Results - 2gqnA: Difference between revisions
No edit summary |
|||
| Line 104: | Line 104: | ||
=='''Annotations for tRNA isopentenyltransferase'''== | |||
[[Image:annotation.JPG]] | [[Image:annotation.JPG]] | ||
Revision as of 09:23, 3 June 2008
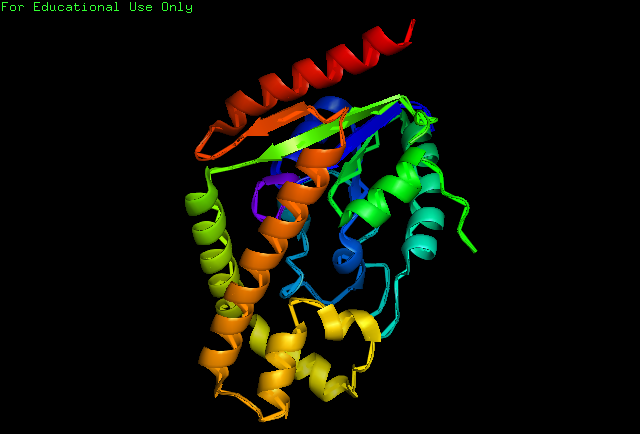
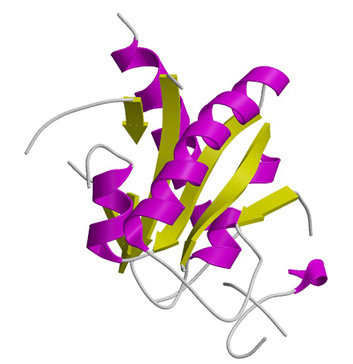
Structure of tRNA isopentenyltransferase
Protein Sequence in FASTA format
>gi|152149497|pdb|2QGN|A Chain A, Crystal Structure Of Trna Isopentenylpyrophosphate Transferase (Bh2366) From Bacillus Halodurans, Northeast Structural Genomics Consortium Target Bhr41. XKEKLVAIVGPTAVGKTKTSVXLAKRLNGEVISGDSXQVYRGXDIGTAKITAEEXDGVPHHLIDIKDPSE SFSVADFQDLATPLITEIHERGRLPFLVGGTGLYVNAVIHQFNLGDIRADEDYRHELEAFVNSYGVQALH DKLSKIDPKAAAAIHPNNYRRVIRALEIIKLTGKTVTEQARHEEETPSPYNLVXIGLTXERDVLYDRINR RVDQXVEEGLIDEAKKLYDRGIRDCQSVQAIGYKEXYDYLDGNVTLEEAIDTLKRNSRRYAKRQLTWFRN KANVTWFDXTDVDFDKKIXEIHNFIAGKLEEKSKLEHHHHHH
Protein Structure
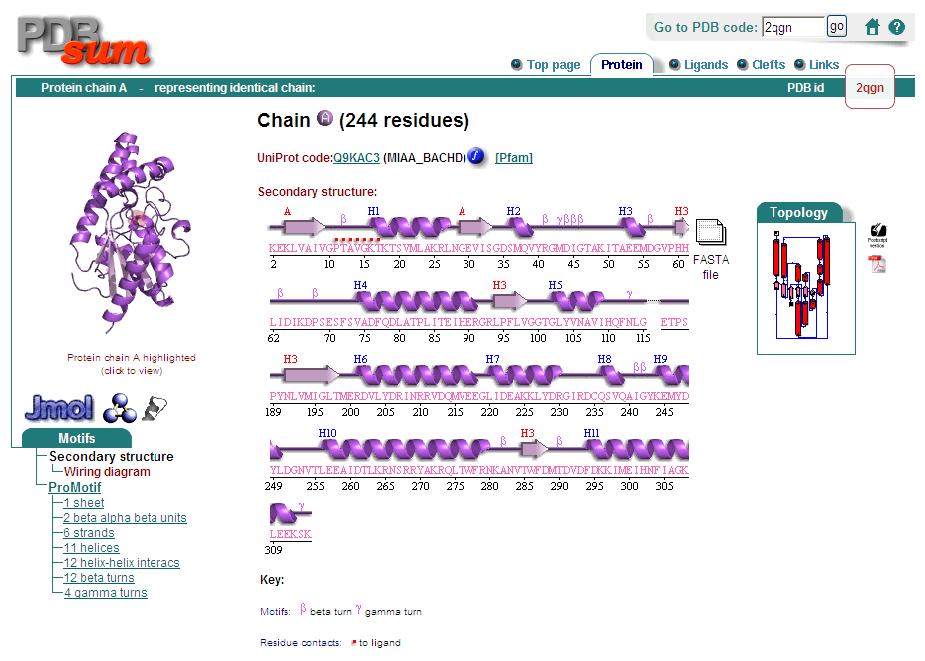
Secondary Structure
Analysis of the secondary structure acquired from Protein Data Bank showed results as displayed below :
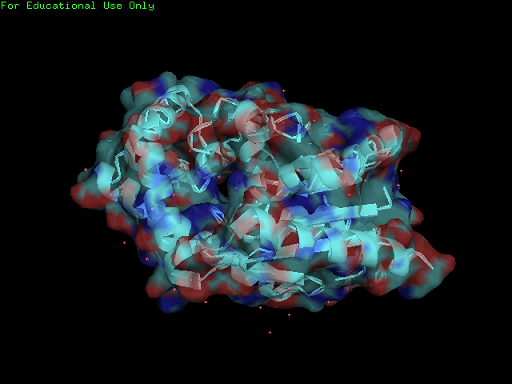
