Structure ERp18: Difference between revisions
John O'Bryen (talk | contribs) No edit summary |
John O'Bryen (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
[[Image: Image_of_CC_residues.png]] | [[Image: Image_of_CC_residues.png]] | ||
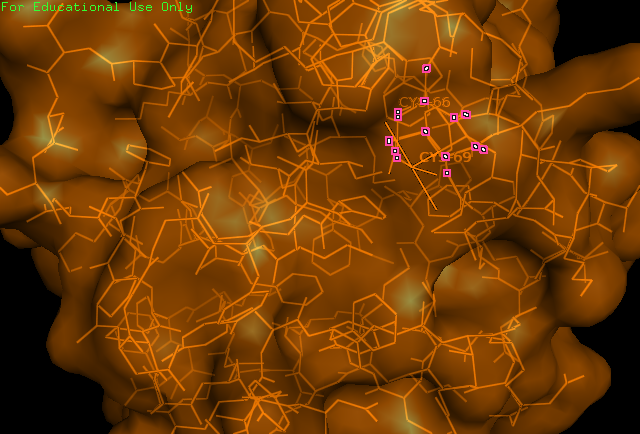
'''Figure 1: PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases. | '''Figure 1:''' PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases. | ||
| Line 7: | Line 7: | ||
[[Image: Image_of_ERp16_only_active_site_no_ions.png]] | [[Image: Image_of_ERp16_only_active_site_no_ions.png]] | ||
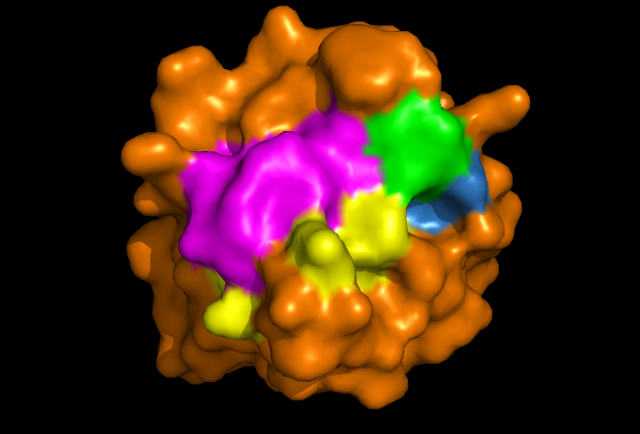
'''Figure 2: PyMOL image displaying conserved surface residues, which correspond to the location of the cysteine residues. These conserved surface residues are hypothesised to form the active site of ERp18. | '''Figure 2:''' PyMOL image displaying conserved surface residues, which correspond to the location of the cysteine residues. These conserved surface residues are hypothesised to form the active site of ERp18. | ||
| Line 13: | Line 13: | ||
[[Image: Image_of_ERp18_with_conserved_residues.png]] | [[Image: Image_of_ERp18_with_conserved_residues.png]] | ||
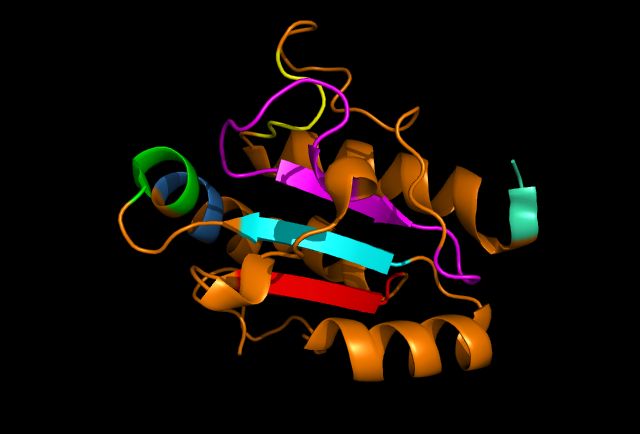
'''Figure 3: PyMOL image displaying the conserved sections of a ribbon diagram of ERp18. Notice the conservation of the three core beta sheets - probably vital for maintaining stability and correct folding of the protein. | '''Figure 3:''' PyMOL image displaying the conserved sections of a ribbon diagram of ERp18. Notice the conservation of the three core beta sheets - probably vital for maintaining stability and correct folding of the protein. | ||
| Line 71: | Line 71: | ||
50: 1ert-A 9.5 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; | 50: 1ert-A 9.5 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; | ||
'''Figure 4: The top 50 hits from a DALI search with 1sen-A. DALI lists proteins in order of their structural similarity. The protein names displayed are almost entirely thioredoxins (synonymous with thiol:disulfide interchange proteins), with the exception of 'putative exported cytochrome c biogenesis-related protein' for 2 and 3, and UBX domain-containing protein 7 for 41. However, hits 2 and 3 contain the CXXC motif and are therefore likely to be members of the thioredoxin family. The motif was not found in hit 41. | '''Figure 4:''' The top 50 hits from a DALI search with 1sen-A. DALI lists proteins in order of their structural similarity. The protein names displayed are almost entirely thioredoxins (synonymous with thiol:disulfide interchange proteins), with the exception of 'putative exported cytochrome c biogenesis-related protein' for 2 and 3, and UBX domain-containing protein 7 for 41. However, hits 2 and 3 contain the CXXC motif and are therefore likely to be members of the thioredoxin family. The motif was not found in hit 41. | ||
Revision as of 08:38, 7 June 2009
Figure 1: PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases.
