Evolution: Difference between revisions
JasonCheong (talk | contribs) No edit summary |
|||
| (25 intermediate revisions by 2 users not shown) | |||
| Line 6: | Line 6: | ||
See '''[[Materials and Methods]]''' | See '''[[Materials and Methods]]''' | ||
The following query protein sequence of N-acetylneuraminic Acid was obtained from GenBank: | The following query protein sequence of N-acetylneuraminic Acid was obtained from GenBank: | ||
| Line 12: | Line 11: | ||
{| border="1" cellpadding="15" cellspacing="0" | {| border="1" cellpadding="15" cellspacing="0" | ||
|+''' | |+'''arylsulfatase K'''|} | ||
|} | |||
'''Fig. 1:''' '''Amino Acid Sequence (260 aa)''' of N-acetylneuraminic Acid (2gfh) protein from ''Mus musculus'' (House Mouse) | '''Fig. 1:''' '''Amino Acid Sequence (260 aa)''' of N-acetylneuraminic Acid (2gfh) protein from ''Mus musculus'' (House Mouse) | ||
| Line 35: | Line 24: | ||
The bacteria sequence matches used for multiple sequence analysis and phylogenetic tree mapping were representative of the other baterial sequences not selected. These selected sequences had the highest bit scores and E-values. | The bacteria sequence matches used for multiple sequence analysis and phylogenetic tree mapping were representative of the other baterial sequences not selected. These selected sequences had the highest bit scores and E-values. | ||
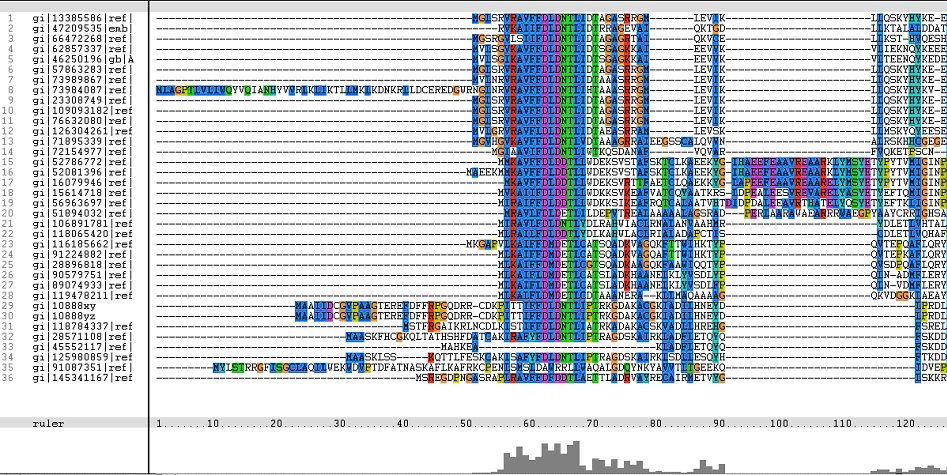
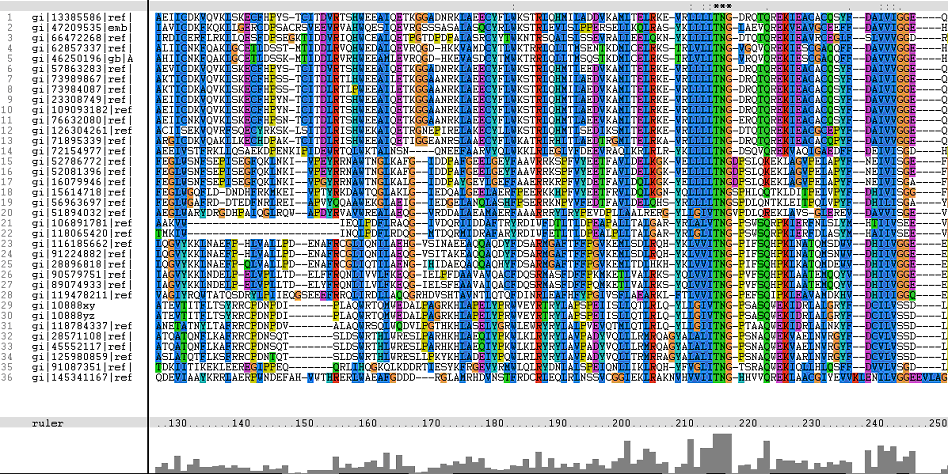
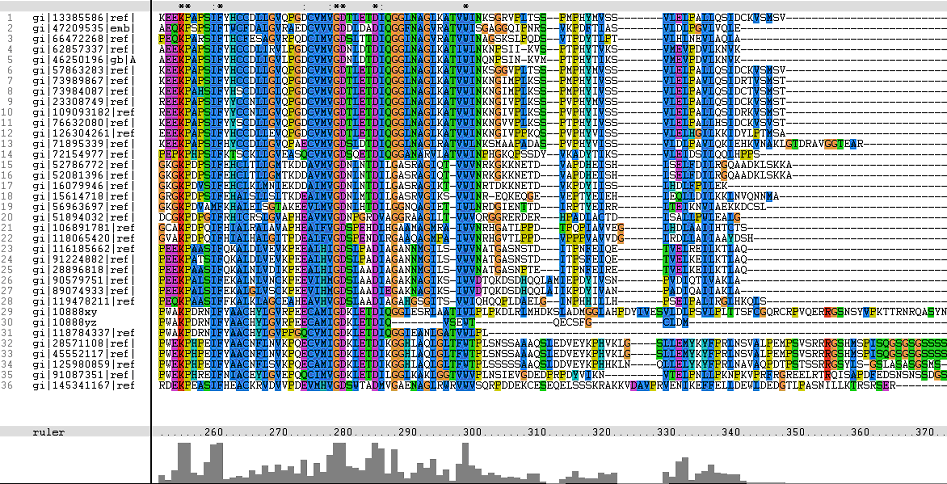
=='''Multiple Sequence Alignments (msa)'''== | =='''Multiple Sequence Alignments (msa)'''== | ||
| Line 54: | Line 41: | ||
'''The topmost sequence belongs to the query protein'''. | '''The topmost sequence belongs to the query protein'''. | ||
| Line 95: | Line 83: | ||