Surface Properties of 2qgn
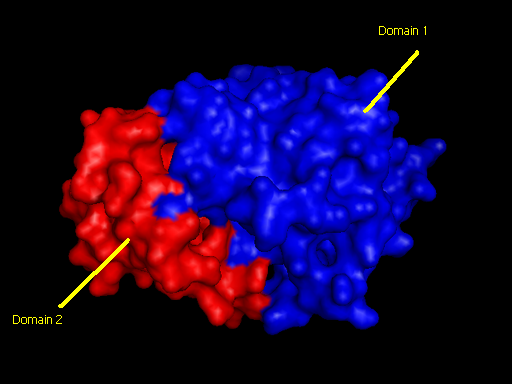
Domains
2qgnA is composed of two main domains. CATH analysis of 2qgn resulted in the finding of two main domains composing 2qgnA.
Domain 1 ranges from residue 2-200 and residue 283-314. Domain 2 encompasses residues stretching from 201-282.
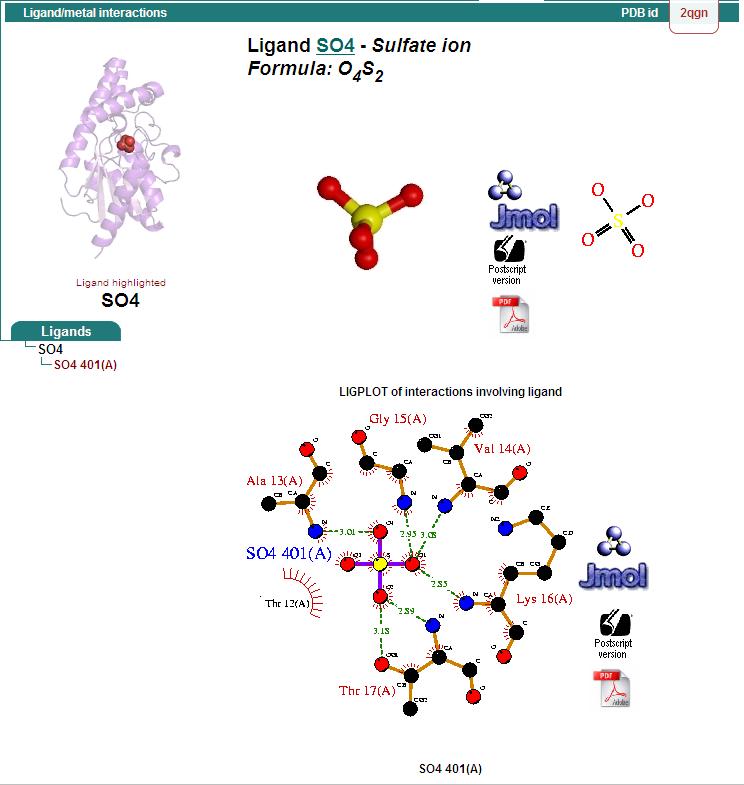
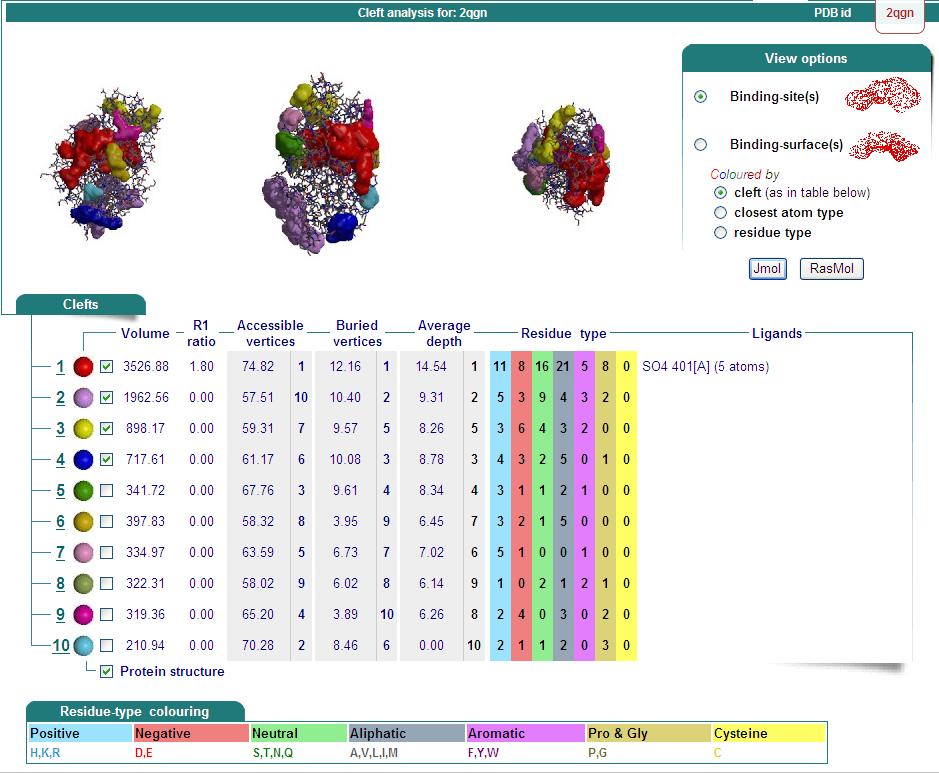
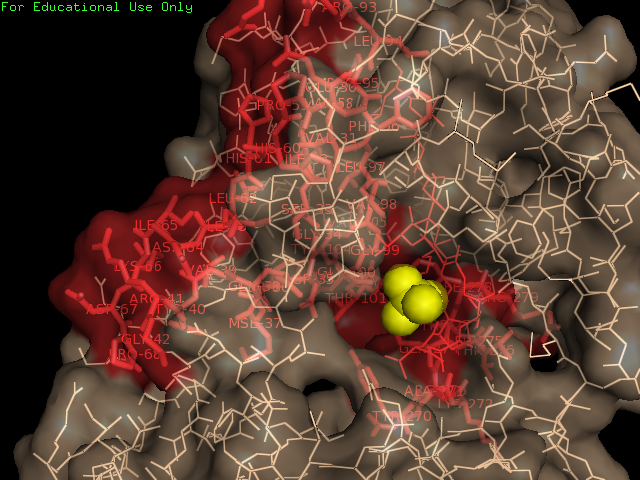
Ligand Binding Sites and Surface Clefts
Protein-ligand interaction
Hydrophillic binding sites
Bridged-H-bond binding sites
Hydrophobic binding sites
