Uncategorized files
From MDWiki
Jump to navigationJump to search
Showing below up to 250 results in range #1,001 to #1,250.
- MsbA Petsko07.pdf 0 × 0; 144 KB
- MultipleBlastProtSeq.txt ; 9 KB
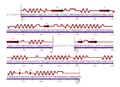
- Multipleestructuralalignmenta.jpeg 825 × 229; 54 KB
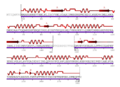
- Multiplestructuralalgnmentb.jpeg 893 × 214; 56 KB
- Multiseq align.jpg 839 × 331; 170 KB
- Mut4.gif 400 × 244; 10 KB
- Mutation1.png 220 × 77; 1 KB
- Mutation2.png 220 × 75; 1 KB
- Mutation3.png 240 × 75; 1 KB
- Mutation4.png 519 × 135; 5 KB
- N-terminus of LOC144557.txt ; 84 KB
- NKxD.jpg 146 × 653; 52 KB
- NKxD2.jpg 146 × 653; 52 KB
- NKxD3.jpg 468 × 742; 151 KB
- NKxD4.jpg 468 × 742; 151 KB
- NKxD5.jpg 514 × 736; 144 KB
- NUBP2BACK.jpg 200 × 200; 12 KB
- Names.jpg 1,051 × 437; 59 KB
- Nature encode.jpg 150 × 198; 32 KB
- Neighbortree.txt ; 100 KB
- Neighbourgene.jpg 770 × 680; 96 KB
- Neighbourtree2.txt ; 100 KB
- Nest.PNG 873 × 602; 79 KB
- Nest analysis result.PNG 545 × 320; 63 KB
- Nest analysis result.bmp 545 × 320; 511 KB
- Net.png 471 × 428; 29 KB
- Net image e 2H7YkMhiRnz6.png 471 × 456; 80 KB
- NewBOOT1000tree.png 934 × 720; 95 KB
- NewBOOTtree.png 1,050 × 812; 113 KB
- NewBOOTtree2.png 934 × 720; 95 KB
- Noble PLOSComputBiol2009.pdf 0 × 0; 130 KB
- NonredundantMSA.pdf 0 × 0; 454 KB
- NtermVS2oew.txt ; 208 KB
- NtermVs2cbi.txt ; 305 KB
- Nubp2front.jpg 200 × 200; 9 KB
- Nubp2top.jpg 200 × 200; 9 KB
- OG binding site.png 640 × 480; 26 KB
- Occurence.PNG 803 × 1,092; 57 KB
- Occurence.jpg 891 × 767; 48 KB
- Okamoto09.pdf 0 × 0; 305 KB
- Olf motif graph.GIF 623 × 382; 15 KB
- Olf motif table.GIF 915 × 994; 49 KB
- Olf trans pathway.gif 680 × 661; 39 KB
- Olfactory cGMP PDE2.PDF 0 × 0; 1.03 MB
- Ontology.jpg 608 × 468; 34 KB
- P1010632.jpg 1,620 × 2,236; 1.37 MB
- PBB GE DHRS1 213279 at fs.png 732 × 530; 12 KB
- PBB GE SELENBP1 214433 s at fs.png 732 × 530; 11 KB
- PDB.jpg 763 × 305; 41 KB
- PDB1.jpg 763 × 305; 41 KB
- PDBSUM.PNG 931 × 733; 145 KB
- PDBSUM.png 931 × 733; 145 KB
- PDBSum pblA.PNG 460 × 498; 31 KB
- PDB structure.jpg 250 × 250; 15 KB
- PDB sum 2.PNG 448 × 353; 38 KB
- PDBsum.JPG 440 × 652; 63 KB
- PDBsum.bmp 720 × 661; 1.36 MB
- PDBsum11.PNG 720 × 661; 90 KB
- PDBsum chain.JPG 878 × 734; 105 KB
- PDBsum cleft.PNG 803 × 681; 157 KB
- PDBsums.png 947 × 677; 69 KB
- PFam domains.png 638 × 477; 61 KB
- PHYH expression.png 520 × 790; 19 KB
- PICT1438.JPG 2,816 × 2,112; 1.11 MB
- PICT1438 copy.jpg 2,816 × 2,112; 1.11 MB
