Unused files
From MDWiki
Jump to navigationJump to search
The following files exist but are not embedded in any page. Please note that other web sites may link to a file with a direct URL, and so may still be listed here despite being in active use.
Showing below up to 423 results in range #1 to #423.
- Thomas125.jpg 125 × 125; 28 KB
- 11 Structure2005.pdf 0 × 0; 894 KB
- Profunc NAR2005.pdf 0 × 0; 1.95 MB
- WIKI.png 730 × 579; 237 KB
- PDB structure.jpg 250 × 250; 15 KB
- 2gfh asym r 500.jpg 500 × 500; 35 KB
- 1zkd asym r 250.jpg 250 × 250; 17 KB
- Fasta seq output of entrez.txt ; 31 KB
- Similar sequences.txt ; 165 KB
- MultipleBlastProtSeq.txt ; 9 KB
- Acession numbers for blast results.txt ; 256 bytes
- HumanOrthologProtSeq.txt ; 534 bytes
- HumanOrthologProtSeqBlast1.txt ; 52 KB
- FASTA.txt ; 249 bytes
- Sequences2.txt ; 6 KB
- Ids2.txt ; 879 bytes
- 2gfh.jpg 115 × 115; 4 KB
- COASYAcobridgedhbond.jpg 517 × 531; 63 KB
- 2gfh 3 bottom.jpg 240 × 240; 16 KB
- 2gfh 3 main.pg.jpg 240 × 240; 12 KB
- 2gfh 3 right.pg.jpg 240 × 240; 16 KB
- Acobridgedhbond.jpg 517 × 531; 63 KB
- CoA Synthesis localistion.png 520 × 610; 14 KB
- 2ho4.850.jpg 850 × 850; 166 KB
- Protein-result.txt ; 6 KB
- Chemical reaction.JPG 640 × 512; 9 KB
- Tree.JPG 819 × 668; 39 KB
- C-terminus of LOC144557.txt ; 94 KB
- N-terminus of LOC144557.txt ; 84 KB
- CoAA - Pantothenate kinase expression data (mouse).png 520 × 610; 14 KB
- 2ex4 asym r 250.jpg 250 × 250; 15 KB
- 1im8 bio r 250.jpg 250 × 250; 15 KB
- 1im8 bio r 251.jpg 250 × 250; 16 KB
- LargeSimilarsequences3.txt ; 29 KB
- Alignment 2ifu & 1qqe.JPG 640 × 580; 36 KB
- NtermVs2cbi.txt ; 305 KB
- NtermVS2oew.txt ; 208 KB
- Tree alignment.JPG 5,098 × 6,599; 515 KB
- Bootstraptree.JPG 651 × 950; 85 KB
- Bootstraptreeunrooted.JPG 651 × 950; 75 KB
- Unrootedtree2.JPG 517 × 476; 57 KB
- Tree alignmnet.JPG 1,280 × 998; 96 KB
- Snap-gamma.png 435 × 317; 85 KB
- Radial2.JPG 1,048 × 859; 67 KB
- 2ifu,1qqe alignment.png 454 × 234; 77 KB
- Phylogenic tree.JPG 680 × 563; 47 KB
- COASYTmhmm graph.gif 710 × 350; 3 KB
- Phylogenetic Tree.jpg 1,280 × 1,024; 141 KB
- Tree with names.JPG 1,133 × 813; 104 KB
- MSA.JPG 1,280 × 1,024; 514 KB
- EIF4G w groove.JPG 512 × 384; 9 KB
- Treep.JPG 486 × 267; 30 KB
- Radial.JPG 967 × 839; 71 KB
- Rad2.JPG 1,048 × 859; 67 KB
- Tempalign.gif 650 × 352; 60 KB
- ProFunc Image.jpg 480 × 480; 19 KB
- Pprofunc superfamily.JPG 1,280 × 1,024; 127 KB
- LOC144557(with ligand bound).doc ; 128 KB
- Olfactory cGMP PDE2.PDF 0 × 0; 1.03 MB
- Treetrans.JPG 449 × 257; 15 KB
- Image002.jpg 415 × 175; 18 KB
- Secondary structure comparsion.jpg 1,215 × 480; 70 KB
- Fig 1.bmp 0 × 0; 1.96 MB
- Document2 05.png 229 × 145; 3 KB