Figure 2: PyMOL image displaying conserved surface residues, which correspond to the location of the cysteine residues. These conserved surface residues are hypothesised to form the active site of ERp18.
Figure 3: PyMOL image displaying the conserved sections of a ribbon diagram of ERp18. Notice the conservation of the three core beta sheets - probably vital for maintaining stability and correct folding of the protein.
Dali Summary
No: Chain Z rmsd lali nres %id PDB Description 1: 1sen-A 31.6 0.0 134 134 100 PDB MOLECULE: THIOREDOXIN-LIKE PROTEIN P19; 2: 3f9u-A 13.0 2.2 114 145 12 PDB MOLECULE: PUTATIVE EXPORTED CYTOCHROME C BIOGENESIS- 3: 3f9u-B 12.3 2.2 114 148 12 PDB MOLECULE: PUTATIVE EXPORTED CYTOCHROME C BIOGENESIS- 4: 2fwe-A 11.7 2.4 107 122 21 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 5: 2fwf-A 11.6 2.4 107 123 21 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 6: 1se1-C 11.1 2.1 103 231 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 7: 1vrs-F 11.1 2.1 103 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 8: 1vrs-D 11.1 2.2 102 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 9: 1se1-A 11.1 2.2 102 239 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 10: 1ep7-B 10.9 2.4 102 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 11: 2fwg-A 10.9 2.7 106 122 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 12: 2fwh-A 10.9 2.2 102 117 23 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 13: 1uc7-A 10.8 2.2 104 124 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 14: 1ep8-B 10.8 2.5 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 15: 1ep8-A 10.7 2.5 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 16: 2ju5-A 10.6 2.2 105 144 17 PDB MOLECULE: THIOREDOXIN DISULFIDE ISOMERASE; 17: 1ep7-A 10.6 2.6 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 18: 1vrs-E 10.6 2.2 102 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 19: 1uc7-B 10.5 2.3 104 124 23 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 20: 3d22-A 10.4 2.8 108 129 13 PDB MOLECULE: THIOREDOXIN H-TYPE; 21: 3d21-A 10.4 2.5 101 111 16 PDB MOLECULE: THIOREDOXIN H-TYPE; 22: 3d21-B 10.3 2.5 101 111 16 PDB MOLECULE: THIOREDOXIN H-TYPE; 23: 3fk8-A 10.2 2.6 106 131 17 PDB MOLECULE: DISULPHIDE ISOMERASE; 24: 2vlv-B 10.2 2.3 99 113 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 25: 1se1-B 10.2 2.2 102 243 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 26: 2vlu-B 10.2 2.3 100 112 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 27: 2vm1-A 10.2 2.3 99 110 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 28: 2vlv-A 10.1 2.4 100 111 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 29: 2vm2-A 10.1 2.3 99 109 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 30: 2vlt-B 10.1 2.4 100 110 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 31: 2vlu-A 10.0 2.4 100 111 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 32: 2iwt-A 10.0 2.2 99 108 19 PDB MOLECULE: THIOREDOXIN H ISOFORM 2; 33: 2vm1-B 10.0 2.4 99 110 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 34: 1gh2-A 9.9 2.3 97 107 14 PDB MOLECULE: THIOREDOXIN-LIKE PROTEIN; 35: 2vm1-C 9.8 2.5 100 113 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 36: 2vlt-A 9.8 2.5 100 111 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 37: 2vm1-D 9.8 2.5 100 109 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 38: 2vm2-B 9.7 2.3 97 107 15 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 39: 1auc-A 9.7 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; 40: 1auc 9.7 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; 41: 2dlx-A 9.6 2.5 111 153 16 PDB MOLECULE: UBX DOMAIN-CONTAINING PROTEIN 7; 42: 2vm2-D 9.6 2.5 99 108 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 43: 2hsh-A 9.6 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; 44: 1aiu 9.6 2.3 96 105 18 PDB MOLECULE: THIOREDOXIN; 45: 1aiu-A 9.6 2.3 96 105 18 PDB MOLECULE: THIOREDOXIN; 46: 1txx-A 9.5 2.3 101 108 19 PDB MOLECULE: PROTEIN (THIOREDOXIN); 47: 1erv 9.5 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; 48: 2ifq-A 9.5 2.3 96 105 18 PDB MOLECULE: THIOREDOXIN; 49: 1erv-A 9.5 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN; 50: 1ert-A 9.5 2.4 96 105 17 PDB MOLECULE: THIOREDOXIN;
Figure 4: The top 50 hits from a DALI search with 1sen-A. DALI lists proteins in order of their structural similarity. The protein names displayed are almost entirely thioredoxins (synonymous with thiol:disulfide interchange proteins), with the exception of 'putative exported cytochrome c biogenesis-related protein' for 2 and 3, and UBX domain-containing protein 7 for 41. However, hits 2 and 3 contain the CXXC motif and are therefore likely to be members of the thioredoxin family. The motif was not found in hit 41.
