Structure ERp18: Difference between revisions
John O'Bryen (talk | contribs) No edit summary |
John O'Bryen (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
== Methods == | |||
== Results == | |||
[[Image: Image_of_CC_residues.png]] | [[Image: Image_of_CC_residues.png]] | ||
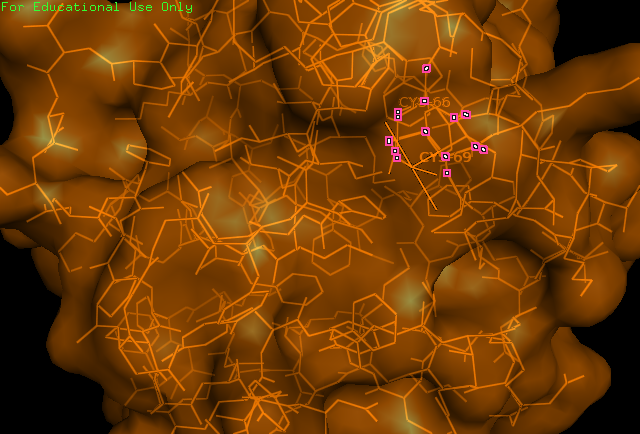
'''Figure 1:''' PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases. | '''Figure 1:''' PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases. | ||
[[Image: Image_of_ERp16_only_active_site_no_ions.png]] | [[Image: Image_of_ERp16_only_active_site_no_ions.png]] | ||
| Line 122: | Line 129: | ||
Related entries: O95881 related db: targetdb | Related entries: O95881 related db: targetdb | ||
== Discussion == | |||
Revision as of 11:25, 7 June 2009
Methods
Results
Figure 1: PyMOL image displaying the catalytic cysteine residues (66,69) of ERp18. These residues match the CXXC motif found in thiol–disulfide oxidoreductases.
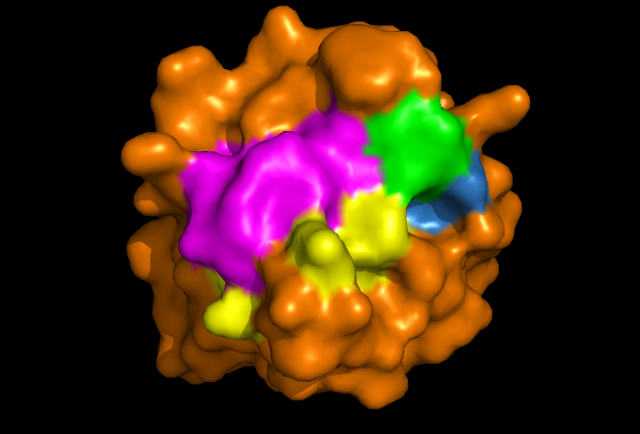
Figure 2: PyMOL image displaying conserved surface residues, which correspond to the location of the cysteine residues. These conserved surface residues are hypothesised to form the active site of ERp18.
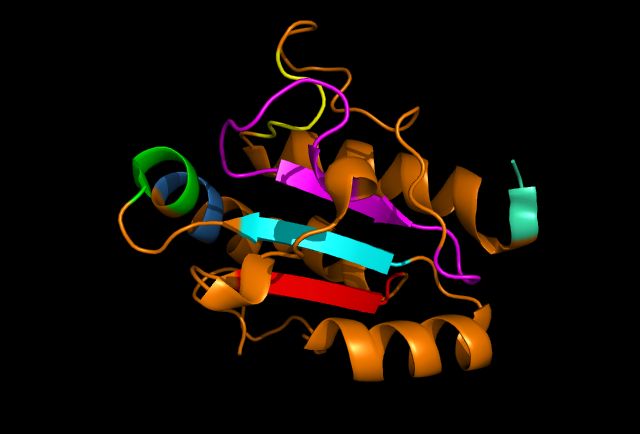
Figure 3: PyMOL image displaying the conserved sections of a ribbon diagram of ERp18. Notice the conservation of the three core beta sheets - probably vital for maintaining stability and correct folding of the protein.
