Arylformamidase Results: Difference between revisions
Thomasparker (talk | contribs) No edit summary |
Thomasparker (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
most similar sequence - catalytic triad, structure with highlighted | |||
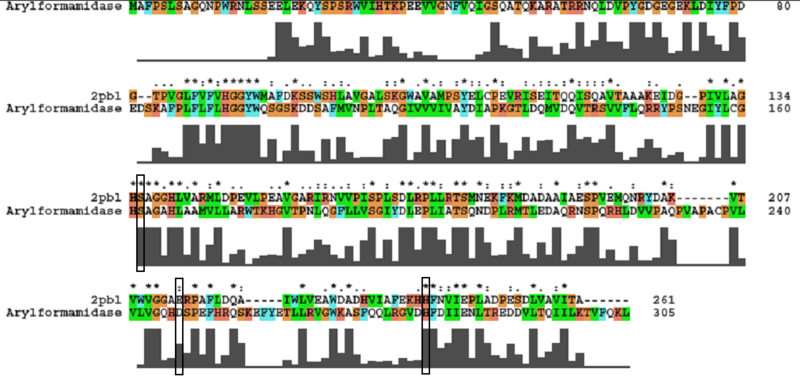
The most similar sequence with functional information available was that of an arylformamidase isolated from the liver of Mus musculus (see figure ...). A functional analysis of this protein has been performed identifying a catalytic triad using site-directed mutagenesis (Pabarcus et al. 2007). Conservation of this catalytic triad with 2pbl was assessed. Both residues S162 and H279 were found to be conserved in relatively conserved regions of the alignment. However, D247 had undergone a semi-conservative substitution. These residues correlated to S136, E214 and H241 of 2pbl which were subsequently located on the tertiary structure and determined to be sufficiently proximal to one another for catalysis (see figure...). | |||
[[Image:arylformamidase_alignment.png|centre|framed|'''Figure 3:''' ''Conservation of the catalytic triad between Arylformamidase and 2pbl.'']] | |||
most similar structure - catalytic triad, structure with highlighted | |||
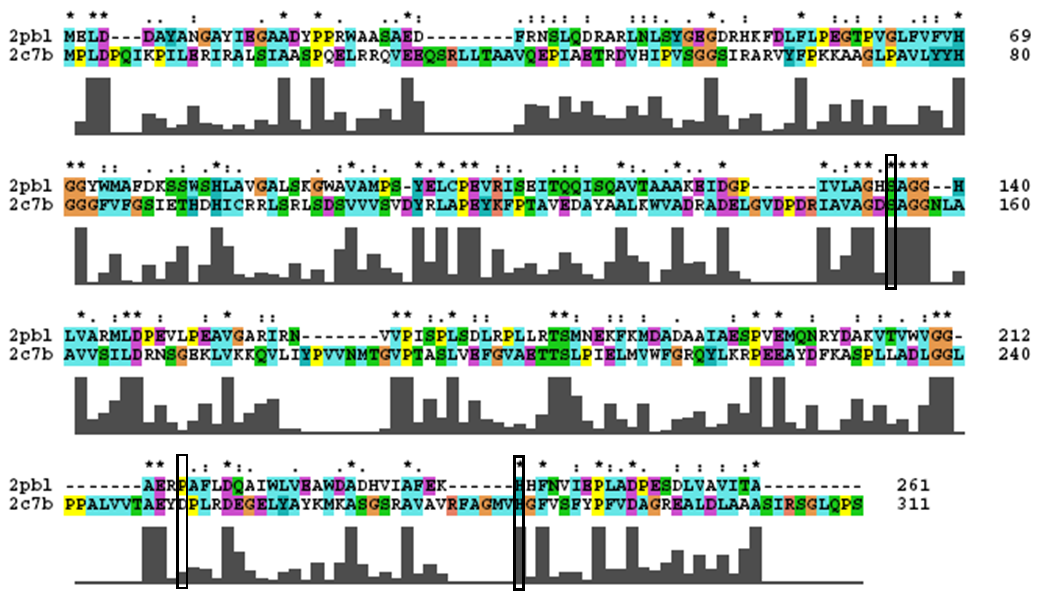
2pbl was found to share most structural similarity with a thermostable carboxylesterase from an uncultured archaeon (PDB ID: 2c7b; see figure ...). 2c7b shares a 16% sequence identity with 2pbl. From its structure, a catalytic triad has been identified (how?). To substantiate any functional similarity between 2pbl and 2c7b, conservation of the 2c7b catalytic triad was analysed (see figure ...). All three residues were found to be conserved, though H... and E... were found to match is less conserved regions. | |||
[[Image:2c7b_alignment.png|centre|framed|'''Figure 1:''' ''Conservation of the catalytic triad between 2cb7 and 2pbl.'']] | |||
presentation of tree and alignments | |||
[[Arylformamidase | Return to the main page...]] | [[Arylformamidase | Return to the main page...]] | ||
Revision as of 05:02, 3 June 2008
most similar sequence - catalytic triad, structure with highlighted
The most similar sequence with functional information available was that of an arylformamidase isolated from the liver of Mus musculus (see figure ...). A functional analysis of this protein has been performed identifying a catalytic triad using site-directed mutagenesis (Pabarcus et al. 2007). Conservation of this catalytic triad with 2pbl was assessed. Both residues S162 and H279 were found to be conserved in relatively conserved regions of the alignment. However, D247 had undergone a semi-conservative substitution. These residues correlated to S136, E214 and H241 of 2pbl which were subsequently located on the tertiary structure and determined to be sufficiently proximal to one another for catalysis (see figure...).
most similar structure - catalytic triad, structure with highlighted
2pbl was found to share most structural similarity with a thermostable carboxylesterase from an uncultured archaeon (PDB ID: 2c7b; see figure ...). 2c7b shares a 16% sequence identity with 2pbl. From its structure, a catalytic triad has been identified (how?). To substantiate any functional similarity between 2pbl and 2c7b, conservation of the 2c7b catalytic triad was analysed (see figure ...). All three residues were found to be conserved, though H... and E... were found to match is less conserved regions.
presentation of tree and alignments