- PO4 400.gif 400 × 400; 1 KB
- PO4 interactions.png 389 × 406; 16 KB
- PPmotif.PNG 771 × 180; 27 KB
- PROFUNC Match To Existing PDB Stuctures.jpg 1,043 × 214; 45 KB
- PROFUNC Nest Analysis.jpg 645 × 363; 51 KB
- PROFUNC SSM.jpg 804 × 190; 43 KB
- PROKNOW-biological process.png 929 × 1,112; 20 KB
- PROKNOW2- molecular function.png 512 × 824; 9 KB
- PTS-1.jpg 282 × 66; 11 KB
- PTS-2.jpg 397 × 71; 15 KB
- PUBSUM.jpg 430 × 365; 18 KB
- Pastore febs07.pdf 0 × 0; 1.43 MB
- Pathway.jpg 773 × 261; 62 KB
- Pathways.png 421 × 590; 67 KB
- Pattern0.txt ; 31 bytes
- Pbio GOS 120.gif 120 × 120; 5 KB
- Pbl.png 640 × 480; 182 KB
- Pdb cartoon 2ece.png 680 × 846; 12 KB
- Pdb sum.png 864 × 648; 185 KB
- Pdbsum 1senA.gif 430 × 233; 11 KB
- Pdbsum 3fpu.gif 430 × 256; 13 KB
- Pdbsums archeal.PNG 894 × 528; 66 KB
- Pep 1.PNG 804 × 696; 91 KB
- Peptide Folding- When Simulation Meets Experiment.pdf 0 × 0; 270 KB
- Petri.jpg 388 × 385; 76 KB
- Pfam-a.jpg 1,066 × 108; 85 KB
- Pfam result.jpg 960 × 720; 114 KB
- Phillips04.pdf 0 × 0; 395 KB
- Phylo radial1.png 947 × 644; 30 KB
- Phylo radial1a.png 863 × 587; 42 KB
- Phylo rect1.png 1,007 × 517; 39 KB
- Phylo tree.jpg 1,146 × 633; 61 KB
- Phylogenetic Tree.jpg 1,280 × 1,024; 141 KB
- Phylogenic tree.JPG 680 × 563; 47 KB
- Phylogenic tree3.jpg 680 × 563; 47 KB
- Phylogram.png 1,280 × 768; 50 KB
- Pi helix.gif 58 × 20; 1 KB
- Picture1.jpg 793 × 655; 42 KB
- Picture2.jpg 333 × 437; 7 KB
- Picture3.jpg 412 × 395; 14 KB
- Picture4.png 1,465 × 941; 2.81 MB
- Picture5.jpg 1,502 × 811; 337 KB
- Picture6.jpg 1,427 × 704; 176 KB
- Picture 1.png 495 × 455; 180 KB
- Picture 2.png 514 × 407; 40 KB
- Plasm.png 939 × 461; 202 KB
- Plasmodium.png 939 × 461; 202 KB
- Platypus.png 1,261 × 629; 120 KB
- Po4 ligand.JPG 763 × 879; 72 KB
- Pocket.gif 640 × 480; 1.65 MB
- Pocket.jpg 1,062 × 570; 74 KB
- Pocket surface.gif 640 × 480; 3.36 MB
- Possiblecatalyticresidues.png 640 × 434; 92 KB
- Pprofunc superfamily.JPG 1,280 × 1,024; 127 KB
- Prestegard ms1.pdf 0 × 0; 36 KB
- Prestegard ms2.pdf 0 × 0; 67 KB
- Pretty.png 640 × 434; 79 KB
- Pretty protein.PNG 613 × 660; 387 KB
- ProFunc Image.jpg 480 × 480; 19 KB
- ProFunc ImageJen.jpg 480 × 480; 19 KB
- Profunc, GO.png 960 × 720; 226 KB
- Profunc.jpg 775 × 582; 74 KB
- Profunc2.jpg 1,089 × 261; 73 KB
- ProfuncNest2.PNG 651 × 577; 41 KB
- ProfuncNests.JPG 651 × 614; 235 KB
- Profunc GO.jpg 960 × 720; 102 KB
- Profunc NAR2005.pdf 0 × 0; 1.95 MB
- Profunc jmb2007.pdf 0 × 0; 908 KB