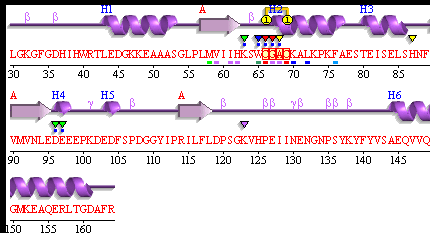
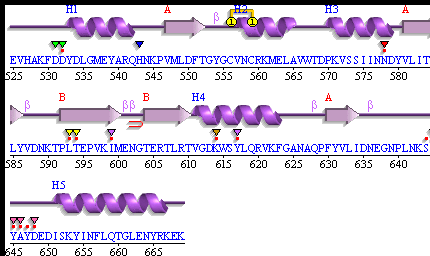
PDBsum of 1senA Secondary Structure
PDBsum of 3f9u Secondary Structure
SCOP Summary
Protein: Thioredoxin-like protein p19, TLP19 from Human (Homo sapiens) [TaxId: 9606] Lineage:
1. Root: scop
2. Class: Alpha and beta proteins (a/b) [51349]
Mainly parallel beta sheets (beta-alpha-beta units)
3. Fold: Thioredoxin fold [52832]
core: 3 layers, a/b/a; mixed beta-sheet of 4 strands, order 4312; strand 3 is antiparallel to the rest
4. Superfamily: Thioredoxin-like [52833]
5. Family: Thioltransferase [52834]
6. Protein: Thioredoxin-like protein p19, TLP19 [110604]
7. Species: Human (Homo sapiens) [TaxId: 9606] [110605]
PDB Entry Domains:
1. 1sen picpic
Structural genomics target
complexed with cl, pt
1. chain a [105465] picpiclink
PDBsum
PDB id: 1sen Name: Structural genomics, unknown function Title: Endoplasmic reticulum protein rp19 o95881
Structure: Thioredoxin-like protein p19. Chain: a. Synonym: endoplasmic reticulum protein erp19. Engineered: yes
Source: Homo sapiens. Human. Organism_taxid: 9606. Gene: tlp19. Expressed in: escherichia coli. Expression_system_taxid: 562. Other_details: the protein was cloned, expressed and purified by the secsg human protein production group (t.A. Dailey, m. Mayer) under the direction of h.A. Dailey.
UniProt: O95881 (TXD12_HUMAN) SAS Seq: 172 a.a. Struc: 135 a.a. Key: PfamB domain Secondary structure CATH domain
Enzyme class: E.C.1.8.4.2 [IntEnz] [ExPASy] [KEGG] [BRENDA]
Reaction: 2 glutathione + protein-disulfide = glutathione disulfide + protein- dithiol (see diagram below)
Resolution: 1.20Å
R-factor: 0.162
R-free: 0.183
Authors: Z.-J.Liu,L.Chen,W.Tempel,A.Shah,D.Lee,T.A.Dailey,M.R.Mayer, J.P.Rose,D.C.Richardson,J.S.Richardson,H.A.Dailey,B.-C.Wang Southeast Collaboratory For Structural Genomics (Secsg)
Key ref: z.-j.liu et al. Endoplasmic reticulum protein Rp19. To be Published, xsi:nil="true" />.
Date: 17-Feb-04
Release date: 13-Jul-04
Related entries: O95881 related db: targetdb