From these short conservation regions, the functions or even structure of the encoded proteins could have significance in its evolutionary pattern. | From these short conservation regions, the functions or even structure of the encoded proteins could have significance in its evolutionary pattern. | ||
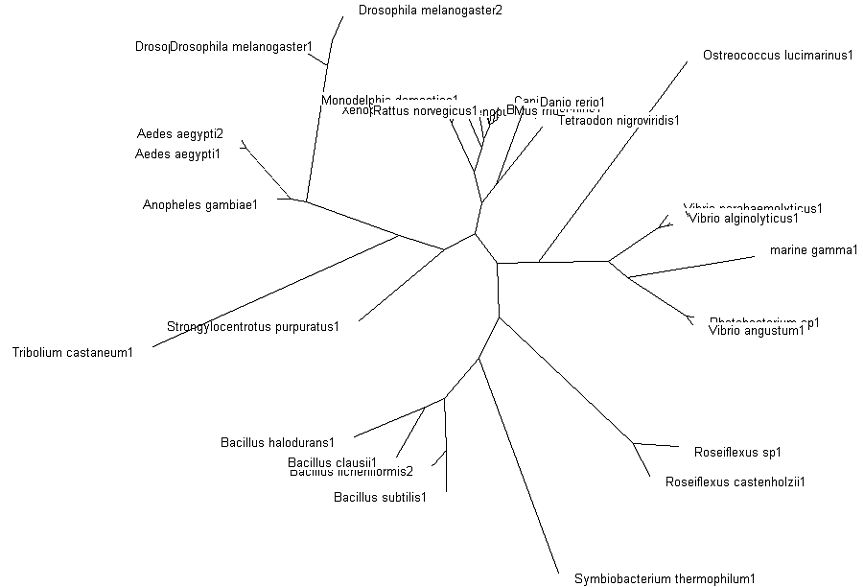
=='''Phylogenetic Map (Tree)'''== | =='''Phylogenetic Map (Tree)'''== | ||
| Line 119: | Line 105: | ||
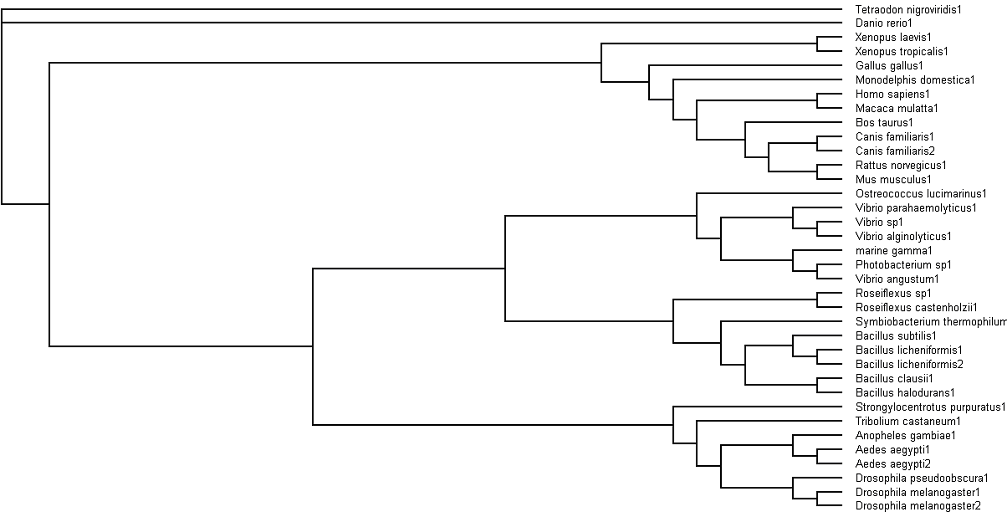
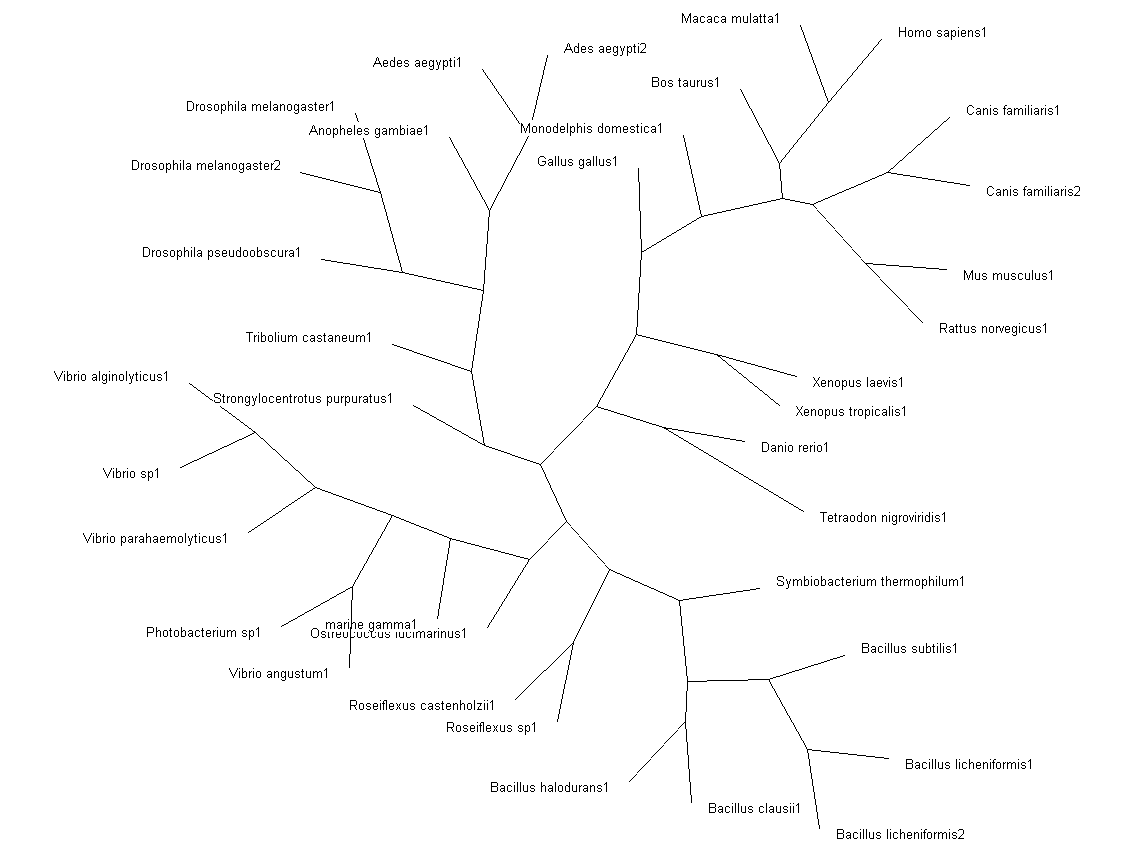
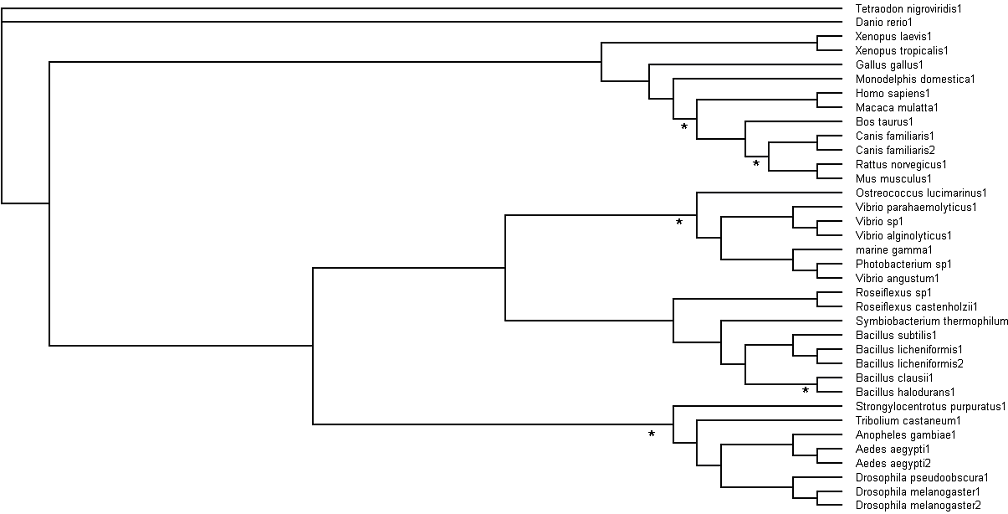
'''Fig. 3: Phylogenetic tree''' showing organisms with related protein sequence homology in Radial Tree view (top); and Rectangular Cladogram view (bottom) | '''Fig. 3: Phylogenetic tree''' showing organisms with related protein sequence homology in Radial Tree view (top); and Rectangular Cladogram view (bottom) | ||
[[Image:untitled.png|Strapboot Tree]] | |||
| Line 164: | Line 157: | ||
The bootstrap values (in percentage) obtained on each branch, signify branching confidence. Bootstrap values of 95% equate to full branching confidence; 75% value equates to 95% branching confidence; 60% value equates to much lowered branching confidence; while 50% value would render no branching confidence. | The bootstrap values (in percentage) obtained on each branch, signify branching confidence. Bootstrap values of 95% equate to full branching confidence; 75% value equates to 95% branching confidence; 60% value equates to much lowered branching confidence; while 50% value would render no branching confidence. | ||
---- | |||
Proceed to '''[http://compbio.chemistry.uq.edu.au/mediawiki/index.php/Function Function]''' | |||
---- | |||
Back to '''[http://compbio.chemistry.uq.edu.au/mediawiki/index.php/N-acetylneuraminic_acid_phosphatase Home Page]''' | |||
Latest revision as of 05:49, 21 May 2008
Sequence Homology
The following query protein sequence of N-acetylneuraminic Acid was obtained from GenBank: