2gnx Results: Difference between revisions
From MDWiki
Jump to navigationJump to search
No edit summary |
No edit summary |
||
| Line 60: | Line 60: | ||
14: 3257-A 2hje-A 4.6 3.0 75 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2 sensor kinasePHOSPHATA | 14: 3257-A 2hje-A 4.6 3.0 75 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2 sensor kinasePHOSPHATA | ||
15: 3257-A 2uv0-E 4.5 3.5 93 159 9 0 0 12 S TRANSCRIPTION transcriptional activator protein lasr | 15: 3257-A 2uv0-E 4.5 3.5 93 159 9 0 0 12 S TRANSCRIPTION transcriptional activator protein lasr | ||
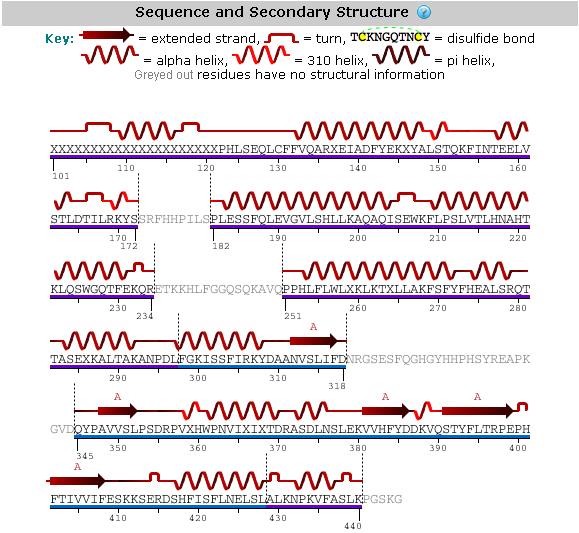
An analysis of the secondary structure of our protein (Figure 1) shows the secondary structural arrangement of different regions of our protein | |||
[[Image:Mel's_picture_of_secondary_str..jpg]] | [[Image:Mel's_picture_of_secondary_str..jpg]] | ||
Revision as of 06:04, 11 June 2007
Structural analysis
A Dali analysis (Table 1) of our protein was highly inconclusive and there were no significant structural matches to our hypothetical protein.
Table 1: A Dali analysis of the 2GNX protein
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3023-A 2gnx-A 42.9 0.0 280 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3023-A 2cmr-A 5.7 3.5 114 192 11 0 0 11 S IMMUNOGLOBULIN COMPLEX d5 (fab heavy chain) d5 (fab li 3: 3023-A 1j3w-A 5.7 3.2 99 134 12 0 0 9 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION giding protein-m 4: 3023-A 1jmr-A 5.5 3.0 94 246 9 0 0 12 S 5: 3023-A 1f5m-B 5.5 5.0 107 177 9 0 0 13 S SIGNALING PROTEIN gaf (saccharomyces cerevisiae) yeas 6: 3023-A 1vcs-A 5.0 4.7 82 102 9 0 0 8 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION vesicle transpor 7: 3023-A 1kt0-A 4.9 2.8 81 357 6 0 0 7 S ISOMERASE 51 kda fk506-binding protein (fkbp51) Mutant 8: 3023-A 1e2a-A 4.9 4.5 80 102 9 0 0 6 S TRANSFERASE enzyme iia (enzyme iii, lactose-specific i 9: 3023-A 2d2s-A 4.8 3.1 75 217 11 0 0 5 S ENDOCYTOSIS/EXOCYTOSIS exocyst complex component exo84 10: 3023-A 2oew-A 4.7 2.8 119 358 8 0 0 12 S PROTEIN TRANSPORT programmed cell death 6-interacting 11: 3023-A 1h3q-A 4.7 4.2 92 140 4 0 0 11 S TRANSPORT sedlin (sedl) (mus musculus) mouse S.B.Jan 12: 3023-A 2oev-A 4.5 36.5 151 697 7 0 0 14 S PROTEIN TRANSPORT programmed cell death 6-interacting 13: 3023-A 2cwy-A 4.5 2.4 82 92 20 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 14: 3023-A 2c5i-T 4.5 2.8 75 93 11 0 0 5 S PROTEIN TRANSPORT/COMPLEX t-snare affecting a late gol 15: 3023-A 3nul 4.4 3.4 93 130 5 0 0 11 S ACTIN-BINDING PROTEIN profilin i (arabidopsis thalian
A Dali analysis carried out separately with only the N-terminal domain (Table 2) of our protein also did not produce any significant structural matches.
