DHRS1 Results: Difference between revisions
Mattlovell (talk | contribs) |
Mattlovell (talk | contribs) |
||
| Line 19: | Line 19: | ||
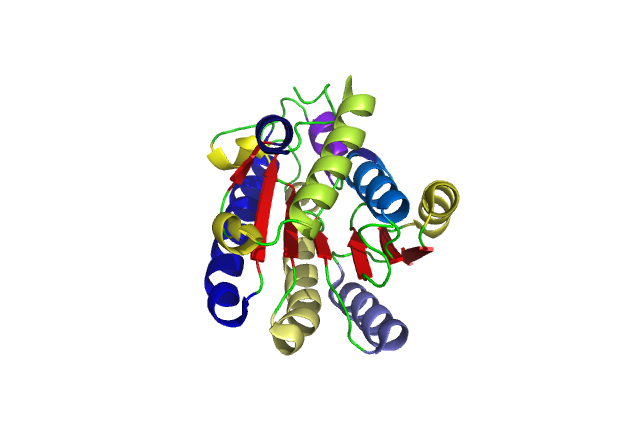
[[Image:pretty.png|framed|'''Figure 2'''<BR>Cartoon of a single DHRS1 unit with different structural features highlighted in different colours. |none]]<BR> | [[Image:pretty.png|framed|'''Figure 2'''<BR>Cartoon of a single DHRS1 unit with different structural features highlighted in different colours. |none]]<BR> | ||
A single monomeric unit of DHRS1 contains 10 α helices 1 central β-sheet region | A single monomeric unit of DHRS1 contains 10 α helices 1 central β-sheet region consisting of 7 strands. It is thought to exist biologically as a dimer. | ||
Dehydrogenase/reductase SDR family member 1 has highly conserved structure compared across the SDR family. | Dehydrogenase/reductase SDR family member 1 has highly conserved structure compared across the SDR family. | ||
Revision as of 05:41, 3 June 2008
Results/Evolution
