2gnx Results
Structural analysis
A Dali analysis (Table 1) of our protein was highly inconclusive and there were no significant structural matches to our hypothetical protein.
Table 1: A Dali analysis of the 2GNX protein
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3023-A 2gnx-A 42.9 0.0 280 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3023-A 2cmr-A 5.7 3.5 114 192 11 0 0 11 S IMMUNOGLOBULIN COMPLEX d5 (fab heavy chain) d5 (fab li 3: 3023-A 1j3w-A 5.7 3.2 99 134 12 0 0 9 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION giding protein-m 4: 3023-A 1jmr-A 5.5 3.0 94 246 9 0 0 12 S 5: 3023-A 1f5m-B 5.5 5.0 107 177 9 0 0 13 S SIGNALING PROTEIN gaf (saccharomyces cerevisiae) yeas 6: 3023-A 1vcs-A 5.0 4.7 82 102 9 0 0 8 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION vesicle transpor 7: 3023-A 1kt0-A 4.9 2.8 81 357 6 0 0 7 S ISOMERASE 51 kda fk506-binding protein (fkbp51) Mutant 8: 3023-A 1e2a-A 4.9 4.5 80 102 9 0 0 6 S TRANSFERASE enzyme iia (enzyme iii, lactose-specific i 9: 3023-A 2d2s-A 4.8 3.1 75 217 11 0 0 5 S ENDOCYTOSIS/EXOCYTOSIS exocyst complex component exo84 10: 3023-A 2oew-A 4.7 2.8 119 358 8 0 0 12 S PROTEIN TRANSPORT programmed cell death 6-interacting 11: 3023-A 1h3q-A 4.7 4.2 92 140 4 0 0 11 S TRANSPORT sedlin (sedl) (mus musculus) mouse S.B.Jan 12: 3023-A 2oev-A 4.5 36.5 151 697 7 0 0 14 S PROTEIN TRANSPORT programmed cell death 6-interacting 13: 3023-A 2cwy-A 4.5 2.4 82 92 20 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 14: 3023-A 2c5i-T 4.5 2.8 75 93 11 0 0 5 S PROTEIN TRANSPORT/COMPLEX t-snare affecting a late gol 15: 3023-A 3nul 4.4 3.4 93 130 5 0 0 11 S ACTIN-BINDING PROTEIN profilin i (arabidopsis thalian
A Dali analysis carried out separately with only the N-terminal domain (Table 2) of our protein also did not produce any significant structural matches.
Table 2: Dali analysis of N-terminal domain
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3256-A 2gnx-A 23.2 0.0 173 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3256-A 1e2a-A 7.5 4.5 80 102 9 0 0 6 S TRANSFERASE enzyme iia (enzyme iii, lactose-specific i 3: 3256-A 1kt0-A 7.4 2.8 81 357 6 0 0 7 S ISOMERASE 51 kda fk506-binding protein (fkbp51) Mutant 4: 3256-A 2d2s-A 7.3 3.1 75 217 11 0 0 5 S ENDOCYTOSIS/EXOCYTOSIS exocyst complex component exo84 5: 3256-A 1vcs-A 7.3 4.7 78 102 9 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION vesicle transpor 6: 3256-A 2cmr-A 6.9 3.2 104 192 11 0 0 9 S IMMUNOGLOBULIN COMPLEX d5 (fab heavy chain) d5 (fab li 7: 3256-A 2c5i-T 6.9 2.8 75 93 11 0 0 5 S PROTEIN TRANSPORT/COMPLEX t-snare affecting a late gol 8: 3256-A 2h7o-A 6.8 3.0 81 270 5 0 0 7 S SIGNALING PROTEIN protein kinase ypka fragment (protei 9: 3256-A 2h7v-C 6.6 4.2 76 269 13 0 0 5 S SIGNALING PROTEIN migration-inducing protein 5 (ras-re 10: 3256-A 2dnx-A 6.5 4.9 80 130 6 0 0 6 S TRANSPORT PROTEIN syntaxin-12 fragment (homo sapiens) 11: 3256-A 1hg5-A 6.5 3.2 85 263 9 0 0 6 S ENDOCYTOSIS clathrin assembly protein short form frag 12: 3256-A 1a17 6.4 2.5 71 159 3 0 0 5 S HYDROLASE serineTHREONINE PROTEIN PHOSPHATASE 5 fragme 13: 3256-A 2if4-A 6.3 2.5 82 258 7 0 0 7 S SIGNALING PROTEIN atfkbp42 fragment (twd1 (twisted dwa 14: 3256-A 1owa-A 6.2 3.3 76 156 12 0 0 6 S CYTOKINE spectrin alpha chain, erythrocyte fragment (e 15: 3256-A 2oew-A 6.1 2.8 119 358 8 0 0 12 S PROTEIN TRANSPORT programmed cell death 6-interacting
However, a Dali analysis (Table 3) carried out with the C-terminal domain of the protein produced one significant structural match, this being the GAF signalling protein, i.e the 4th result in the Dali analysis.