- Fig 2.PNG 556 × 354; 7 KB
- MSAfigure2.JPG 899 × 864; 335 KB
- ClustalX image.JPG 1,203 × 618; 390 KB
- ClustalX.jpg 1,203 × 618; 381 KB
- Fig1 chains.jpg 707 × 571; 66 KB
- Fig2 properties.jpg 960 × 535; 110 KB
- Table1.bmp 1,067 × 543; 69 KB
- NKxD.jpg 146 × 653; 52 KB
- NKxD2.jpg 146 × 653; 52 KB
- Msaseg1a.JPG 863 × 433; 144 KB
- Msaseg1b.PNG 947 × 475; 148 KB
- Phylo radial1.png 947 × 644; 30 KB
- WalkerAB.jpg 736 × 653; 262 KB
- TREE.jpg 628 × 525; 47 KB
- 2ifuA secondary structure.png 478 × 395; 22 KB
- Ligand stucture and environment.png 1,347 × 416; 209 KB
- Cleft analysis 2i2O.jpg 0 × 0; 1.1 MB
- 2ifuA interpro .png 586 × 304; 20 KB
- NKxD3.jpg 468 × 742; 151 KB
- NKxD4.jpg 468 × 742; 151 KB
- Fig1 properties.JPG 962 × 1,112; 176 KB
- Secondary structure.jpg 540 × 390; 23 KB
- 2ifua hydrophobes and hydrophilics.png 772 × 660; 301 KB
- Pep 1.PNG 804 × 696; 91 KB
- WalkerAB2.jpg 933 × 754; 323 KB
- Mel's picture of secondary str..bmp 0 × 0; 906 KB
- Dotlet.PNG 508 × 336; 85 KB
- IfuA-structure.png 1,144 × 344; 241 KB
- GOterm.JPG 962 × 748; 77 KB
- 2gnx.jpg 250 × 250; 9 KB
- 2GNX.doc ; 20 KB
- Domain.jpg 804 × 696; 93 KB
- Domains.jpg 804 × 696; 53 KB
- Fold.bmp 0 × 0; 339 KB
- Ligand structure visualisation.jpg 1,161 × 827; 280 KB
- Alignment.bmp 0 × 0; 1.62 MB
- Alignment.jpeg 1,001 × 565; 191 KB
- Tree.jpeg 1,001 × 1,013; 36 KB
- MSAAnnotated.aln ; 59 KB
- MG2-.jpg 800 × 300; 7 KB
- Domain.JPG 395 × 104; 7 KB
- Bash-Beginners-Guide.pdf 0 × 0; 1.16 MB
- Red x.png 25 × 28; 3 KB
- Green tick.png 24 × 24; 2 KB
- Install.txt ; 15 KB
- Rdc key equations.pdf 0 × 0; 271 KB
- Biomosagain.jpg 794 × 1,163; 340 KB
- Alanwithteddy.jpg 1,141 × 794; 340 KB
- Schultz pnas02.pdf 0 × 0; 562 KB
- Ito pnas07.pdf 0 × 0; 1.02 MB
- Structure of protein.doc ; 525 KB
- Structure.jpg 660 × 738; 1.39 MB
- 2ece bio r 500 structure.jpg 500 × 500; 57 KB
- 2ECE.fasta.txt ; 497 bytes
- 2qgnA.png 640 × 434; 85 KB
- Arylformamidase function.jpg 445 × 111; 7 KB
- Chain A.PNG 286 × 364; 89 KB
- Whole protein.png 710 × 745; 310 KB
- Confidence interaction with names.PNG 727 × 540; 72 KB
- HomoCLUSTAL.aln ; 131 KB
- BacterCLUSTAL.aln ; 50 KB
- CARBOXYLASE.PNG 830 × 699; 421 KB
- Carboxylase A chain.bmp 0 × 0; 960 KB
- Carboxylase A chain.PNG 640 × 512; 135 KB
- Carboxylesterase Chain A.txt ; 2 KB
- PDBSUM.png 931 × 733; 145 KB
- PDBSUM.PNG 931 × 733; 145 KB
- Topology.gif 353 × 706; 13 KB
- 640-730.bmp 0 × 0; 2.25 MB
- PDBsums.png 947 × 677; 69 KB
- 1-170.png 1,280 × 1,024; 196 KB
- 1-170aa.jpg 1,024 × 820; 249 KB
- 511-680.png 1,280 × 1,024; 92 KB
- HomoBLAST.aln ; 2.93 MB
- Example1.jpg ; 131 KB
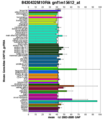
- PBB GE DHRS1 213279 at fs.png 732 × 530; 12 KB