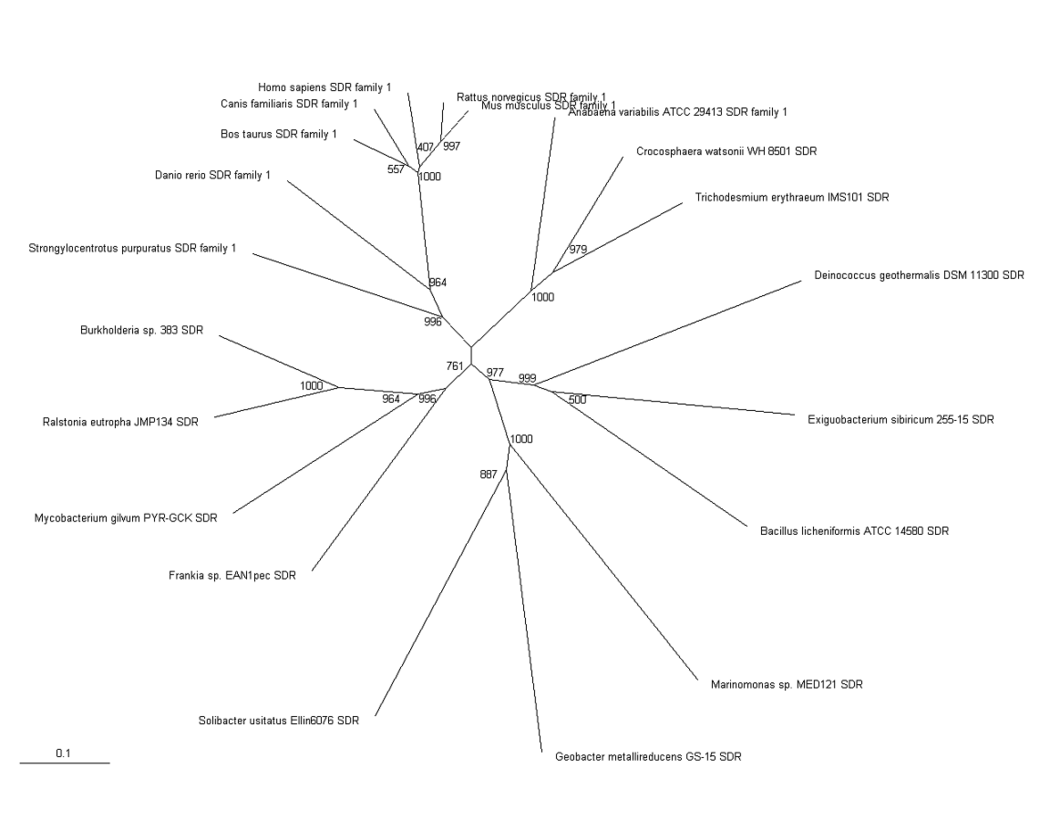
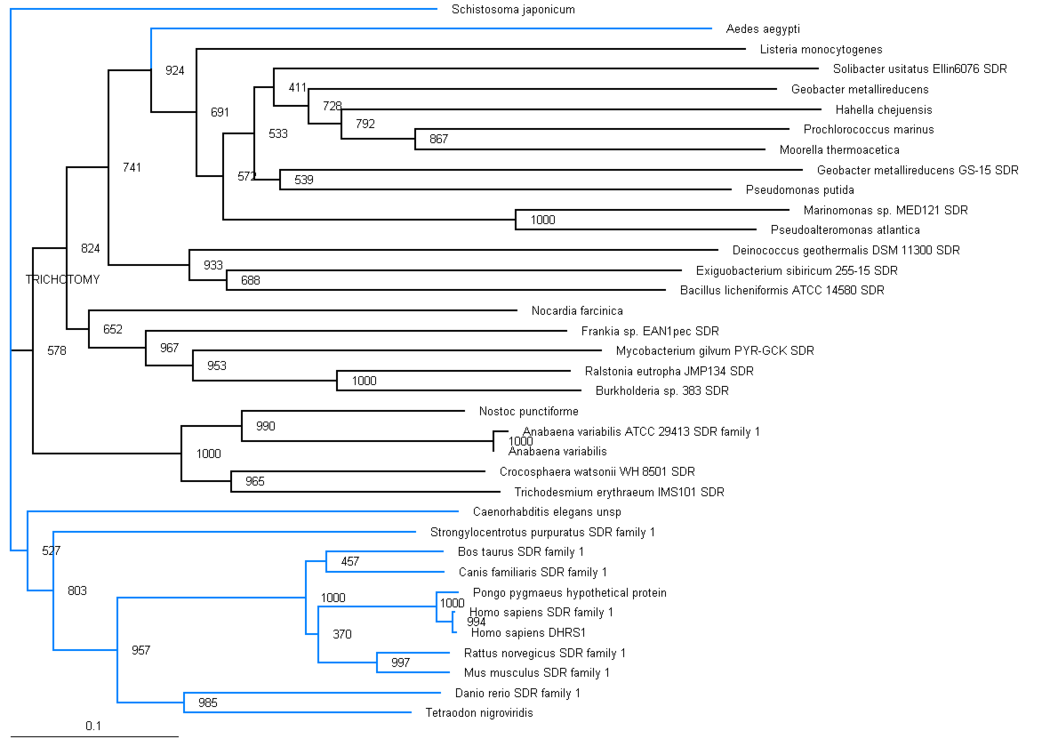
The phylogenic tree shows a small section of the family is contained in land based higher organisms (Bos taurus through to Mus Muluscus), a smaller section is found in sea based eukaryotes (Danio rerio and Tetraodon nigroviridis) and the majority of the family is round in bacteria. The majority of bacteria found to possess this family are proteobacteria, although there are also gram positive rod (eg. bacillus) and cyanobacteria (eg. anabena). This suggests that this particular family has been present for a long time and has evolved from bacteria, most likely proteobacteria.
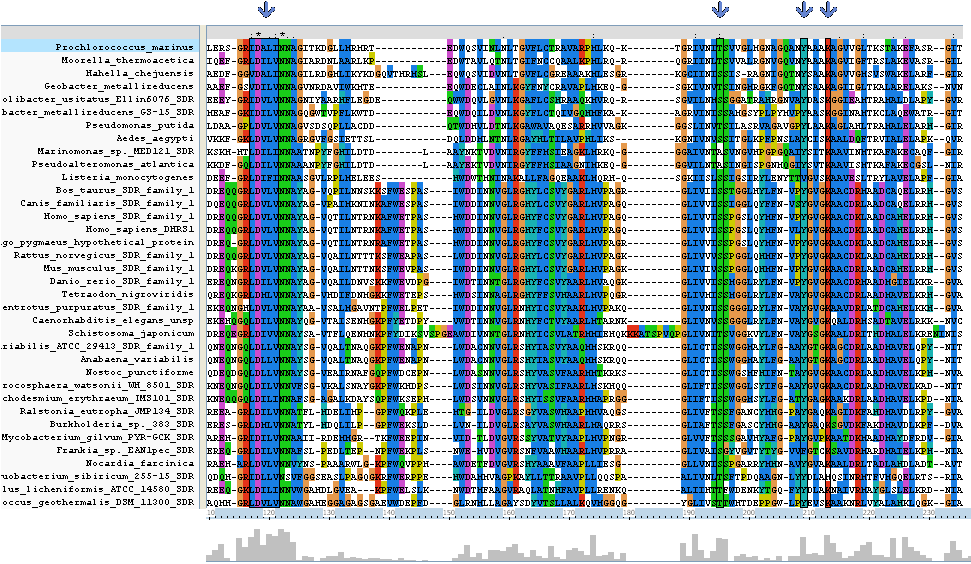
Clustal Alignment
Clustal alignment shows that Homo_sapiens_SDR_family_1 (an exact match for our target protein) with regards to protein sequence, contains all eight key residues (locations 117 - 121, 195,209 and 213). It also contains a sequence K-[A,S]-F-W-E-x-P-A-S at location 138 - 146 which is conserved only between land based animals.
