ATP binding domain 4 Structures: Difference between revisions
Anharmustafa (talk | contribs) No edit summary |
Anharmustafa (talk | contribs) No edit summary |
||
| Line 8: | Line 8: | ||
SCOP Work: http://scop.mrc-lmb.cam.ac.uk/scop/data/scop.b.d.cj.c.b.bb.html | SCOP Work: http://scop.mrc-lmb.cam.ac.uk/scop/data/scop.b.d.cj.c.b.bb.html | ||
General information from PDB indicates that : | |||
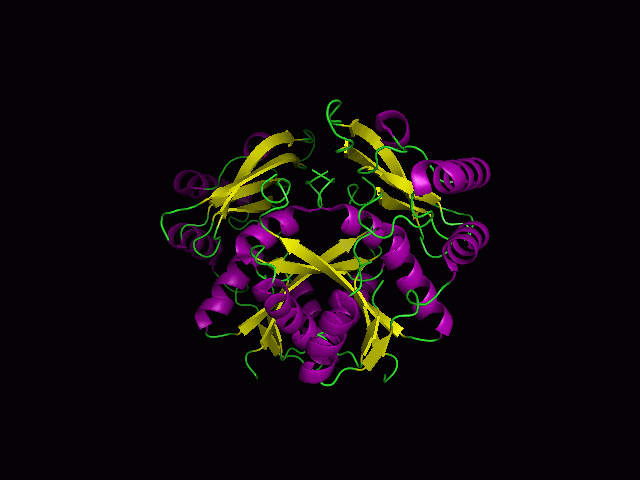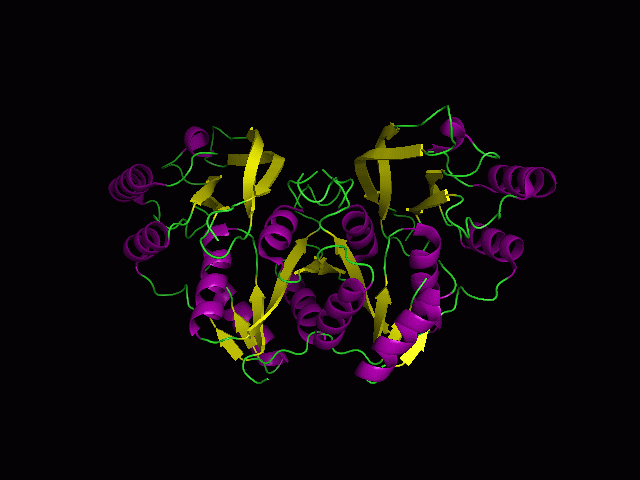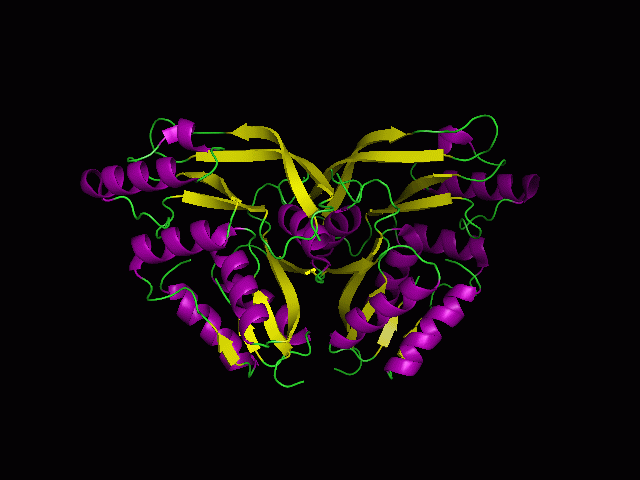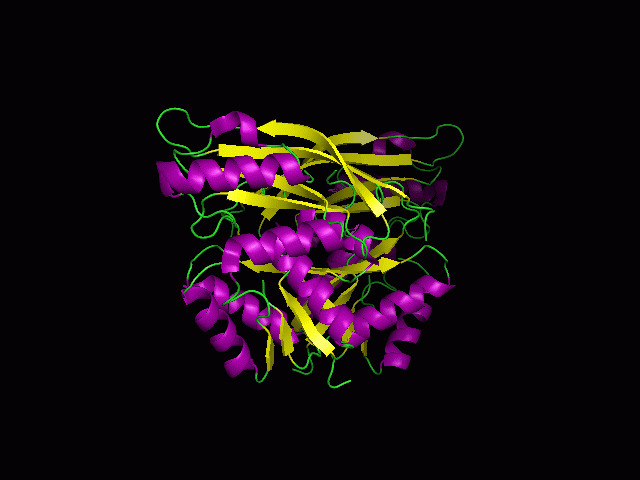
(a) 1RU8 is a putative n-type ATP pyrophosphatase isolated from Pyrococcus furiosus, expressed in Escherichia Coli. | |||
(b) Is a member of clan PP-loop | |||
(c) Resolution of 2.7 angstroms, with an r-value of 0.218. | |||