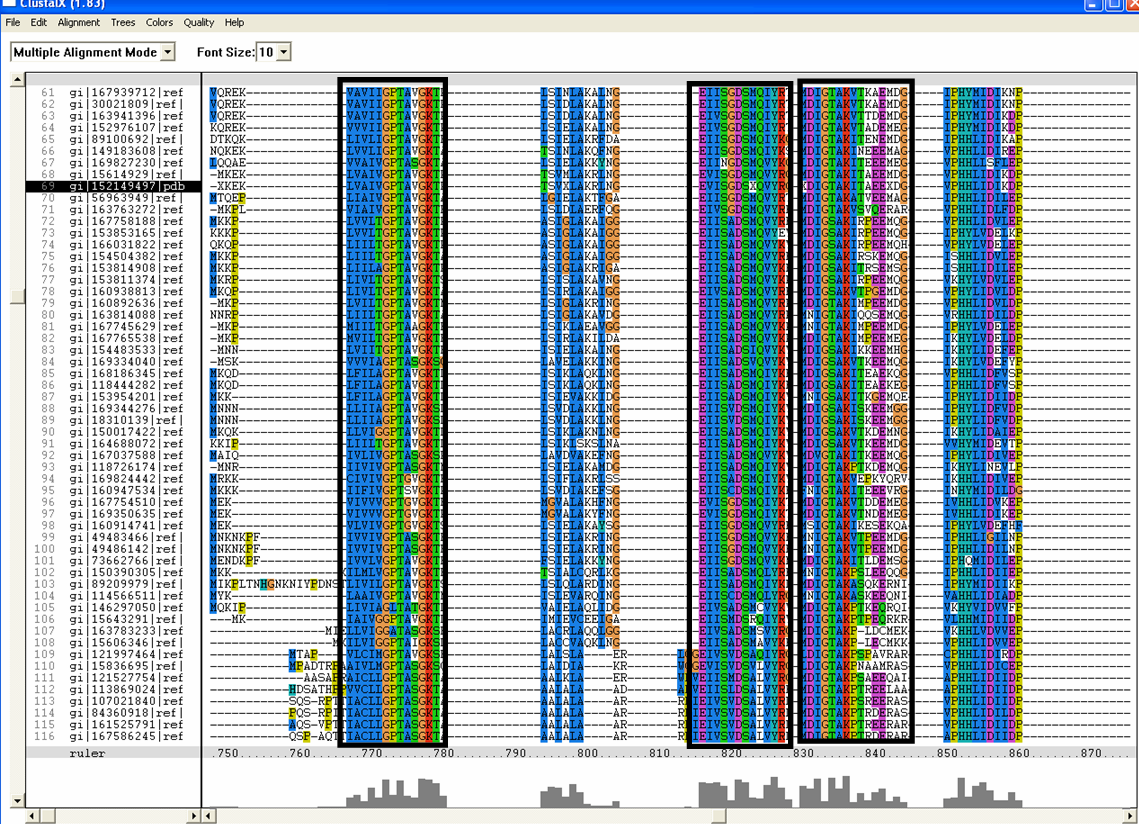
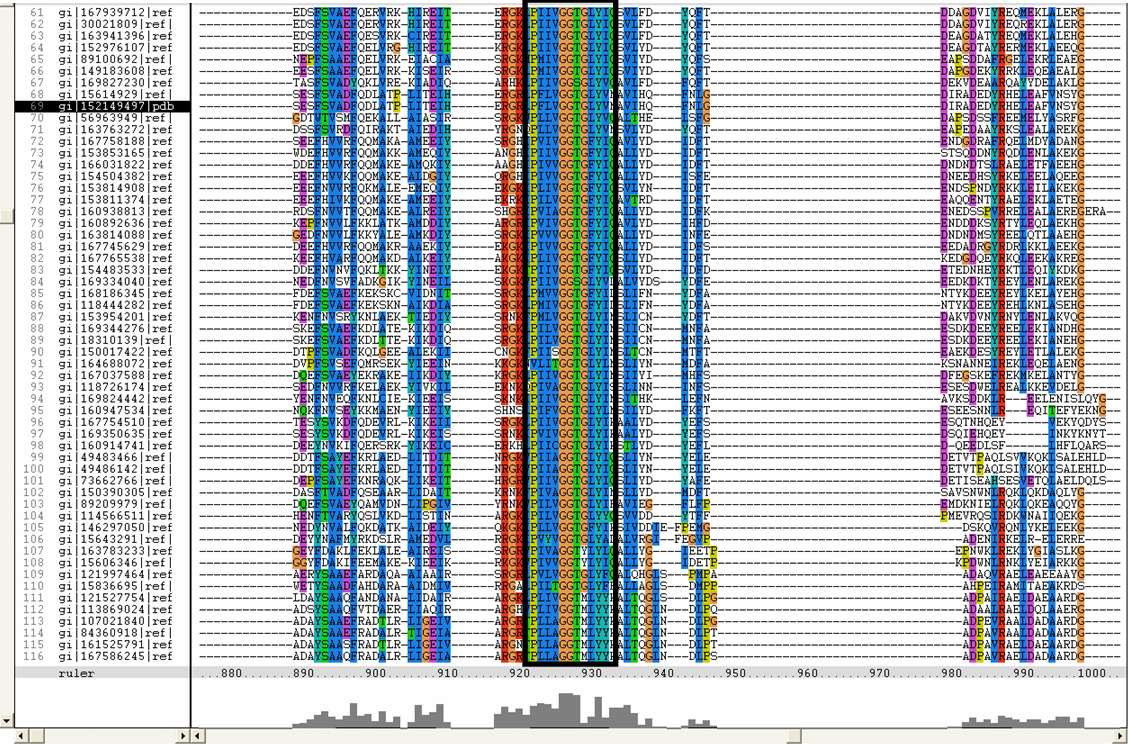
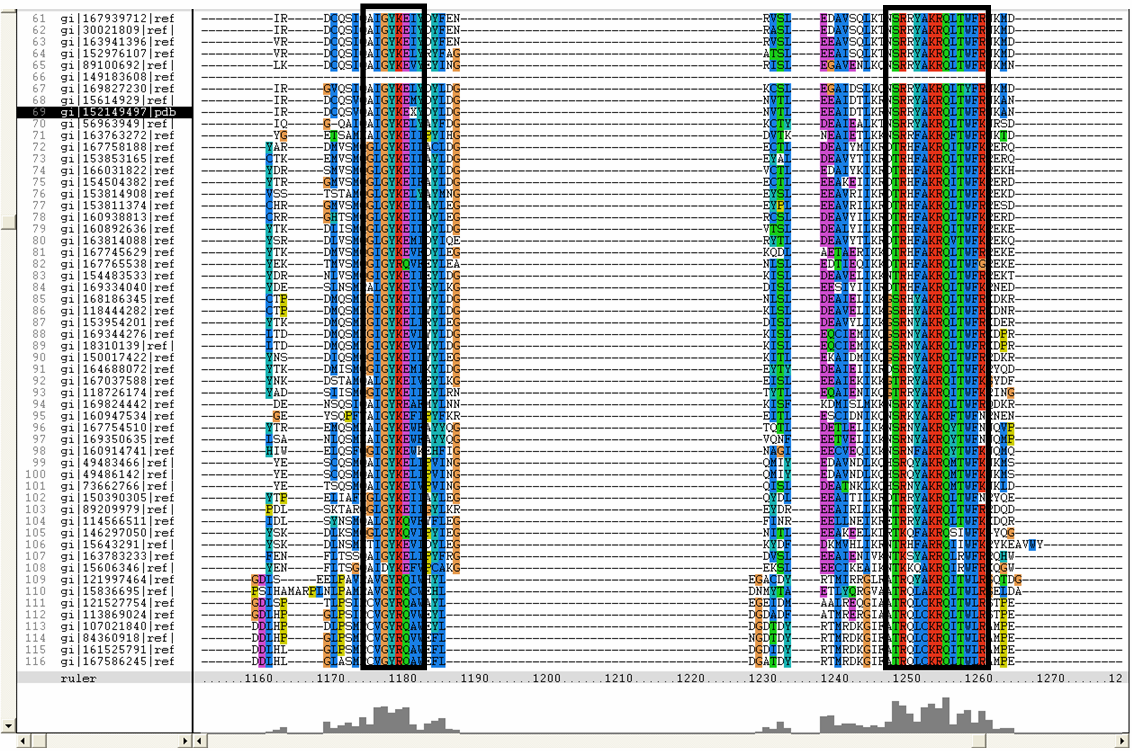
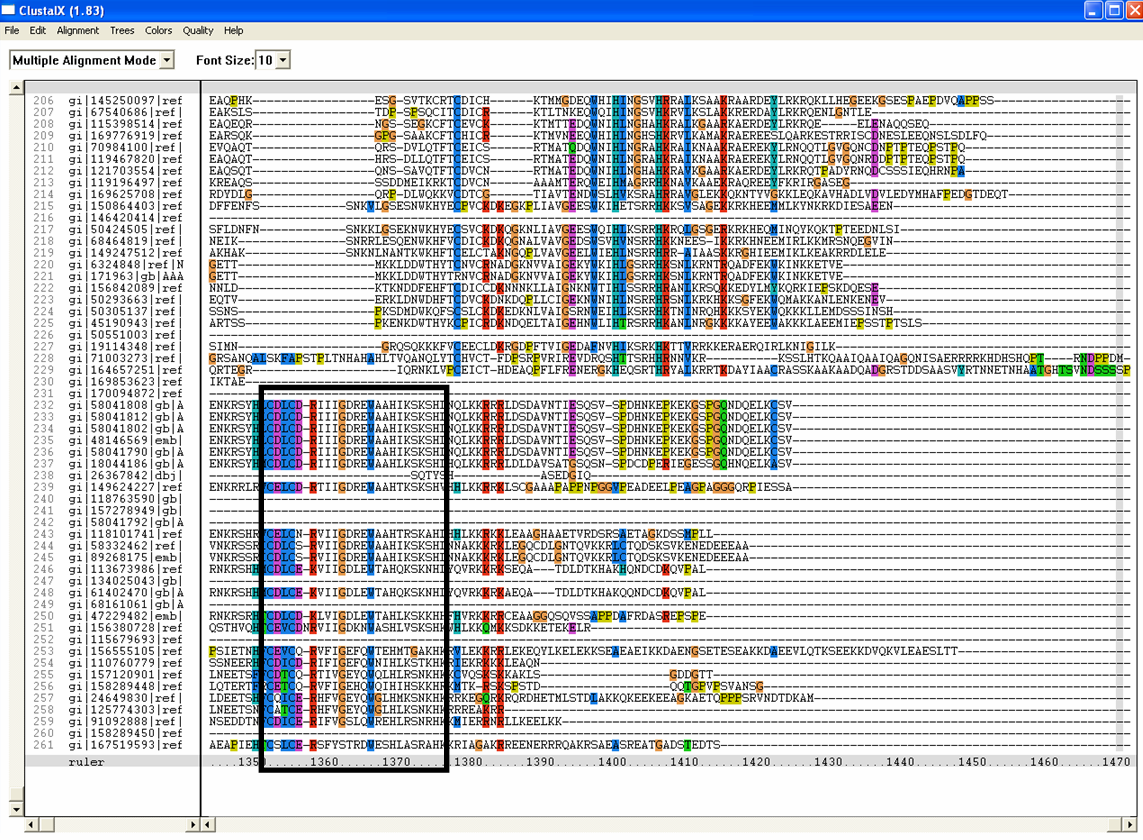
Conserved residues for tRNA isopentenyl transferase from Clustal alignment
Multiple sequence alignment from ClustalX allowed conserved regions in 2qgn and related species to be found.
Structural Alignment
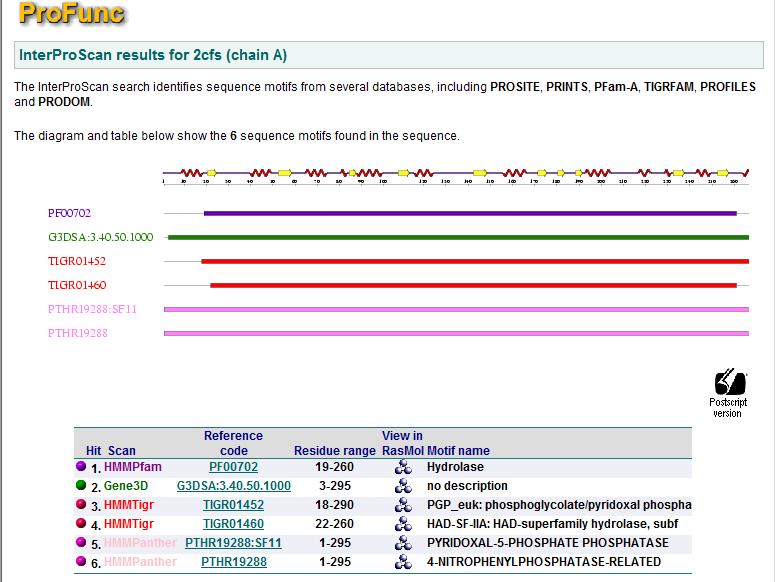
Profunc
Related protein sequences
Proteins with similar fold retrived from SSM (Secondary Structure Matching)
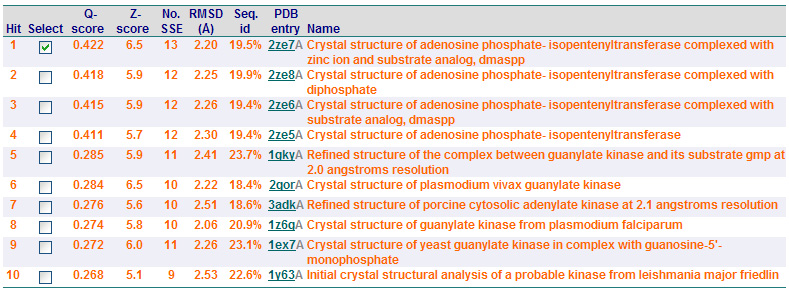