- StrappedTree.bmp 0 × 0; 3.75 MB
- Xxxxx1.PNG 1,280 × 1,024; 136 KB
- Clustal Alignment top 5.txt ; 4 KB
- StrappedTree.png 1,152 × 922; 92 KB
- DanielTopology2.gif 430 × 367; 19 KB
- MSA11.jpg 1,280 × 889; 116 KB
- MSA2.jpg 1,267 × 850; 193 KB
- MSA3.jpg 1,264 × 847; 161 KB
- MSA4.jpg 1,264 × 849; 131 KB
- MSA6.jpg 1,263 × 849; 151 KB
- MSA5.jpg 1,267 × 847; 199 KB
- MSA7.jpg 1,267 × 844; 128 KB
- MSA8.jpg 1,266 × 850; 101 KB
- Secondary Structure.png 537 × 623; 42 KB
- Dehydrogenasereductase (SDR family) member 1 phylogram.png 835 × 608; 51 KB
- Examplex.jpg 1,006 × 179; 30 KB
- 2c7b DALI alignment.png 910 × 851; 301 KB
- BacterTREE.jpg 856 × 621; 55 KB
- Tree3.png 1,280 × 1,024; 59 KB
- Zn Cl n pocket.png 904 × 638; 299 KB
- Crystal Structure 2cfsA.jpg 240 × 226; 10 KB
- Geneatlas human.png 520 × 790; 19 KB
- Sequencemsa.pdf 0 × 0; 202 KB
- 2NXF showing all four major cavities.png 480 × 380; 98 KB
- Net image e 2H7YkMhiRnz6.png 471 × 456; 80 KB
- Treepic1.JPG 5,105 × 6,601; 732 KB
- StrappedTree.PNG 1,181 × 840; 41 KB
- Error creating thumbnail: File with dimensions greater than 12.5 MPTreegif.gif 5,105 × 6,601; 44 KB
- Surface prop.png 512 × 384; 185 KB
- Consensetree.jpg 1,887 × 1,187; 80 KB
- Consense.jpg 1,887 × 1,187; 134 KB
- Msa1.bmp 0 × 0; 1.81 MB
- 2qgn3.png 384 × 288; 70 KB
- 2c7b alignment.jpg 640 × 602; 71 KB
- 2cb7alignment.jpg 640 × 602; 71 KB
- Error creating thumbnail: convert: Expected 8192 bytes; found 8119 bytes `/var/www/mediawiki/upload/2/29/Pymol.PNG.png' @ warning/png.c/MagickPNGWarningHandler/1669. convert: Read Exception `/var/www/mediawiki/upload/2/29/Pymol.PNG.png' @ error/png.c/MagickPNGErrorHandler/1643. convert: no images defined `/tmp/transform_c342f5ee39aa.png' @ error/convert.c/ConvertImageCommand/3229. Error code: 1Pymol.PNG.png 1,060 × 906; 713 KB
- NewBOOTtree.png 1,050 × 812; 113 KB
- Arylformamidase.txt ; 302 KB
- ConservedRegion 1.bmp 0 × 0; 4.47 MB
- File.jpg 0 × 0; 4.47 MB
- Tree name group.bmp 0 × 0; 59 KB
- ConservedRegion 2.png 1,021 × 681; 89 KB
- ConservedRegion 1.png 1,024 × 687; 95 KB
- ConservedRegion 3.png 1,024 × 685; 80 KB
- Tree name group.png 987 × 640; 109 KB
- Dali output.jpg 721 × 345; 166 KB
- 2qgn4.jpg 640 × 480; 109 KB
- 2qgn42.jpg 410 × 307; 81 KB
- 2qgn42.png 410 × 307; 81 KB
- Motif.jpg 754 × 472; 63 KB
- Wholepic.jpg 850 × 850; 68 KB
- Domain-1.jpg 510 × 510; 61 KB
- Domain pic.jpg 448 × 304; 71 KB
- Domain pic2.jpg 512 × 384; 112 KB
- Active Site.JPG 640 × 480; 20 KB
- Active Site of 2cfsA.JPG 531 × 403; 17 KB
- AS2CFS.JPG 640 × 480; 20 KB
- Arylformamidase alignment.jpg 1,116 × 524; 231 KB
- 2CFS Active Site.JPG 640 × 480; 20 KB
- KW Active Site.jpg 640 × 480; 20 KB