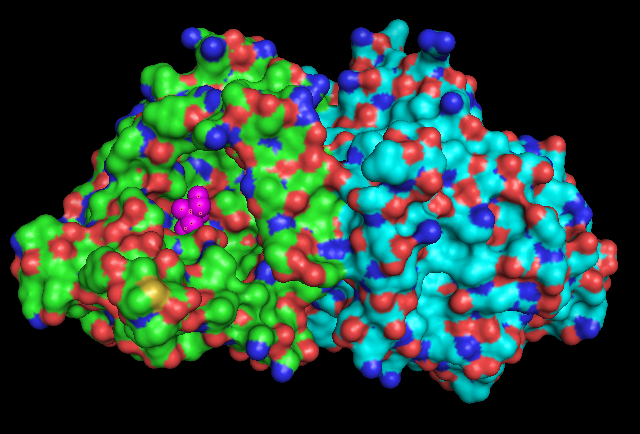
(d) Ligand chemical component identified as TRS (2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL). | |||
Revision as of 03:48, 30 May 2009
Useful info
Here is some useful paper. check it out Link to Paper: http://pfam.sanger.ac.uk/family?acc=PF00764 Dali: http://ekhidna.biocenter.helsinki.fi/dali_server/results/20090512-0051-3897fbdf5431ad2e808e66dde1070610/index.html#alignment-7 Interpro: http://www.ebi.ac.uk/interpro/DisplayIproEntry?ac=IPR002761 (align 1RU8 & i. 11-16, 2NZ2 & i. 11-15)
SCOP Work: http://scop.mrc-lmb.cam.ac.uk/scop/data/scop.b.d.cj.c.b.bb.html
General information from PDB indicates that :
(a) 1RU8 is a putative n-type ATP pyrophosphatase isolated from Pyrococcus furiosus, expressed in Escherichia Coli.