Dali Summary
No: Chain Z rmsd lali nres %id PDB Description 1: 1sen-A 31.6 0.0 134 134 100 PDB MOLECULE: THIOREDOXIN-LIKE PROTEIN P19; 2: 3f9u-A 13.0 2.2 114 145 12 PDB MOLECULE: PUTATIVE EXPORTED CYTOCHROME C BIOGENESIS- 3: 3f9u-B 12.3 2.2 114 148 12 PDB MOLECULE: PUTATIVE EXPORTED CYTOCHROME C BIOGENESIS- 4: 2fwe-A 11.7 2.4 107 122 21 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 5: 2fwf-A 11.6 2.4 107 123 21 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 6: 1se1-C 11.1 2.1 103 231 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 7: 1vrs-F 11.1 2.1 103 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 8: 1vrs-D 11.1 2.2 102 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 9: 1se1-A 11.1 2.2 102 239 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 10: 1ep7-B 10.9 2.4 102 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 11: 2fwg-A 10.9 2.7 106 122 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 12: 2fwh-A 10.9 2.2 102 117 23 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 13: 1uc7-A 10.8 2.2 104 124 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 14: 1ep8-B 10.8 2.5 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 15: 1ep8-A 10.7 2.5 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 16: 2ju5-A 10.6 2.2 105 144 17 PDB MOLECULE: THIOREDOXIN DISULFIDE ISOMERASE; 17: 1ep7-A 10.6 2.6 103 112 13 PDB MOLECULE: THIOREDOXIN CH1, H-TYPE; 18: 1vrs-E 10.6 2.2 102 118 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 19: 1uc7-B 10.5 2.3 104 124 23 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 20: 3d22-A 10.4 2.8 108 129 13 PDB MOLECULE: THIOREDOXIN H-TYPE; 21: 3d21-A 10.4 2.5 101 111 16 PDB MOLECULE: THIOREDOXIN H-TYPE; 22: 3d21-B 10.3 2.5 101 111 16 PDB MOLECULE: THIOREDOXIN H-TYPE; 23: 3fk8-A 10.2 2.6 106 131 17 PDB MOLECULE: DISULPHIDE ISOMERASE; 24: 2vlv-B 10.2 2.3 99 113 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 25: 1se1-B 10.2 2.2 102 243 22 PDB MOLECULE: THIOL:DISULFIDE INTERCHANGE PROTEIN DSBD; 26: 2vlu-B 10.2 2.3 100 112 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 27: 2vm1-A 10.2 2.3 99 110 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 28: 2vlv-A 10.1 2.4 100 111 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.; 29: 2vm2-A 10.1 2.3 99 109 16 PDB MOLECULE: THIOREDOXIN H ISOFORM 1.; 30: 2vlt-B 10.1 2.4 100 110 22 PDB MOLECULE: THIOREDOXIN H ISOFORM 2.;
Figure 4: The top thirty hits from a DALI search with 1sen-A. DALI lists proteins in order of their structural similarity. The protein names displayed are almost entirely thioredoxins (synonymous with thiol:disulfide interchange proteins), with the exception of 'putative exported cytochrome c biogenesis-related protein' for 2 and 3. However, hits 2 and 3 contain the CXXC motif and are therefore likely to be members of the thioredoxin family.
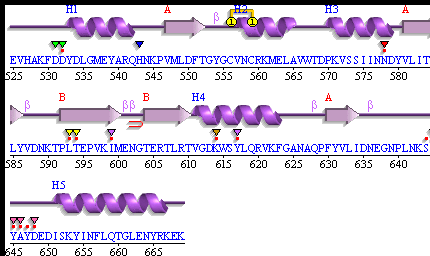
PDBsum of 1senA Secondary Structure
PDBsum of 3f9u Secondary Structure
SCOP Summary
Protein: Thioredoxin-like protein p19, TLP19 from Human (Homo sapiens) [TaxId: 9606] Lineage:
1. Root: scop
2. Class: Alpha and beta proteins (a/b) [51349]
Mainly parallel beta sheets (beta-alpha-beta units)
3. Fold: Thioredoxin fold [52832]
core: 3 layers, a/b/a; mixed beta-sheet of 4 strands, order 4312; strand 3 is antiparallel to the rest
4. Superfamily: Thioredoxin-like [52833]
5. Family: Thioltransferase [52834]
6. Protein: Thioredoxin-like protein p19, TLP19 [110604]
7. Species: Human (Homo sapiens) [TaxId: 9606] [110605]
PDB Entry Domains:
1. 1sen picpic
Structural genomics target
complexed with cl, pt
1. chain a [105465] picpiclink
PDBsum
PDB id: 1sen Name: Structural genomics, unknown function Title: Endoplasmic reticulum protein rp19 o95881
Structure: Thioredoxin-like protein p19. Chain: a. Synonym: endoplasmic reticulum protein erp19. Engineered: yes
Source: Homo sapiens. Human. Organism_taxid: 9606. Gene: tlp19. Expressed in: escherichia coli. Expression_system_taxid: 562. Other_details: the protein was cloned, expressed and purified by the secsg human protein production group (t.A. Dailey, m. Mayer) under the direction of h.A. Dailey.
UniProt: O95881 (TXD12_HUMAN) SAS Seq: 172 a.a. Struc: 135 a.a. Key: PfamB domain Secondary structure CATH domain
Enzyme class: E.C.1.8.4.2 [IntEnz] [ExPASy] [KEGG] [BRENDA]
Reaction: 2 glutathione + protein-disulfide = glutathione disulfide + protein- dithiol (see diagram below)
Resolution: 1.20Å
R-factor: 0.162
R-free: 0.183
Authors: Z.-J.Liu,L.Chen,W.Tempel,A.Shah,D.Lee,T.A.Dailey,M.R.Mayer, J.P.Rose,D.C.Richardson,J.S.Richardson,H.A.Dailey,B.-C.Wang Southeast Collaboratory For Structural Genomics (Secsg)
Key ref: z.-j.liu et al. Endoplasmic reticulum protein Rp19. To be Published, xsi:nil="true" />.
Date: 17-Feb-04
Release date: 13-Jul-04
Related entries: O95881 related db: targetdb