- Error creating thumbnail: File missing2CFTA Structure.jpg 700 × 562; 53 KB
- Ssm.jpg 789 × 293; 190 KB
- Rootedtree1.jpg 1,170 × 618; 53 KB
- Rootedtreeboot.jpg 1,170 × 618; 50 KB
- Chain A.jpg 499 × 514; 193 KB
- ChainA.PNG 499 × 514; 193 KB
- UQGreatCourt.JPG 320 × 240; 66 KB
- MDgroup2.JPG 480 × 360; 144 KB
- MDgroup.JPG 480 × 360; 133 KB
- Archeaon Carboxylesterase.PNG 438 × 484; 169 KB
- Mdgroup2.jpg 480 × 360; 144 KB
- Cattriad 2c7b.PNG 477 × 438; 150 KB
- Carton pocket 1.png 640 × 480; 80 KB
- Cartoon behind pocket 1.png 640 × 480; 79 KB
- NewBOOTtree2.png 934 × 720; 95 KB
- Iron binding 2opwB.jpg 640 × 480; 12 KB
- Pyridoxal phosphatase MSA Page 1.jpg 783 × 377; 59 KB
- Pyridoxal phosphatase MSA Page 2.jpg 783 × 377; 170 KB
- Pyridoxal phosphatase MSA Page 3.jpg 783 × 377; 202 KB
- Pyridoxal phosphatase MSA Page 4.jpg 306 × 369; 73 KB
- Region2.png 1,272 × 929; 153 KB
- Region3.png 1,272 × 929; 138 KB
- Conserved.jpg 512 × 384; 316 KB
- Pyridoxal Phosphatase MSA sequences.pdf 0 × 0; 694 KB
- Clustalx1.jpg 0 × 0; 3.75 MB
- Region1.png 1,280 × 1,024; 972 KB
- CR1.png 1,139 × 824; 882 KB
- CR2.png 1,129 × 744; 720 KB
- CR3.png 1,132 × 751; 625 KB
- ZINC.png 1,143 × 840; 528 KB
- Tree name group tRNA.PNG 1,181 × 745; 403 KB
- Tree.PNG 1,181 × 745; 403 KB
- Annotation.bmp 0 × 0; 389 KB
- Annotation.JPG 426 × 311; 30 KB
- 2CFS MG Active Site.jpg 638 × 478; 31 KB
- Adenine modified.jpg 659 × 783; 25 KB
- Reaction.png 866 × 783; 20 KB
- Clustalexample.bmp 366 × 270; 41 KB
- Conserved regions.JPG 420 × 475; 24 KB
- Tree in paint.jpg 0 × 0; 3.75 MB
- Tree in paint.bmp 0 × 0; 3.75 MB
- ASKsite.jpg 606 × 542; 203 KB
- ASAcat1.png 643 × 584; 196 KB
- ASAsite.png 607 × 540; 172 KB
- Trees.jpg 1,020 × 864; 69 KB
- Annotated image1.jpg 640 × 434; 60 KB
- Annotatedimage.jpg 640 × 434; 60 KB
- Uniprotsequence.jpg 791 × 432; 36 KB
- Uniprotname.jpg 1,051 × 437; 59 KB
- Uniprotontology.jpg 608 × 468; 34 KB
- Prositescan.png 477 × 599; 23 KB
- AlignmentC.png 809 × 522; 36 KB
- SBP with Sele Atom.png 640 × 480; 161 KB
- SBP with sele atom.png 640 × 480; 161 KB
- EMI.jpg 0 × 0; 879 KB
- INTER.jpg 593 × 333; 34 KB
- Hydrophobic.png 582 × 411; 82 KB
- Comparison 2ijz and 1vhe sequence.JPG 576 × 627; 107 KB
- Dali output2.jpg 721 × 345; 89 KB
- HSL thermophillic alignment.png 680 × 617; 60 KB
- Pyridoxal phosphatase Rectangular Phylogeny.pdf 0 × 0; 176 KB
- STSsite1.png 660 × 629; 190 KB
- ASKsite3.png 616 × 577; 225 KB
- Pyridoxal phosphatase BLAST.pdf 0 × 0; 250 KB
- Plasmodium.png 939 × 461; 202 KB
- IMG.jpg 1,613 × 1,063; 291 KB
- 2c7b alignment.png 1,049 × 595; 347 KB
- String2.jpg 828 × 400; 79 KB
- Blast.txt ; 1.03 MB
- Domain111.jpg 640 × 400; 12 KB
