ATP binding domain 4 Structures: Difference between revisions
Anharmustafa (talk | contribs) |
Anharmustafa (talk | contribs) |
||
| Line 39: | Line 39: | ||
== Structure Similarities == | == Structure Similarities == | ||
[[Image:Dali33.PNG|framed|none|'''Figure | [[Image:Dali33.PNG|framed|none|'''Figure 4'''. The list of structural similarities to 1RU8 obtained using DALI. Highlighted by the box is the most similar structure that has known function which are used in this investigation.]] | ||
''Z score'' , the statistical significance of the similarity between protein-of-interest and other neighbourhood proteins. The program optimizes a weighted sum of similarities of intramolecular distances. | ''Z score'' , the statistical significance of the similarity between protein-of-interest and other neighbourhood proteins. The program optimizes a weighted sum of similarities of intramolecular distances. | ||
Revision as of 12:57, 2 June 2009
General Properties
General information from PDB indicates that :
(a) 1RU8 is a putative n-type ATP pyrophosphatase isolated from Pyrococcus furiosus, expressed in Escherichia Coli.
(b) Is a member of clan PP-loop
(c) Resolution of 2.7 angstroms, with an r-value of 0.218.
(d) Ligand chemical component identified as TRS (2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL).
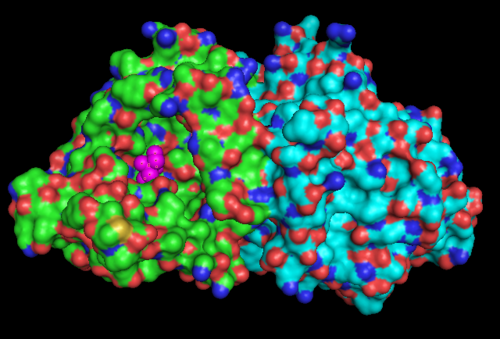
Surface Structure
Pymol Visualization
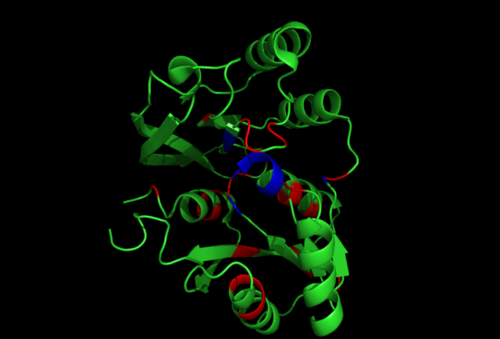
Secondary Structure and Location of PP-loop
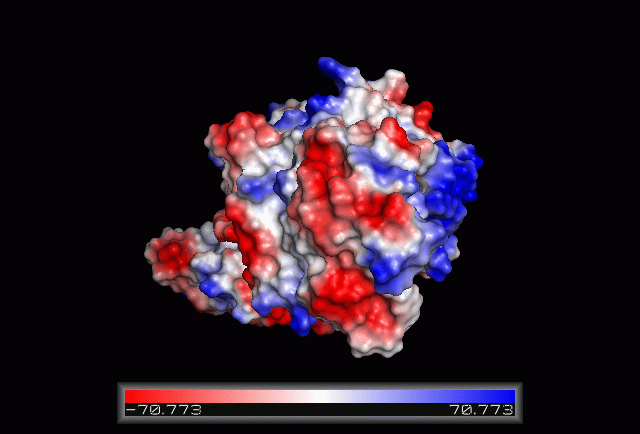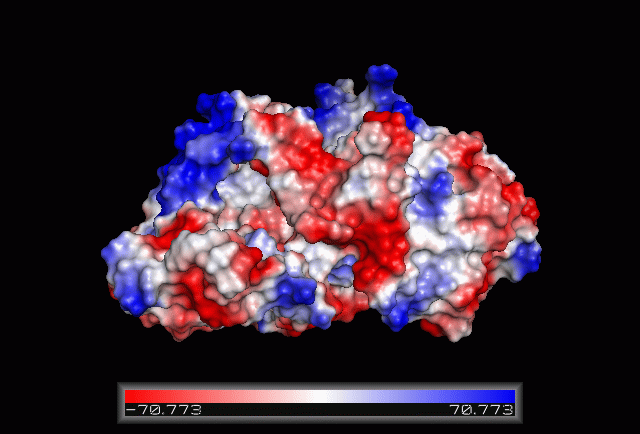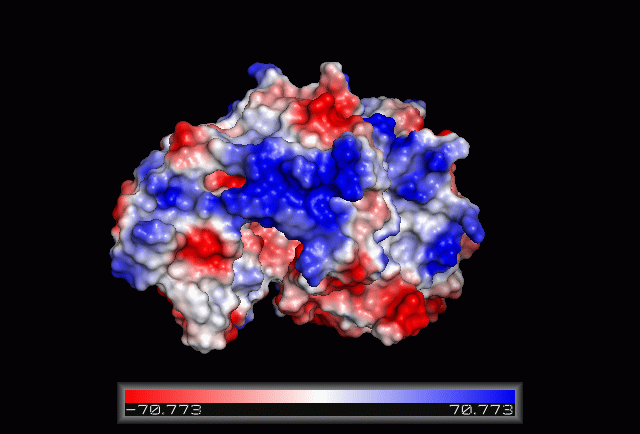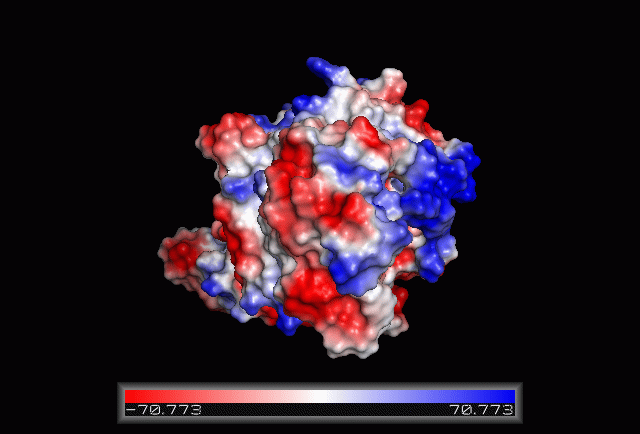
Electrostatic Surface Potential
Structure Similarities
Z score , the statistical significance of the similarity between protein-of-interest and other neighbourhood proteins. The program optimizes a weighted sum of similarities of intramolecular distances.
