ATP binding domain 4 Structures
Useful info
Here is some useful paper. check it out Link to Paper: http://pfam.sanger.ac.uk/family?acc=PF00764 Dali: http://ekhidna.biocenter.helsinki.fi/dali_server/results/20090512-0051-3897fbdf5431ad2e808e66dde1070610/index.html#alignment-7 Interpro: http://www.ebi.ac.uk/interpro/DisplayIproEntry?ac=IPR002761 (align 1RU8 & i. 11-16, 2NZ2 & i. 11-15)
SCOP Work: http://scop.mrc-lmb.cam.ac.uk/scop/data/scop.b.d.cj.c.b.bb.html
General information from PDB indicates that :
(a) 1RU8 is a putative n-type ATP pyrophosphatase isolated from Pyrococcus furiosus, expressed in Escherichia Coli.
(b) Is a member of clan PP-loop
(c) Resolution of 2.7 angstroms, with an r-value of 0.218.
(d) Ligand chemical component identified as TRS (2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL).
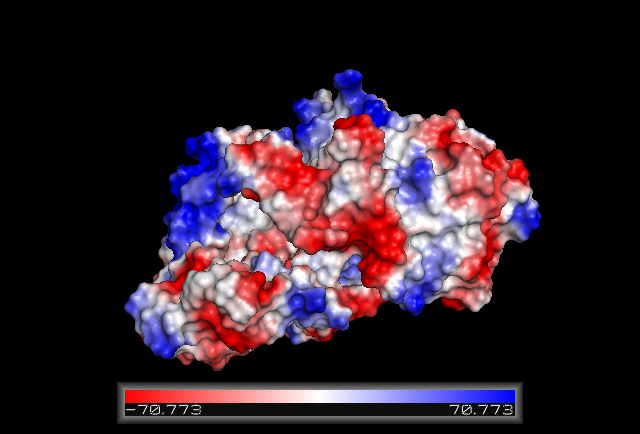
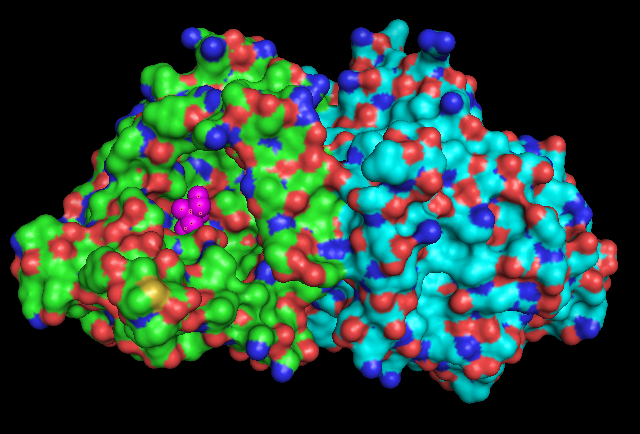
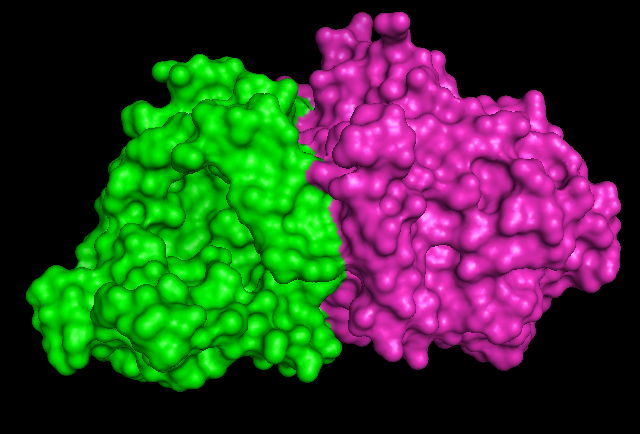
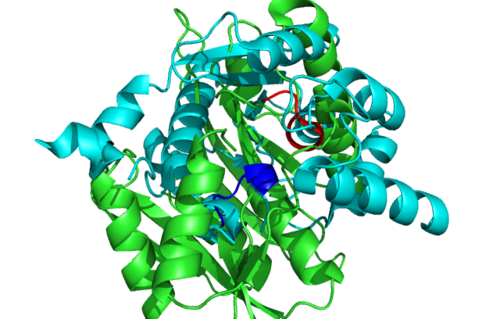
Surface Structure
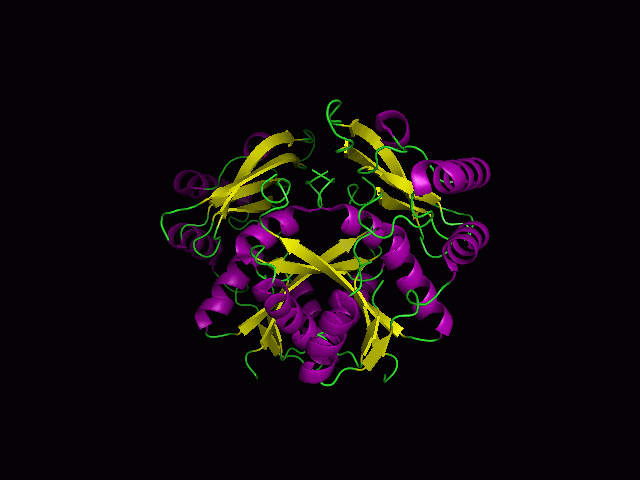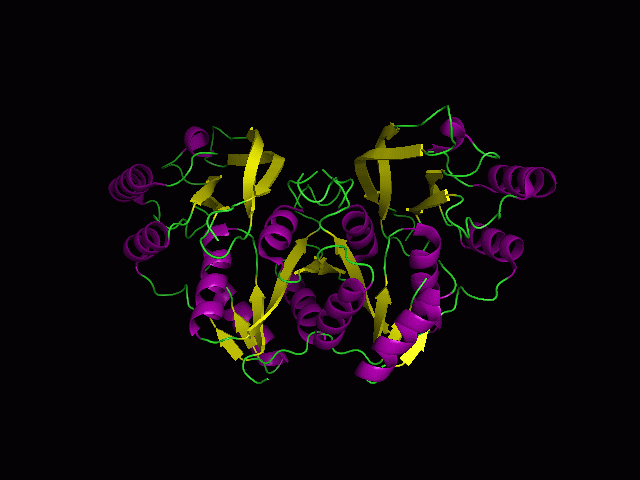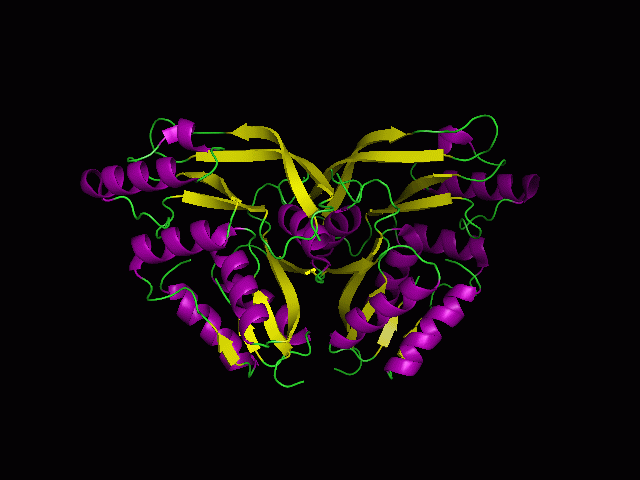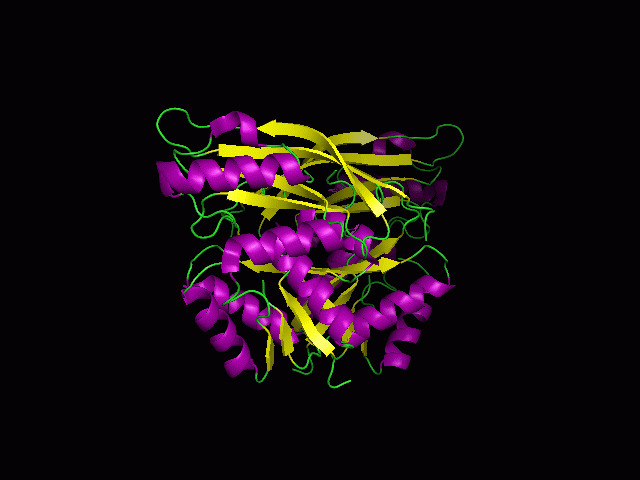
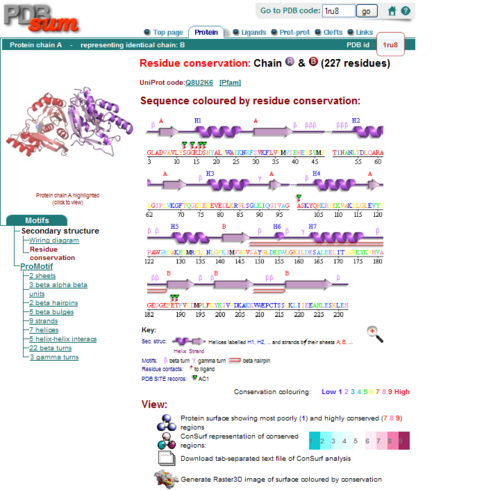
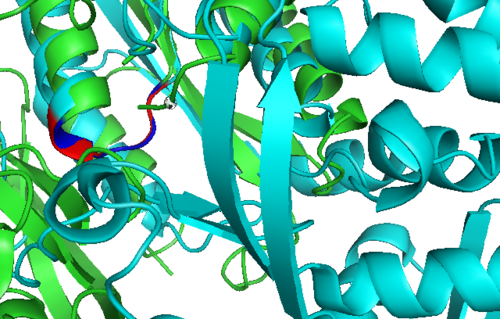
Secondary Structure and Location of P-loop (ATP binding site)
1RU8 is a dimer
Electrostatic Surface Potential
Alignment of 1RU8 and 2NZ2