Table 3: Dali analysis of C-terminal domain
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3257-A 2gnx-A 24.3 0.0 118 280 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 2: 3257-A 1jmr-A 7.6 3.0 94 246 9 0 0 12 S 3: 3257-A 1j3w-A 7.5 2.9 91 134 13 0 0 7 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION giding protein-m 4: 3257-A 1f5m-B 6.8 2.9 95 177 9 0 0 10 S SIGNALING PROTEIN gaf (saccharomyces cerevisiae) yeas 5: 3257-A 1h3q-A 6.6 4.2 92 140 4 0 0 11 S TRANSPORT sedlin (sedl) (mus musculus) mouse S.B.Jan 6: 3257-A 3nul 6.3 3.4 93 130 5 0 0 11 S ACTIN-BINDING PROTEIN profilin i (arabidopsis thalian 7: 3257-A 1mc0-A 5.8 4.1 99 341 8 0 0 11 S HYDROLASE 3',5'-cyclic nucleotide phosphodiesterase 2a 8: 3257-A 2h28-A 5.4 2.8 75 106 8 0 0 10 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 9: 3257-A 2p7j-A 5.0 2.9 79 262 13 0 0 11 S TRANSCRIPTION putative sensory boxGGDEF FAMILY PROTEIN 10: 3257-A 2dmw-A 5.0 3.3 85 131 7 0 0 11 S MEMBRANE PROTEIN synaptobrevin-like 1 variant fragment 11: 3257-A 2avx-A 4.8 3.6 93 171 5 0 0 10 S TRANSCRIPTION regulatory protein sdia Mutant (escheri 12: 3257-A 2j3t-C 4.7 5.2 83 141 7 0 0 8 S PROTEIN TRANSPORT trafficking protein particle complex 13: 3257-A 2hj9-C 4.7 3.3 76 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2-binding periplasmic pr 14: 3257-A 2hje-A 4.6 3.0 75 210 5 0 0 9 S SIGNALING PROTEIN autoinducer 2 sensor kinasePHOSPHATA 15: 3257-A 2uv0-E 4.5 3.5 93 159 9 0 0 12 S TRANSCRIPTION transcriptional activator protein lasr
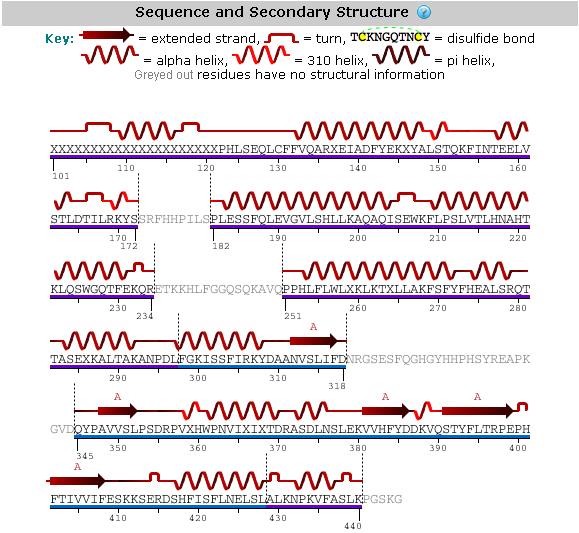
An analysis of the secondary structure of our protein from its amino acid sequence (Figure 1) shows the secondary structural arrangement of different regions of our protein
Figure 1: Secondary structure analysis of hypothetical protein LOC144557
Figure 2. Dotlet Analysis for 2GNX
The Dotlet analysis showed that there was no internally homologous repeats in 2GNX.
The comparison of 2GNX with the published cGMP bingding site