Structural analysis of 2qgn: Difference between revisions
Choojinhsien (talk | contribs) No edit summary |
Choojinhsien (talk | contribs) |
||
| Line 19: | Line 19: | ||
==Domains== | ==Domains== | ||
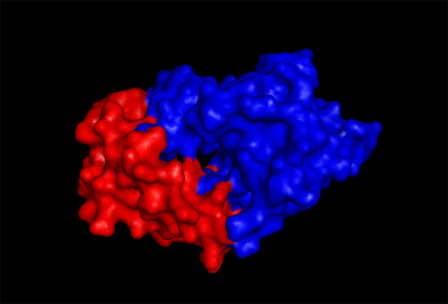
2qgnA is composed of two main domains. | |||
[[Image:domain pic.jpg|framed|'''Figure 4''' <BR>Two main domains exhibited by tRNA isopentenyl transferase(2qgnA). Blue regions denote one domain while Red regions underlying another domain.|none]]<BR> | [[Image:domain pic.jpg|framed|'''Figure 4''' <BR>Two main domains exhibited by tRNA isopentenyl transferase(2qgnA). Blue regions denote one domain while Red regions underlying another domain.|none]]<BR> | ||
| Line 25: | Line 25: | ||
[[Image:domain--2.jpg]] | [[Image:domain--2.jpg]] | ||
[[Image:domain--1.jpg]] | [[Image:domain--1.jpg]] | ||
==Ligand Binding Sites and Surface Clefts== | ==Ligand Binding Sites and Surface Clefts== | ||