Results/structure
A single monomeric unit of DHRS1 contains 10 α helices 1 central β-sheet region consisting of 7 strands. It is thought to exist biologically as a dimer.
Dehydrogenase/reductase SDR family member 1 has highly conserved structure compared across the SDR family.
SDR family is part of the Super Family; NAD(P)-binding Rossmann-fold domain proteins all of which have the Rossmann-fold domain, which is characterised a central β-sheet surrounded by α-helicies. This puts them in the Alpha and Beta proteins (α/β) class.
Some key residues are conserved across the entire family. Notable a Tyrosine that binds NAD(P). It is part of a larger motif [S-x(12)-Y-x(3)-K] that includes two other residues involved in binding NAD(P).
There is another highly conserved motif (LDVLD) involved in the initial folding of the protein. It forms part of the hydrophobic core most notably a strand of the β-sheet that is part of the Rossmann-fold.
Structure comparison
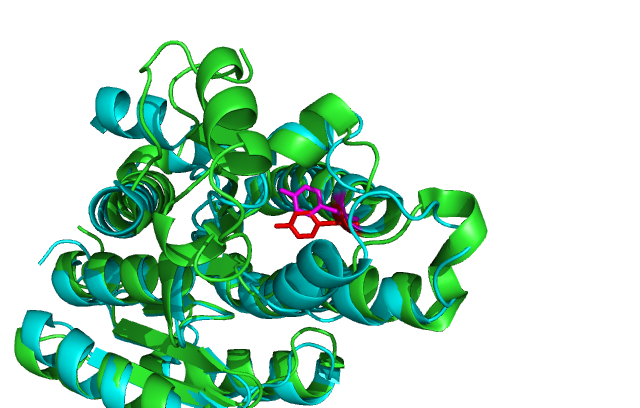
Comparison of DHRS1 to 3-OXOACYL-(ACYL-CARRIER-PROTEIN) REDUCTASE its closest structurally related protein, finds that whilst it shares only 27% of its sequence, it is structurally very similar (Fig 2). They share many structural features including the central β-sheet region and many α helices. Key conserved residues such as the tyrosine shown are also in very similar positions eluding to a similar function. Importantly the key catalytic triad (S-Y-K) is still in the same place with the tips of the residues only moved 2-3 Angstroms (fig 3) .
Table 1
PDB/chain identifiers and structural alignment statistics from DALI search
No: Chain Z rmsd lali nres %id Description 1: 2qq5-A 48.1 0.0 238 238 100 MOL: DEHYDROGENASE/REDUCTASE SDR1; 2: 2uvd-A 29.6 2.1 220 246 27 MOL: 3-OXOACYL-(ACYL-CARRIER-PROTEIN) REDUCTASE; 3: 1yde-F 29.3 2.1 216 256 29 MOL: RETINAL DEHYDROGENASE/REDUCTASE 3; 4: 1vl8-B 29.1 2.0 220 252 27 MOL: GLUCONATE 5-DEHYDROGENASE; 5: 2bgk-A 29.0 2.1 219 267 25 MOL: RHIZOME SECOISOLARICIRESINOL DEHYDROGENASE; 6: 2q2q-D 28.9 2.0 217 255 26 MOL: BETA-D-HYDROXYBUTYRATEDEHYDROGENASE; 7: 1rwb-F 28.8 2.2 221 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 8: 1rwb-A 28.8 2.3 222 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 9: 1gee-A 28.8 2.3 222 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 10: 1gco-A 28.8 2.3 222 261 24 MOL: GLUCOSE DEHYDROGENASE; 11: 2zat-A 28.7 2.1 221 251 23 MOL: DEHYDROGENASE/REDUCTASE SDR4; 12: 1gee-B 28.7 2.3 221 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 13: 1gco-E 28.7 2.3 221 261 24 MOL: GLUCOSE DEHYDROGENASE;