(b) Is a member of clan PP-loop
(c) Resolution of 2.7 angstroms, with an r-value of 0.218.
(d) Ligand chemical component identified as TRS (2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL).
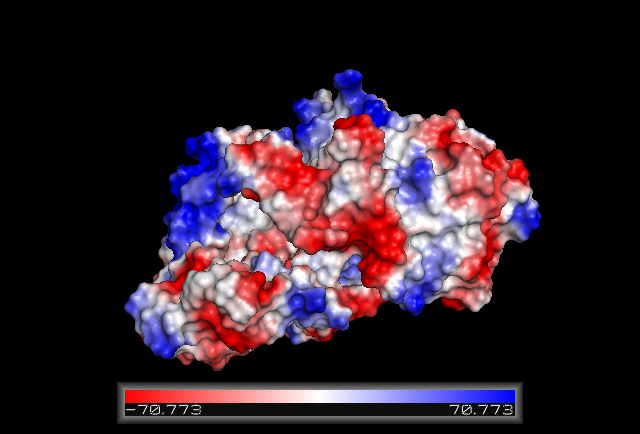
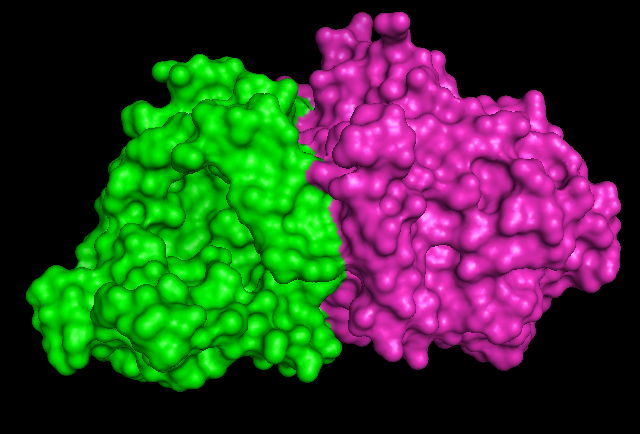
Surface Structure
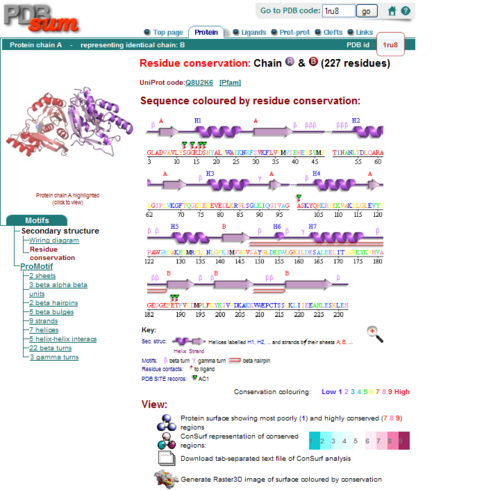
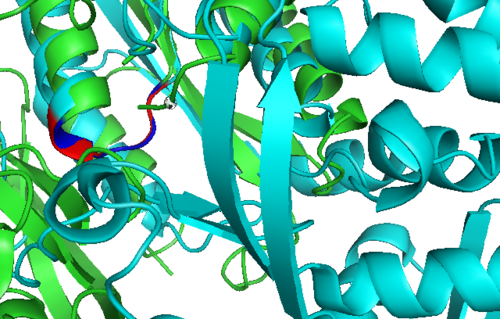
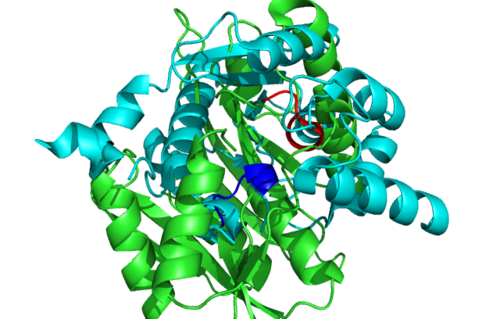
Secondary Structure and Location of P-loop (ATP binding site)
1RU8 is a dimer
Electrostatic Surface Potential
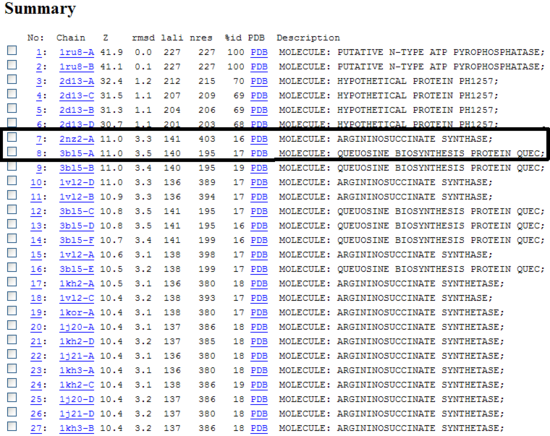
Alignment of 1RU8 and 2NZ2
1RU8 and 2NZ2
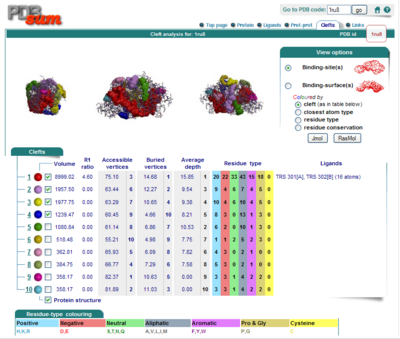
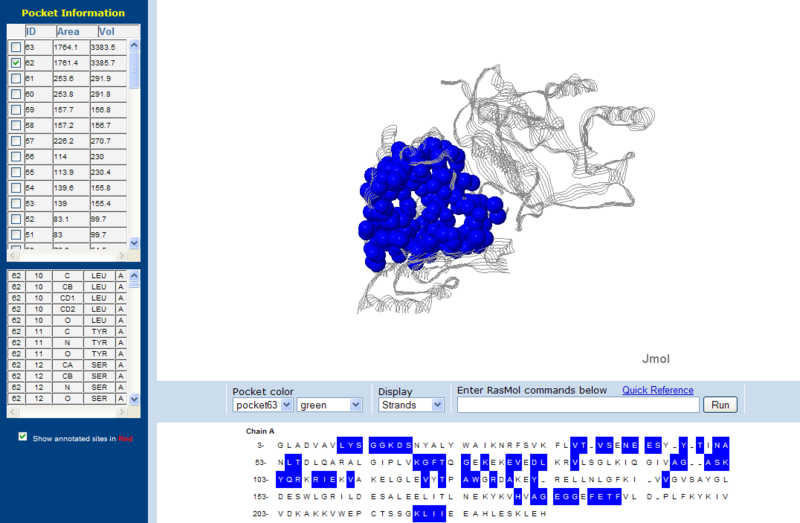
Surface Clefts
Surface Topography