507: 2ha0-A 5.8 10.8 180 535 14 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 508: 2jge-A 4.9 3.5 143 535 13 MOLECULE: ACETYLCHOLINESTERASE; SYNONYM:ACHE;
| |
| 509: 1uxo-A 4.7 3.4 135 186 11 MOLECULE: YDEN PROTEIN;
| |
| 510: 3be8-B 4.4 8.0 168 534 11 MOLECULE: NEUROLIGIN-4, X-LINKED;
| |
| 511: 2jgg-A 3.7 3.6 136 535 15 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 512: 2jgl-B 2.3 12.2 185 533 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 513: 2jgm-B 2.2 12.1 186 530 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
| 514: 2jez-A 2.0 4.0 122 535 11 MOLECULE: ACETYLCHOLINESTERASE;
| |
|
| |
|
| | ''Sections of alignment showing the conserved residues across bacterial and eukaryotic species.'' |
|
| |
|
| Notation:
| | [[Image:Alignment1.jpg]] |
|
| |
|
| * Chain: PDB entry code plus chain identifier
| | |
| * Z: normalized Z-score that depends on the size of the structures. The program optimises a weighted sum of similarities of intramolecular distances.
| | [[Arylformamidase | Return to the main page...]] |
| * rmsd: root-mean-square deviation of C-alpha atoms in the least-squares superimposition of the structurally equivalent C-alpha atoms. The program does not optimise rmsd, this is only reported for your information.
| |
| * lali: number of structurally equivalent residues
| |
| * nres: number of amino acids in the protein
| |
| * %id: percentage of identical amino acids over all structurally equivalent residues
| |
| * Description: the COMPND record from the PDB entry
| |