Revision as of 12:55, 30 May 2008
Protein Sequence in FASTA format
>gi|152149497|pdb|2QGN|A Chain A, Crystal Structure Of Trna Isopentenylpyrophosphate Transferase (Bh2366) From Bacillus Halodurans, Northeast Structural Genomics Consortium Target Bhr41. XKEKLVAIVGPTAVGKTKTSVXLAKRLNGEVISGDSXQVYRGXDIGTAKITAEEXDGVPHHLIDIKDPSE SFSVADFQDLATPLITEIHERGRLPFLVGGTGLYVNAVIHQFNLGDIRADEDYRHELEAFVNSYGVQALH DKLSKIDPKAAAAIHPNNYRRVIRALEIIKLTGKTVTEQARHEEETPSPYNLVXIGLTXERDVLYDRINR RVDQXVEEGLIDEAKKLYDRGIRDCQSVQAIGYKEXYDYLDGNVTLEEAIDTLKRNSRRYAKRQLTWFRN KANVTWFDXTDVDFDKKIXEIHNFIAGKLEEKSKLEHHHHHH
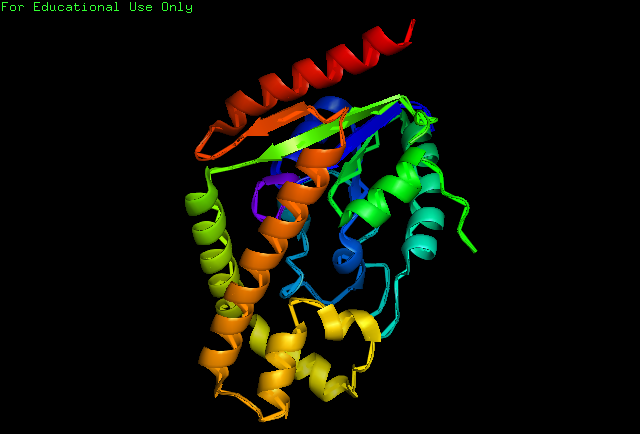
Structure of Protein
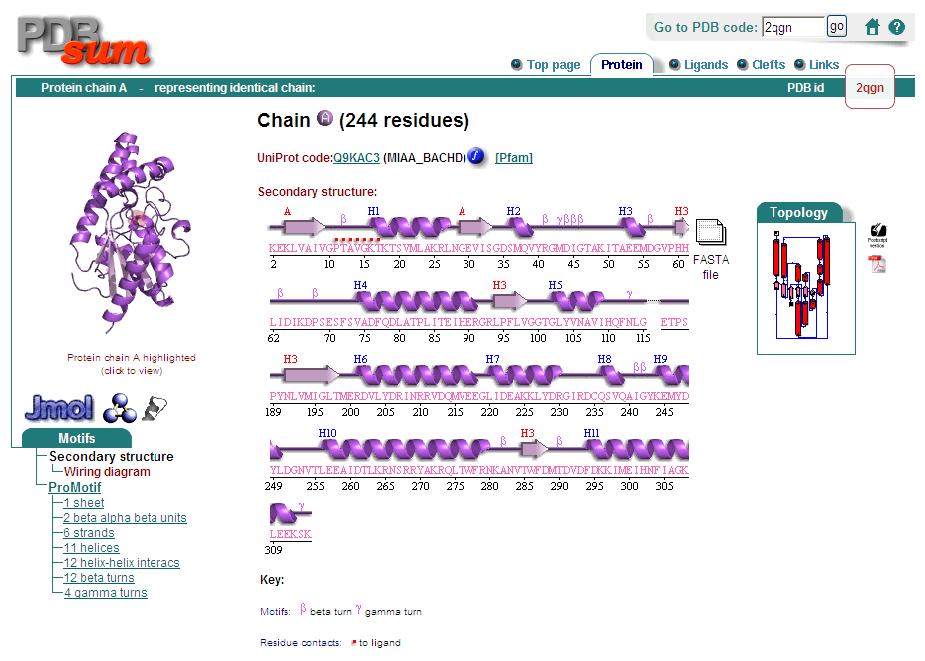
Secondary Structure
Analysis of the secondary structure acquired from Protein Data Bank showed results as displayed below :
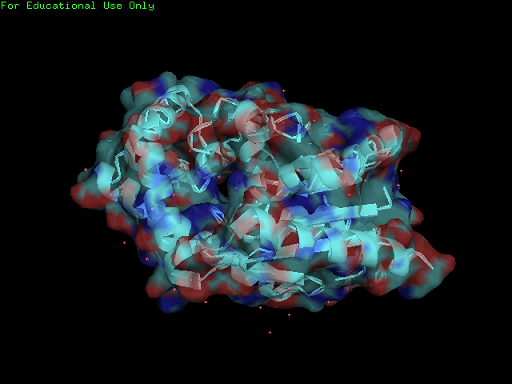