Z score , the statistical significance of the similarity between protein-of-interest and other neighbourhood proteins. The program optimizes a weighted sum of similarities of intramolecular distances.
Root Mean Square Distance (RMSD), root-mean-square deviation of C-alpha atoms in the least-squares superimposition of the structurally equivalent C-alpha atoms. RMSD is not optimized and is only reported for information.
lali, the number of structurally equivalent residues.
nres, or the total number of amino acids in the hit protein.
%id, percentage of identical amino acids over structurally equivalent residues.
1RU8 and 2NZ2
1RU8 and 3BL5
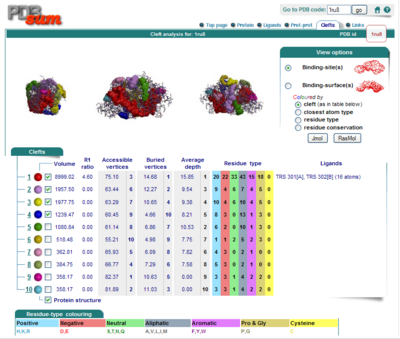
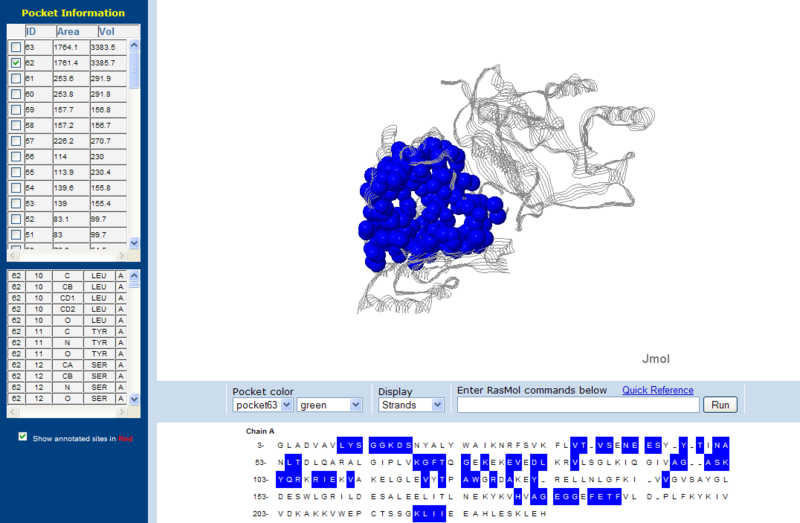
Surface Clefts
Surface Topography