- Profunc nest.PNG 622 × 195; 13 KB
- Profunc nest.bmp 622 × 195; 356 KB
- Profunc nests.JPG 651 × 614; 84 KB
- Profunc nests.jpg 651 × 614; 84 KB
- Proknow Eisenberg Structure2005.pdf 0 × 0; 480 KB
- Prositeprediction.jpg 477 × 599; 23 KB
- Prositescan.png 477 × 599; 23 KB
- Protdist1.txt ; 1.24 MB
- Protdist2.txt ; 1.24 MB
- Protein-result.txt ; 6 KB
- Protein-results.txt ; 6 KB
- Protein2.JPG 2,102 × 434; 79 KB
- Protein interaction.jpg 471 × 456; 80 KB
- Protein sequence.JPG 606 × 146; 33 KB
- Protein showing zinc pymol.png 1,060 × 906; 232 KB
- PsaA docked to E-cadherin in simulation cell.png 1,044 × 941; 739 KB
- Psiblast.txt ; 1.03 MB
- PyMOL 2oycA vs 2cfsA.jpg 555 × 397; 32 KB
- Error creating thumbnail: convert: Expected 8192 bytes; found 8119 bytes `/var/www/mediawiki/upload/2/29/Pymol.PNG.png' @ warning/png.c/MagickPNGWarningHandler/1669. convert: Read Exception `/var/www/mediawiki/upload/2/29/Pymol.PNG.png' @ error/png.c/MagickPNGErrorHandler/1643. convert: no images defined `/tmp/transform_ee25e6271e32.png' @ error/convert.c/ConvertImageCommand/3229. Error code: 1Pymol.PNG.png 1,060 × 906; 713 KB
- Pymol.png 640 × 434; 167 KB
- Pymol hsp movie0001.png 580 × 482; 53 KB
- Pymol ss1.PNG 406 × 370; 59 KB
- Pyridoxal Phosphatase MSA sequences.pdf 0 × 0; 694 KB
- Pyridoxal Phosphatase Rectangular Tree.jpg 3,508 × 2,480; 485 KB
- Pyridoxal phosphatase BLAST.pdf 0 × 0; 250 KB
- Pyridoxal phosphatase MSA Page 1.jpg 783 × 377; 59 KB
- Pyridoxal phosphatase MSA Page 2.jpg 783 × 377; 170 KB
- Pyridoxal phosphatase MSA Page 3.jpg 783 × 377; 202 KB
- Pyridoxal phosphatase MSA Page 4.jpg 306 × 369; 73 KB
- Pyridoxal phosphatase MSA sequences conserved .jpg 927 × 928; 457 KB
- Pyridoxal phosphatase Radial Cladogram.jpg 3,508 × 2,480; 477 KB
- Pyridoxal phosphatase Rectangular Phylogeny.pdf 0 × 0; 176 KB
- Pyridoxal phosphatase Rectangular Tree discussion1.jpg 1,768 × 704; 97 KB
- Pyridoxal phosphatase conserved image.jpg 640 × 480; 105 KB
- Querol2003.pdf 0 × 0; 827 KB
- Querol2006.pdf 0 × 0; 145 KB
- Question mark.jpg 10 × 18; 539 bytes
- RNA template 1hu3.gif 732 × 377; 33 KB
- RR1.png 798 × 577; 574 KB
- RR2.png 791 × 521; 506 KB
- RR3.png 793 × 526; 451 KB
- Rad.JPG 967 × 839; 71 KB
- Rad2.JPG 1,048 × 859; 67 KB
- Radial.JPG 967 × 839; 71 KB
- Radial2.JPG 1,048 × 859; 67 KB
- Radialtree.JPG 656 × 607; 62 KB
- Radiationtreexx22.jpg 789 × 722; 38 KB
- Ramachandran.jpg 786 × 393; 47 KB
- Ramachandran 1.jpg 816 × 391; 34 KB
- Rat fatty acid binding protein 2IFB.jpg 500 × 500; 47 KB