- Alignment.txt ; 2.67 MB
- FASTA.jpg 890 × 400; 94 KB
- Protdist1.txt ; 1.24 MB
- Neighbortree.txt ; 100 KB
- Align2.txt ; 2.89 MB
- Protdist2.txt ; 1.24 MB
- Neighbourtree2.txt ; 100 KB
- Expressionlev.jpg 640 × 724; 59 KB
- GROMOS tutorial.pdf 0 × 0; 569 KB
- AM 07 01.pdf 0 × 0; 723 KB
- AM 07 01.pdf $wgFileExtensions = .pdf 0 × 0; 723 KB
- Am 02 02.pdf 0 × 0; 239 KB
- AM 91 01.pdf 0 × 0; 228 KB
- 1RU8 align 2NZ2.png 705 × 472; 240 KB
- Clutsalw1.JPG 941 × 831; 132 KB
- Structural alignment.JPG 1,280 × 1,024; 234 KB
- MSA.png 1,265 × 636; 120 KB
- PDBsum chain.JPG 878 × 734; 105 KB
- 1ru8 electro surface.png 640 × 434; 175 KB
- Dalii.PNG 760 × 603; 63 KB
- Topograp.bmp 1,076 × 703; 2.16 MB
- PDBsum.bmp 720 × 661; 1.36 MB
- Sialidase.jpg 960 × 720; 111 KB
- Sialidase.png 960 × 720; 130 KB
- Sialidase2.png 576 × 396; 108 KB
- Galactose oxidase.png 576 × 396; 103 KB
- AATN domain ATPBD4.JPG 723 × 570; 69 KB
- HMM superfamily.JPG 822 × 816; 75 KB
- Profunc nests.JPG 651 × 614; 84 KB
- Profunc nests.jpg 651 × 614; 84 KB
- ProfuncNests.JPG 651 × 614; 235 KB
- PDBsum cleft.PNG 803 × 681; 157 KB
- PDBsum11.PNG 720 × 661; 90 KB
- Image of ERp16.png 640 × 434; 139 KB
- InterProScan result for 1tvg.bmp 748 × 559; 1.2 MB
- Interpro output.bmp 1,280 × 1,024; 3.75 MB
- Interpro.JPG 685 × 512; 44 KB
- Human Ssu72 BLAST result.bmp 652 × 367; 701 KB
- String1.JPG 770 × 513; 30 KB
- String1b.JPG 861 × 456; 75 KB
- Boostrap.PNG 591 × 1,320; 34 KB
- Clustalx.png 859 × 1,320; 140 KB
- Occurence.PNG 803 × 1,092; 57 KB
- Profunc, GO.png 960 × 720; 226 KB
- Profunc GO.jpg 960 × 720; 102 KB
- Blast Results.jpg 596 × 591; 78 KB
- Distance Matrix Calcuation.txt ; 16 KB
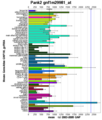
- Pfam result.jpg 960 × 720; 114 KB
- Blast hits.jpg 960 × 720; 139 KB
- Blast search vs uniprot.jpg 960 × 720; 99 KB
- PDB.jpg 763 × 305; 41 KB
- Image of ERp16 only active site no ions.png 640 × 434; 138 KB
- String result.jpg 960 × 720; 77 KB
- Image of ERp18 conserved secondary structure.png 640 × 434; 82 KB
- Image of ERp18 with conserved residues.png 640 × 434; 82 KB
- Image of CC bvbresidues.png 640 × 434; 258 KB
- NonredundantMSA.pdf 0 × 0; 454 KB
- TopNonredundantMSA.pdf 0 × 0; 350 KB
- SwissProtMSA.pdf 0 × 0; 61 KB
- BLAST10.png 1,262 × 737; 692 KB
- Tree final.png 1,273 × 866; 66 KB
- Pdb sum.png 864 × 648; 185 KB
- Pymol ss1.PNG 406 × 370; 59 KB
- Pymol hsp movie0001.png 580 × 482; 53 KB
- Geneologie2.png 667 × 665; 64 KB
- Tree cml.jpg 600 × 524; 26 KB
- Slide1.png 720 × 540; 94 KB
- Picture 2.png 514 × 407; 40 KB
- Fascin bootstrap animalia.insecta.jpg 933 × 845; 84 KB
- 1tvg castp.png 1,133 × 782; 228 KB
- 1tvg castp2.png 856 × 647; 66 KB