Table 2: Dali analysis of N-terminal domain
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3256-A 2gnx-A 23.2 0.0 173 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3256-A 1e2a-A 7.5 4.5 80 102 9 0 0 6 S TRANSFERASE enzyme iia (enzyme iii, lactose-specific i 3: 3256-A 1kt0-A 7.4 2.8 81 357 6 0 0 7 S ISOMERASE 51 kda fk506-binding protein (fkbp51) Mutant 4: 3256-A 2d2s-A 7.3 3.1 75 217 11 0 0 5 S ENDOCYTOSIS/EXOCYTOSIS exocyst complex component exo84 5: 3256-A 1vcs-A 7.3 4.7 78 102 9 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION vesicle transpor 6: 3256-A 2cmr-A 6.9 3.2 104 192 11 0 0 9 S IMMUNOGLOBULIN COMPLEX d5 (fab heavy chain) d5 (fab li 7: 3256-A 2c5i-T 6.9 2.8 75 93 11 0 0 5 S PROTEIN TRANSPORT/COMPLEX t-snare affecting a late gol 8: 3256-A 2h7o-A 6.8 3.0 81 270 5 0 0 7 S SIGNALING PROTEIN protein kinase ypka fragment (protei 9: 3256-A 2h7v-C 6.6 4.2 76 269 13 0 0 5 S SIGNALING PROTEIN migration-inducing protein 5 (ras-re 10: 3256-A 2dnx-A 6.5 4.9 80 130 6 0 0 6 S TRANSPORT PROTEIN syntaxin-12 fragment (homo sapiens) 11: 3256-A 1hg5-A 6.5 3.2 85 263 9 0 0 6 S ENDOCYTOSIS clathrin assembly protein short form frag 12: 3256-A 1a17 6.4 2.5 71 159 3 0 0 5 S HYDROLASE serineTHREONINE PROTEIN PHOSPHATASE 5 fragme 13: 3256-A 2if4-A 6.3 2.5 82 258 7 0 0 7 S SIGNALING PROTEIN atfkbp42 fragment (twd1 (twisted dwa 14: 3256-A 1owa-A 6.2 3.3 76 156 12 0 0 6 S CYTOKINE spectrin alpha chain, erythrocyte fragment (e 15: 3256-A 2oew-A 6.1 2.8 119 358 8 0 0 12 S PROTEIN TRANSPORT programmed cell death 6-interacting
However, a Dali analysis (Table 3) carried out with the C-terminal domain of the protein produced one significant structural match, this being the GAF signalling protein, i.e the 4th result in the Dali analysis.
Table 3: Dali analysis of C-terminal domain
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3257-A 2gnx-A 24.3 0.0 118 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3257-A 1jmr-A 7.6 3.0 94 246 9 0 0 12 S 3: 3257-A 1j3w-A 7.5 2.9 91 134 13 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION giding protein-m 4: 3257-A 1f5m-B 6.8 2.9 95 177 9 0 0 10 S SIGNALING PROTEIN gaf (saccharomyces cerevisiae) yeas 5: 3257-A 1h3q-A 6.6 4.2 92 140 4 0 0 11 S TRANSPORT sedlin (sedl) (mus musculus) mouse S.B.Jan 6: 3257-A 3nul 6.3 3.4 93 130 5 0 0 11 S ACTIN-BINDING PROTEIN profilin i (arabidopsis thalian 7: 3257-A 1mc0-A 5.8 4.1 99 341 8 0 0 11 S HYDROLASE 3',5'-cyclic nucleotide phosphodiesterase 2a 8: 3257-A 2h28-A 5.4 2.8 75 106 8 0 0 10 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 9: 3257-A 2p7j-A 5.0 2.9 79 262 13 0 0 11 S TRANSCRIPTION putative sensory boxGGDEF FAMILY PROTEIN 10: 3257-A 2dmw-A 5.0 3.3 85 131 7 0 0 11 S MEMBRANE PROTEIN synaptobrevin-like 1 variant fragment 11: 3257-A 2avx-A 4.8 3.6 93 171 5 0 0 10 S TRANSCRIPTION regulatory protein sdia Mutant (escheri 12: 3257-A 2j3t-C 4.7 5.2 83 141 7 0 0 8 S PROTEIN TRANSPORT trafficking protein particle complex 13: 3257-A 2hj9-C 4.7 3.3 76 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2-binding periplasmic pr 14: 3257-A 2hje-A 4.6 3.0 75 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2 sensor kinasePHOSPHATA 15: 3257-A 2uv0-E 4.5 3.5 93 159 9 0 0 12 S TRANSCRIPTION transcriptional activator protein lasr
An analysis of the secondary structure of our protein (Figure 1) shows the secondary structural arrangement of different regions of our protein