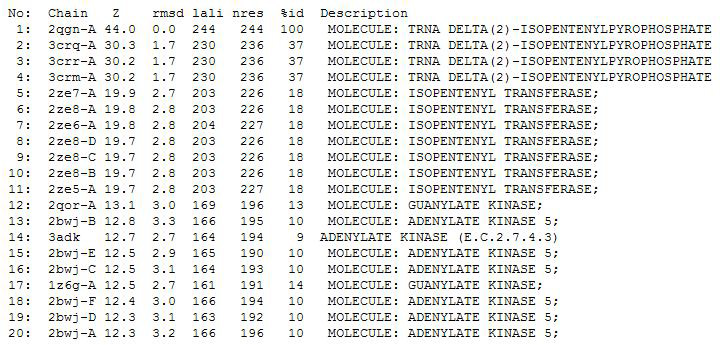
Dali Output
PDB entry code for 2qgn was loaded onto DALI server to search for structurally similar neighbours. Displayed below are the results from DALI search :-
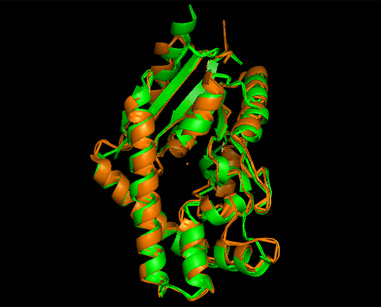
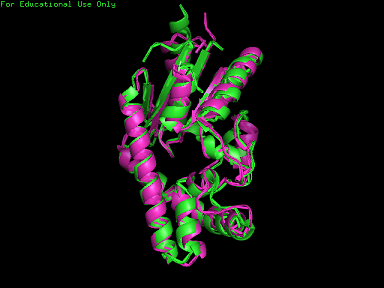
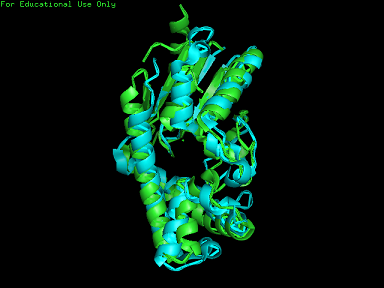
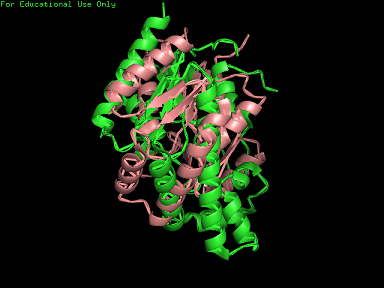
A total of 527 hits were found from DALI search, nonetheless only the first 20 hits that may be of significance were shown on the figure. Based on the outcome of DALI and Profunc, PDB files of each