- 1tvg residue metal.png 640 × 434; 303 KB
- Nest analysis result.bmp 545 × 320; 511 KB
- Nest analysis result.PNG 545 × 320; 63 KB
- Msafinal.png 1,268 × 846; 716 KB
- Tree.bmp 1,270 × 861; 3.13 MB
- 1tvg castp2.PNG 794 × 548; 77 KB
- Nest.PNG 873 × 602; 79 KB
- 1tvg nests.png 448 × 304; 76 KB
- Profunc nest.bmp 622 × 195; 356 KB
- Dali.bmp 925 × 724; 1.92 MB
- Blue hills.jpg 800 × 600; 28 KB
- Sunset.jpg 800 × 600; 70 KB
- 2qmqA.png 620 × 438; 77 KB
- 2qmqm.png 640 × 434; 7 KB
- Getimg.gif 435 × 393; 23 KB
- Untitled.bmp 640 × 512; 960 KB
- Untitled3.PNG 850 × 800; 89 KB
- Untitled5.PNG 986 × 896; 169 KB
- Untitled11.bmp 306 × 216; 194 KB
- CASTp front.PNG 940 × 1,050; 132 KB
- Gene Ontology.bmp 380 × 86; 96 KB
- Gene ontology.PNG 755 × 400; 11 KB
- Hydrophobicity.png 888 × 637; 100 KB
- Hydrophobicity.png.jpeg 592 × 425; 40 KB
- Schematic.jpg 590 × 444; 35 KB
- SwissProtMSA2.pdf 0 × 0; 56 KB
- Clustal.jpg 394 × 329; 21 KB
- Fewerseqforalign.txt ; 7 KB
- 195.pdf 0 × 0; 242 KB
- Ba5125.pdf 0 × 0; 1.12 MB
- He5437.pdf 0 × 0; 473 KB
- Intropic.jpg 587 × 223; 19 KB
- SelNonredundantMSA.jpg 2,120 × 1,590; 1.45 MB
- Export.png 1,463 × 848; 40 KB
- Gurad final.bmp 718 × 285; 600 KB
- Gen posre.txt ; 2 KB
- Secondary structure.txt ; 559 bytes
- Zfs snapshots.pdf 0 × 0; 605 KB
- Guvench PLOSVomputBiol2009.pdf 0 × 0; 427 KB
- GPCR srt 1 day ProtSci09.pdf 0 × 0; 495 KB
- Group tasks.doc ; 46 KB
- DesaltTrypticPeptide.pdf 0 × 0; 45 KB
- InGelDigProtocol.pdf 0 × 0; 13 KB
- Guanidination.pdf 0 × 0; 83 KB
- 1.jpg 800 × 600; 118 KB
- 4720639103 a4e46f2f4d b.jpg 1,024 × 683; 247 KB
- 2hkz crystal.png 1,200 × 1,000; 379 KB
- 2hkz jacs.png 1,200 × 1,000; 359 KB
- Alpesh lychee.png 258 × 399; 130 KB
- UQ GPCR alpesh.png 400 × 800; 708 KB
- Background alpesh.png 2,400 × 2,400; 632 KB
- Atb.png 300 × 131; 24 KB
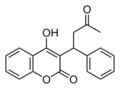
- Warfarin.png 220 × 163; 4 KB
- Warfarin 2.jpg 278 × 192; 20 KB
- Hessian.png 393 × 252; 5 KB
- SSP.png 1,216 × 445; 49 KB
- SAMPL.png 522 × 132; 49 KB
- Boron.png 424 × 600; 23 KB
- An2690.gif 70 × 52; 616 bytes
- Gila.jpg 500 × 424; 61 KB
- Byetta.jpg 320 × 240; 28 KB
- Test.jpg 1,003 × 1,728; 636 KB
- DSC 0127.jpeg 276 × 354; 85 KB
- DSC 0127.jpg 276 × 354; 85 KB
- Warfarin tautomers.png 333 × 255; 16 KB
- Artemisinin.png 300 × 300; 29 KB
- PICT1438.JPG 2,816 × 2,112; 1.11 MB
- PICT1438 copy.jpg 2,816 × 2,112; 1.11 MB
- P1010632.jpg 1,620 × 2,236; 1.37 MB
- Alpesh.JPG 768 × 768; 271 KB
- Drug Follow The Rules.pdf 0 × 0; 577 KB
- PsaA docked to E-cadherin in simulation cell.png 1,044 × 941; 739 KB
- Doxorubicin bound surface POPC CLR bilayer.png 719 × 721; 1 MB
- EPOR-EMP33 asu.png 1,181 × 816; 1.02 MB