Root Mean Square Distance (RMSD), root-mean-square deviation of C-alpha atoms in the least-squares superimposition of the structurally equivalent C-alpha atoms. RMSD is not optimized and is only reported for information.
lali, the number of structurally equivalent residues.
nres, or the total number of amino acids in the hit protein.
%id, percentage of identical amino acids over structurally equivalent residues.
Structure Alignment
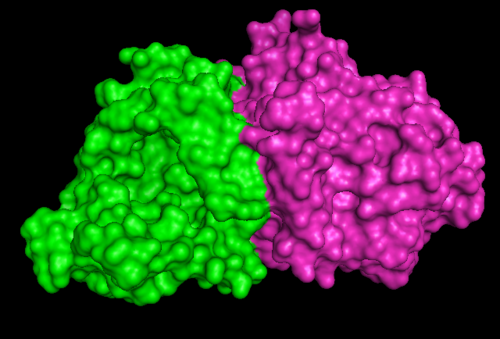
1RU8 and 2NZ2
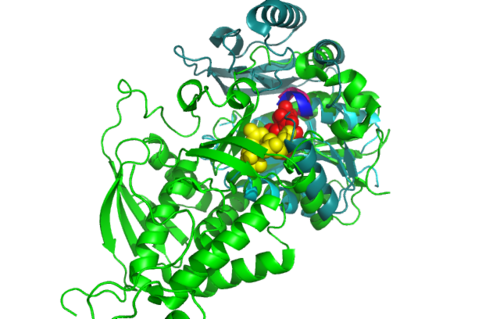
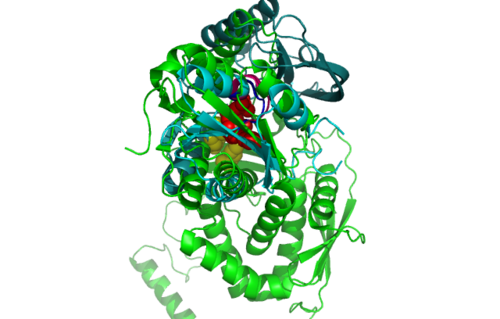
Highlighted in green is the 2NZ2 while in magenta is the 1RU8. Since 1RU8 has 2 domain, domain B is coloured darker in magenta for differentiation. There are 2 ligand indicated by the yellow (Citrulline) and red (ATP) spheres from 2NZ2. Highlighted also is the conserved region of PP-loop in pink and blue for 1RU8 and 2NZ2 respectively.
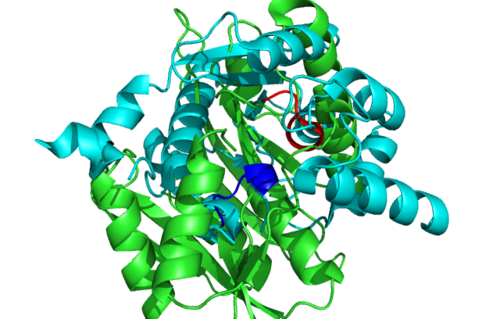
1RU8 and 3BL5
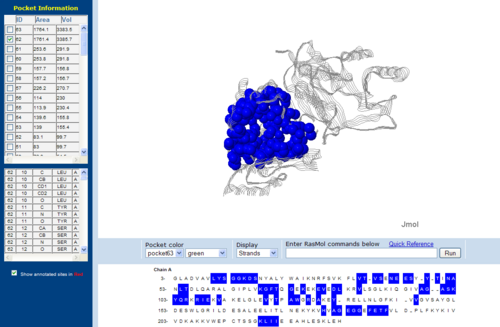
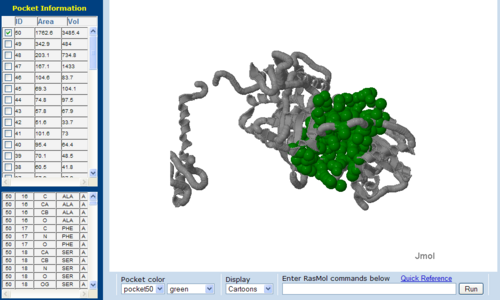
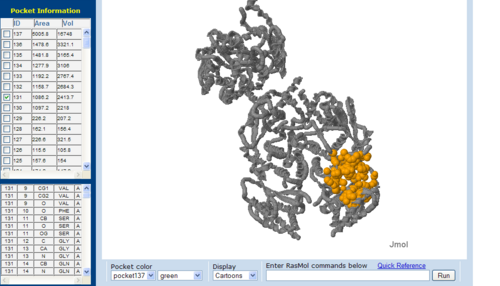
Surface Cleft and Topography
The volume and size of the cleft of between protein indicated similarity between 1RU8 and 2NZ2. While 3BL5 has less similarity when compared to both 1RU8 and 2NZ2