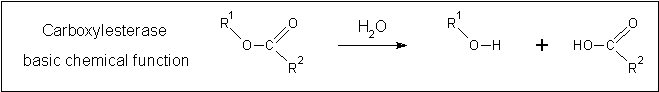Arylformamidase Function: Difference between revisions
Thomasparker (talk | contribs) No edit summary |
Thomasparker (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
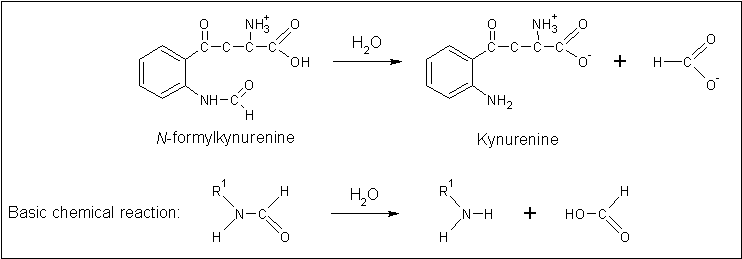
The putative annotation which was initially provided ''on the basis of...'' prompted a literature search using the term 'arylformamidase'. Pabarcus et al. (2007) have analysed the function of an arylformamidase present in the liver of ''Mus musculus''. Arylformamidase has... | The putative annotation which was initially provided ''on the basis of...'' prompted a literature search using the term 'arylformamidase'. Pabarcus et al. (2007) have analysed the function of an arylformamidase present in the liver of ''Mus musculus''. Arylformamidase has... | ||
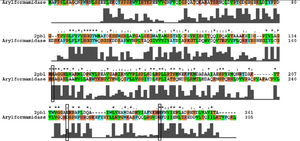
Ironically, from the BLAST results, the most similar sequence to 2pbl for which there was functional information available was the arylformamidase characterised by Pabarcus et al. 2007. Using site-directed mutagenesis, a catalytic triad was identified - S162, D247 and H279. To elucidate any functional similarity | Ironically, from the BLAST results, the most similar sequence to 2pbl for which there was functional information available was the arylformamidase characterised by Pabarcus et al. 2007. Using site-directed mutagenesis, a catalytic triad was identified - S162, D247 and H279. To elucidate any functional similarity between arylformamidase and 2pbl, conservation of the catalytic triad was assessed through a clustalW alignment. Note: the clustalW was found to differ from the alignment as provided as part of the blast results | ||
[[Image:arylformamidase_alignment.png|thumb|none|framed|'''Figure 3:''' ''Conservation of the catalytic triad between Arylformamidase and 2pbl.'']] | [[Image:arylformamidase_alignment.png|thumb|none|framed|'''Figure 3:''' ''Conservation of the catalytic triad between Arylformamidase and 2pbl.'']] | ||
Revision as of 08:35, 31 May 2008
The putative annotation which was initially provided on the basis of... prompted a literature search using the term 'arylformamidase'. Pabarcus et al. (2007) have analysed the function of an arylformamidase present in the liver of Mus musculus. Arylformamidase has...
Ironically, from the BLAST results, the most similar sequence to 2pbl for which there was functional information available was the arylformamidase characterised by Pabarcus et al. 2007. Using site-directed mutagenesis, a catalytic triad was identified - S162, D247 and H279. To elucidate any functional similarity between arylformamidase and 2pbl, conservation of the catalytic triad was assessed through a clustalW alignment. Note: the clustalW was found to differ from the alignment as provided as part of the blast results
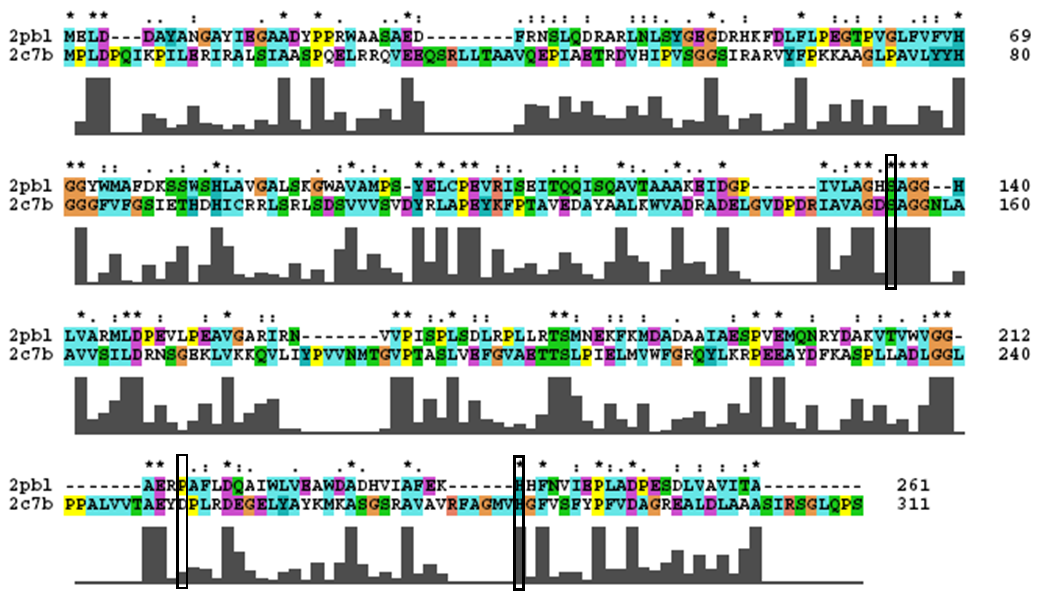
A review of the DALI output revealed many structures similar to 2pbl with little sequence identity. The most similar of these, 2cb7, is a thermophilic and thermostable carboxylesterase. The catalytic triad responsible for its activity has been identified, and in determining whether 2pbl shared similar functionality, conservation of the catalytic triad between the two proteins was assessed (see figure 1).