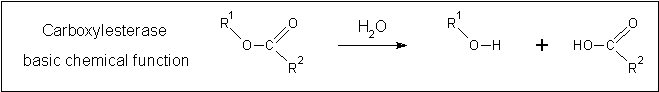Arylformamidase Function Slide 4: Difference between revisions
From MDWiki
Jump to navigationJump to search
Thomasparker (talk | contribs) No edit summary |
Thomasparker (talk | contribs) No edit summary |
||
| Line 13: | Line 13: | ||
== Evidence from Similar Structures == | == Evidence from Similar Structures == | ||
- Catalytic triad identified in paper - | - Catalytic triad identified in paper - Ser154, Asp251, and His281. | ||
- To assess functional similarity, conservation of the catalytic triad was analysed. | - To assess functional similarity, conservation of the catalytic triad was analysed. | ||
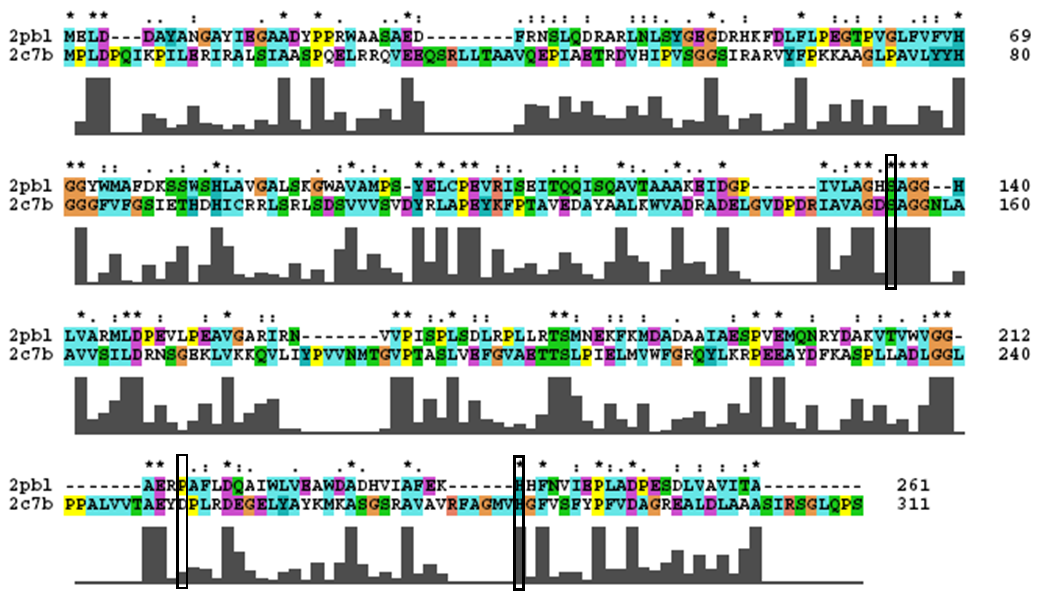
[[Image:2c7b_alignment.png|centre|framed|'''Conservation of the catalytic triad between 2cb7 and 2pbl.''']] | |||
- Catalytic triad conserved. | |||
- Note: difference in clustalW alignment from DALI sequence alignment. Unstable region of alignment? | |||
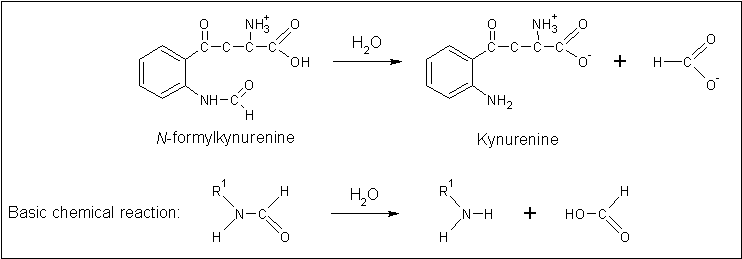
== Similarity in mechanism? == | == Similarity in mechanism? == | ||
Revision as of 12:29, 8 June 2008
Evidence from Similar Structures
- Screening of DALI result (see Basma) for functional information.
- Most similar structure well-characterised:
Byun JS, Rhee JK, Kim ND, Yoon J, Kim DU, Koh E, Oh JW, Cho HS. Crystal structure of hyperthermophilic esterase EstE1 and the relationship between its dimerization and thermostability properties. BMC Struct Biol. 2007 Jul 12;7:47.
- A thermostable carboxylesterase from an uncultured archaeon - isolated in an environmental sample.
Evidence from Similar Structures
- Catalytic triad identified in paper - Ser154, Asp251, and His281.
- To assess functional similarity, conservation of the catalytic triad was analysed.
- Catalytic triad conserved.
- Note: difference in clustalW alignment from DALI sequence alignment. Unstable region of alignment?