Evolution: Difference between revisions
JasonCheong (talk | contribs) |
JasonCheong (talk | contribs) No edit summary |
||
| Line 29: | Line 29: | ||
The bacteria sequence matches used for multiple sequence analysis and phylogenetic tree mapping were representative of the other baterial sequences not selected. These selected sequences had the highest bit scores and E-values. | The bacteria sequence matches used for multiple sequence analysis and phylogenetic tree mapping were representative of the other baterial sequences not selected. These selected sequences had the highest bit scores and E-values. | ||
=='''Multiple Sequence Alignments (msa)'''== | =='''Multiple Sequence Alignments (msa)'''== | ||
Revision as of 17:38, 10 June 2007
Query Sequences (2gfh)
The following query protein sequence of N-acetylneuraminic Acid was obtained from GenBank:
| mgsdkihhhh hhmglsrvra vffdldntli dtagasrrgm levikllqsk yhykeeaeii
cdkvqvklsk ecfhpystci tdvrtshwee aiqetkggad nrklaeecyf lwkstrlqhm iladdvkaml telrkevrll lltngdrqtq rekieacacq syfdaivigg eqkeekpaps ifyhccdllg vqpgdcvmvg dtletdiqgg lnaglkatvw inksgrvplt sspmphymvs svlelpallq sidckvsmsv |
Fig. 1: Amino Acid Sequence (260 aa) of N-acetylneuraminic Acid (2gfh) protein from Mus musculus (House Mouse)
BlastP similarity sequence search was carried out on fixed database found on DVD-rom. A total of 500 proteins were yielded.
Only a 38 proteins were selected for comparison as these had shown higher homology to the query sequence, in contrast with the remainder of the search results.
These proteins were chosen according to their bit scores and E-values. Two more outlier partial sequences contributing to poor overall alignment (huge deletion gaps) were subsequently removed. The remaining 36 sequences were used for the generation of the phylogenetic tree (and bootstrapped tree as well).
The bacteria sequence matches used for multiple sequence analysis and phylogenetic tree mapping were representative of the other baterial sequences not selected. These selected sequences had the highest bit scores and E-values.
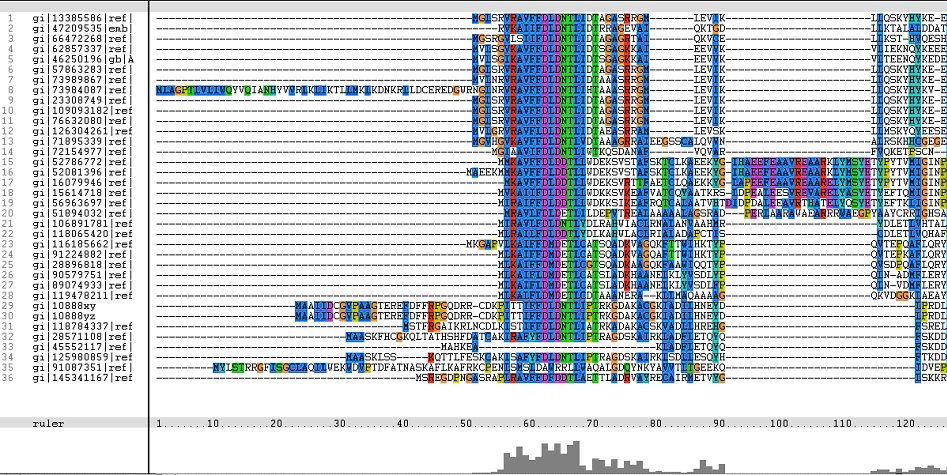
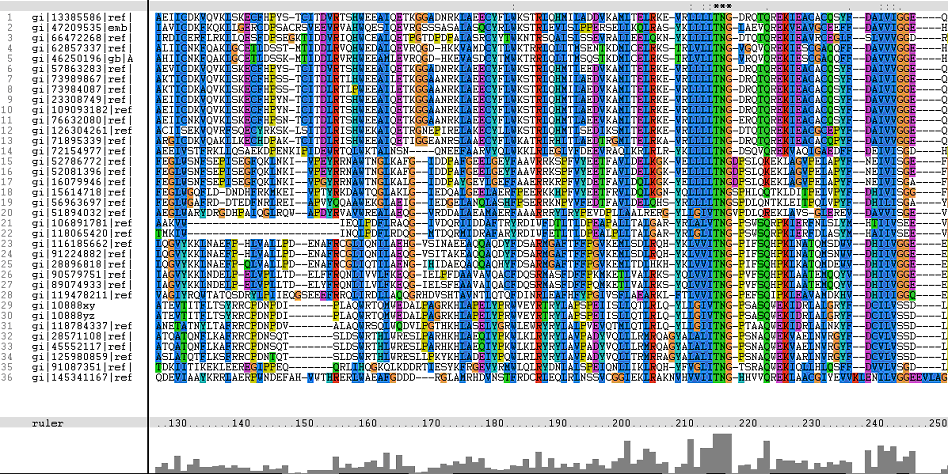
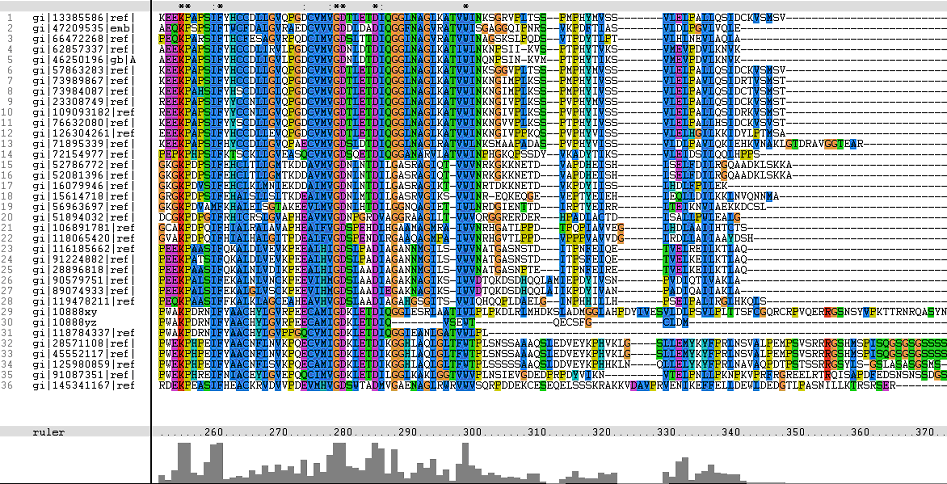
Multiple Sequence Alignments (msa)
Figure 1. Multiple Sequence Alignments of 2gfh amino acid sequences with that of other similar protein sequences in various organisms (A) Glycolysis pathway with substrates that are hydrolyze by HADs: glucose 6-phosphate, fructose 6-phosphate and dihydroxyacetone phosphate. (B) Pentose phostphate pathway with substrates that are hydrolyze by HADs: glucose-6-phosphate, fructose-6-phosphate, dihydroxyacetone phosphate, glyceraldehyde-3-phosphate, gluconate 6-phosphate and erythrose-4-phosphate.
| Catalytic activity | N-acylneuraminate 9-phosphate + H2O = N-acylneuraminate + phosphate |
| Cofactor | Magnesium (By similarity) |
| Enzyme regulation | Inhibited by vanadate and calcium (By similarity) |
| Pathway | Carbohydrate metabolism; aminosugar metabolism |
| Similarity | Belongs to the haloacid dehalogenase-like hydrolase superfamily. NANP family |