structurally similar protein was obtained from PDB. These were each superimposed against 2qgn using the PyMOL software, to compare the structural similiarity. Results are as below :
As indicated by the figures above, each structures were structurally similar to 2qgn, suggesting that they could have functionally similar properties. Nonetheless,notice that 2-qor is only partially similar to 2qgn structure. As the Z-score decreases for the DALI output, the structural similarity decreases as well. For this reason, functional analysis of 2qgn was only done for DALI outputs with lali scores higher than 200.
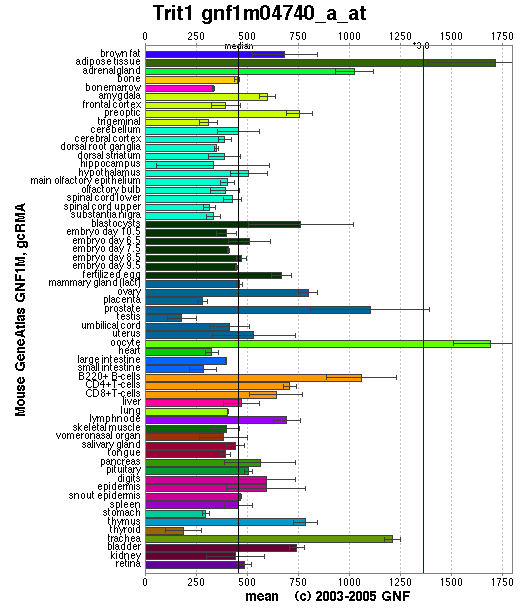
Localisation Expression of tRNA isopentenyltransferase
Generally, this enzyme is expressed in all tissue types since it is important that functional protein are synthesized in each of these tissues. Specifically, it is highly expressed in adipose tissues as well as oocytes. Relatively high amounts of this enzyme is expressed in prostate, adrenal gland, B-cells and trachea. The reason why tRNA-IPT are at higher concentrations in these tissues may reflect higher levels of protein synthesis.
