Fascin Structure: Difference between revisions
| Line 25: | Line 25: | ||
Dali output showing first 33 structurally similar protein results. | Dali output showing first 33 structurally similar protein results. | ||
[[Image:Structural comparrison Fascin FGF.png]] | [[Image:Structural comparrison Fascin FGF.png]] | ||
Pymol | Pymol allignment of Fascin 1 and FGF | ||
== References == | == References == | ||
Revision as of 08:23, 10 June 2009
Fascin structure and domains
The human Fascin protein is a monomeric protein consisting of 8 repeats of the Fascin domain.
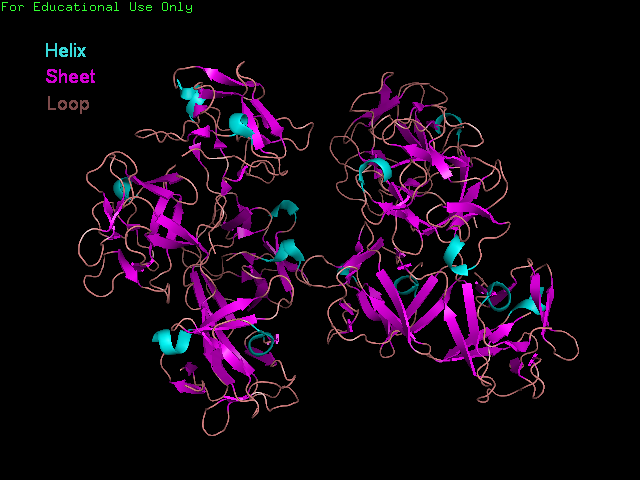
'Front' view showing secondary structure of Fascin.
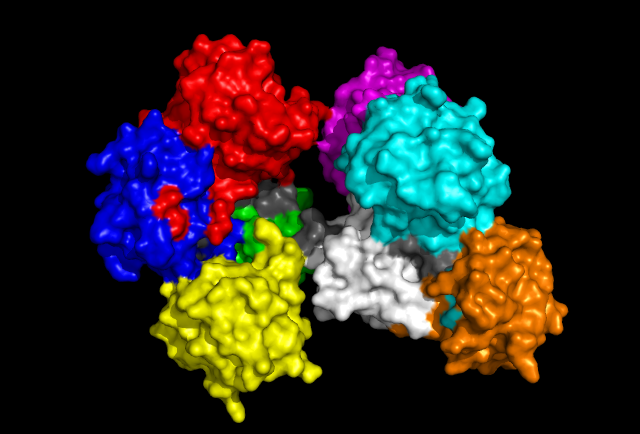
'Front' view of Fascin protein with each Fascin domain coloured. Grey colouration represents conserved residues found in sequence allignment.
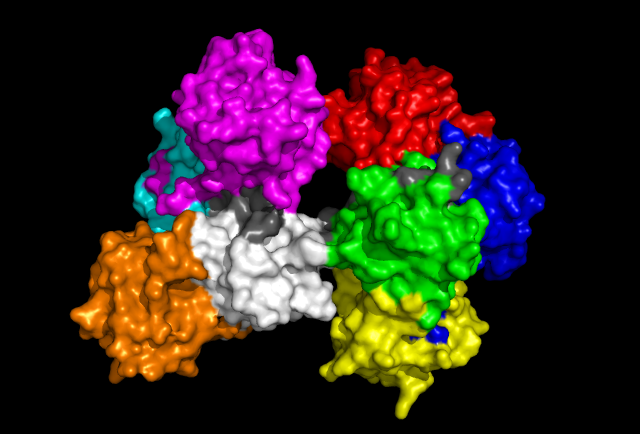
'Back' view of Fascin protein. Same colouration as above.
Structual allignments
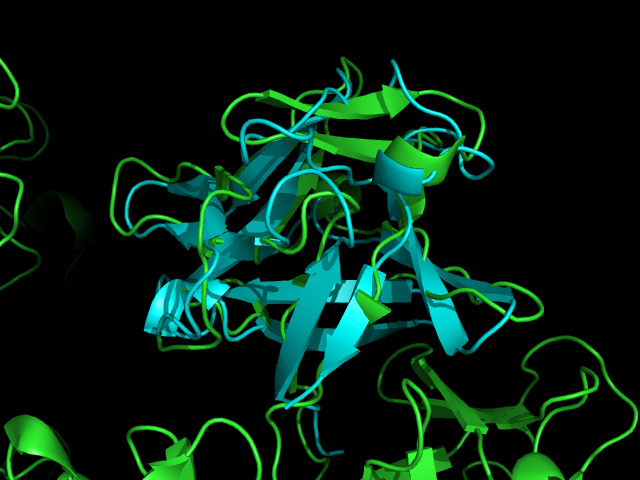
A Dali database search returned several hundred similar structures to Fascin 1. The first page of results can be seen below. Unfortunately none of the structurally similar molecules were actin binding so could not help with actin binding regions. The molecule 1ijt (Fibroblast Growth Factor - FGF) was alligned with Fascin to show structural similarity (even though they are functionally very different and only contain 9% sequence identity). Structurally FGF has much more Beta sheet secondarty structure, but as said before they are still quite similar
Dali output showing first 33 structurally similar protein results.
Pymol allignment of Fascin 1 and FGF
References
Joe Dundas, Zheng Ouyang, Jeffery Tseng, Andrew Binkowski, Yaron Turpaz, and Jie Liang 2006. CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated resiudes.Nucleic Acid Research, 34:W116-W118.
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA. PDB2PQR: an automated pipeline for the setup, execution, and analysis of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res, 32, W665-W667 (2004).