- Rbsfunction.jpg 662 × 201; 31 KB
- Rdc key equations.pdf 0 × 0; 271 KB
- Reaction.jpg 506 × 146; 10 KB
- Reaction.png 866 × 783; 20 KB
- Reaction1.png 530 × 242; 10 KB
- ReactionASK.png 416 × 92; 13 KB
- Reaction Arrow.jpg 26 × 21; 775 bytes
- Red x.png 25 × 28; 3 KB
- Redox Halves.png 892 × 227; 7 KB
- Region1.png 1,280 × 1,024; 972 KB
- Region2.png 1,272 × 929; 153 KB
- Region3.png 1,272 × 929; 138 KB
- Rhodopseudomonas palustris chromosome.jpg 900 × 628; 88 KB
- Ribbon structure.png 640 × 434; 82 KB
- Rootedtree1.jpg 1,170 × 618; 53 KB
- Rootedtreeboot.jpg 1,170 × 618; 50 KB
- Rootedtreecol1.jpg 1,170 × 618; 67 KB
- Rootedtreecol2.jpg 1,170 × 618; 71 KB
- Roux1999 BiophysChem78-1.pdf 0 × 0; 211 KB
- Ruscio2008 PNAS105-9204.pdf 0 × 0; 697 KB
- Russel Structure2004.pdf 0 × 0; 287 KB
- SABLE.gif 550 × 664; 19 KB
- SAMPL.png 522 × 132; 49 KB
- SAS Aligned Sequences.jpg 1,190 × 243; 66 KB
- SBP with Sele Atom.png 640 × 480; 161 KB
- SBP with Sele atom.png 640 × 480; 161 KB
- SBP with sele atom.png 640 × 480; 161 KB
- SCOP.JPG 1,167 × 122; 36 KB
- SECONDARY.jpg 503 × 471; 9 KB
- SPLIT combined alan.pdf 0 × 0; 34.95 MB
- SSAME.png 640 × 434; 97 KB
- SSM1.PNG 791 × 520; 50 KB
- SSM2.PNG 792 × 402; 54 KB
- SSMPhosphatases.jpg 3,124 × 1,013; 551 KB
- SSMPhosphatases.pdf 0 × 0; 21 KB
- SSP.png 1,216 × 445; 49 KB
- STSsite1.png 660 × 629; 190 KB
- STSsite2.png 660 × 629; 190 KB
- STSwhole.png 616 × 603; 215 KB
- SURFACE.jpg 986 × 768; 118 KB
- Schematic.jpg 590 × 444; 35 KB
- Schultz chembiochem02.pdf 0 × 0; 101 KB
- Schultz currentopinionChemBiol05.pdf 0 × 0; 401 KB
- Schultz nature methods07.pdf 0 × 0; 389 KB
- Schultz pnas02.pdf 0 × 0; 562 KB
- Scop.png 404 × 217; 44 KB
- Secondary.jpg 925 × 661; 92 KB
- Secondary Structure.png 537 × 623; 42 KB
- Secondary Structure 2cfsA.jpg 486 × 765; 77 KB
- Secondary Structure 2oycA.jpg 466 × 731; 69 KB
- Secondary structure.gif 430 × 600; 24 KB
- Secondary structure.jpg 540 × 390; 23 KB
- Secondary structure.png 540 × 390; 23 KB
- Secondary structure.txt ; 559 bytes
- Secondary structure comparsion.jpg 1,215 × 480; 70 KB
- Secondary structures.pdf 0 × 0; 13.24 MB
- SelNonredundantMSA.PNG 1,344 × 646; 113 KB
- SelNonredundantMSA.jpg 2,120 × 1,590; 1.45 MB
- SelNonredundantMSA.pdf 0 × 0; 171 KB
- Seminario Hess2FF.pdf 0 × 0; 480 KB
- Seq1.jpg 732 × 78; 4 KB
- Seq2.jpg 593 × 111; 14 KB
- Sequence.jpg 791 × 432; 36 KB
- Sequence1-cml.jpg 750 × 398; 102 KB
- Sequence & secondary structure PDB.bmp 0 × 0; 906 KB