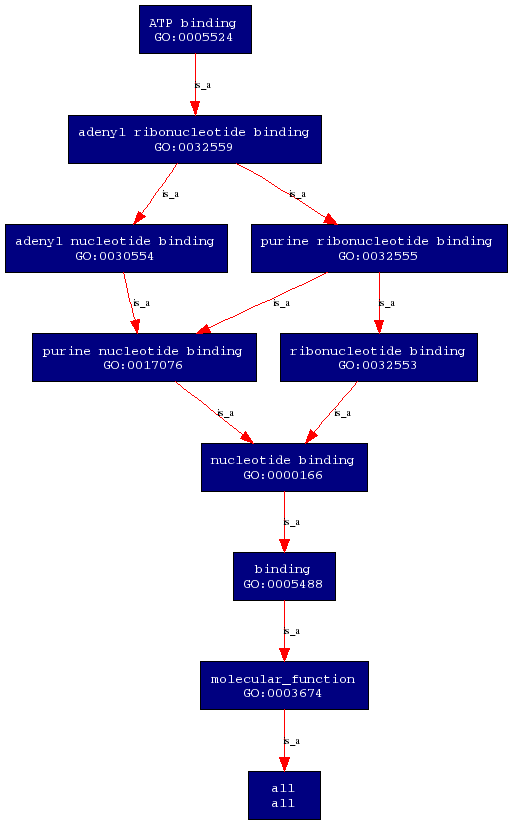
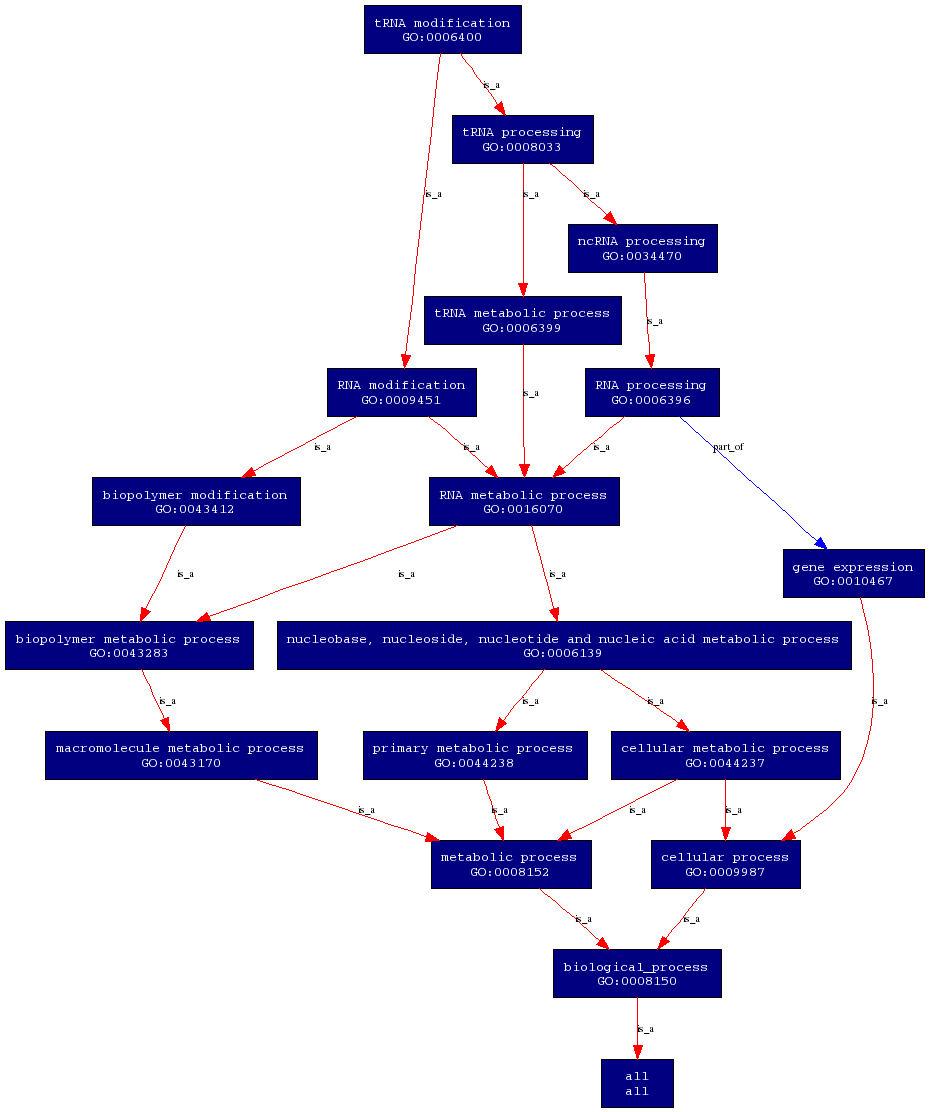
Molecular Function
Biological Process