Surface Properties of 2qgn
Domains
2qgnA is composed of two main domains.
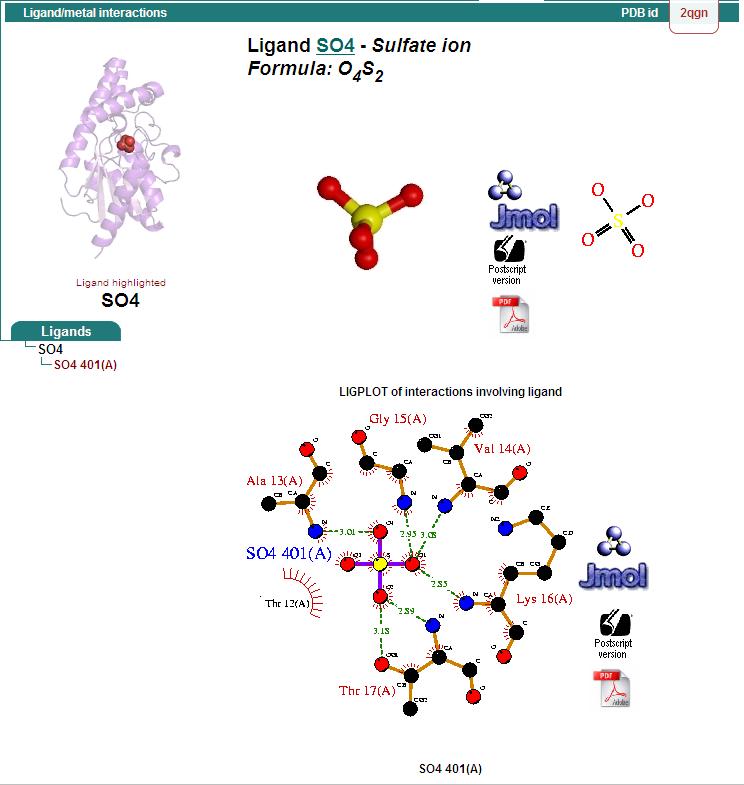
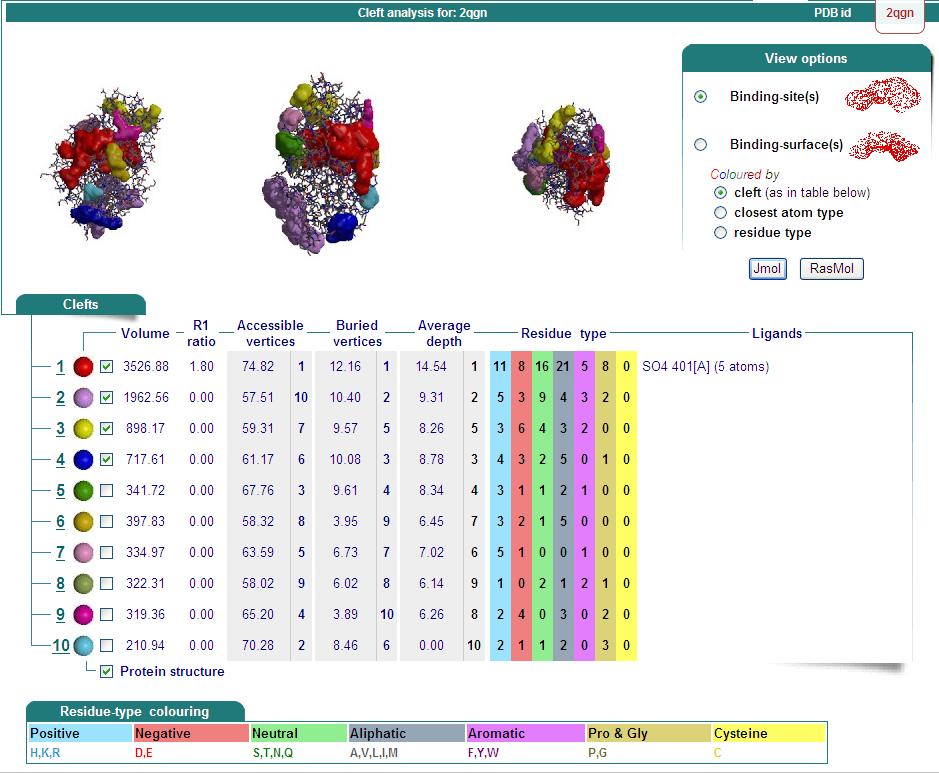
Ligand Binding Sites and Surface Clefts
Structural Alignment
Dali Output
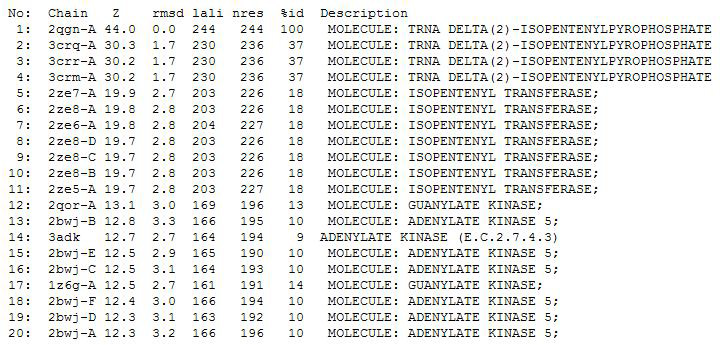
PDB entry code for 2qgn was loaded onto DALI to search for structurally similar neighbours. Displayed below are the results from DALI search :-
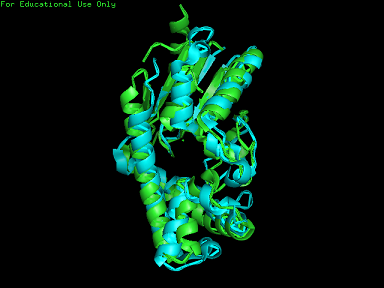
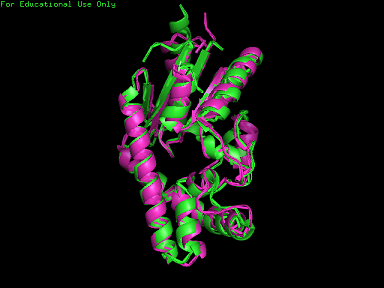
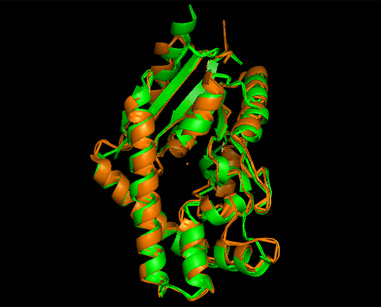
A total of 527 hits were found from DALI search, thus only the first 20 hits that may be of significance were shown on the figure. Based on the outcome of DALI, PDB files of each structurally similar protein was obtained from PDB. These were each superimposed with 2qgn on PyMOL, to compare the structural similiarity. Results are as below :