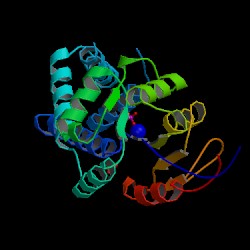
Structure of N-acetylneuraminic acid phosphatase
From MDWiki
Jump to navigationJump to search
CHAIN SEQUENCE
MGSDKIHHHHHHMGLSRVRAVFFDLDNTLIDTAGASRRGMLEVIKLLQSKYHYKEEAEIICDKVQVKLSKECFHPYSTCI TDVRTSHWEEAIQETKGGADNRKLAEECYFLWKSTRLQHMILADDVKAMLTELRKEVRLLLLTNGDRQTQREKIEACACQ SYFDAIVIGGEQKEEKPAPSIFYHCCDLLGVQPGDCVMVGDTLETDIQGGLNAGLKATVWINKSGRVPLTSSPMPHYMVS SVLELPALLQSIDCKVSMSV
DALI SEARCH RESULTS
SUMMARY: PDB/chain identifiers and structural alignment statistics NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3033-A 2gfh-A 41.1 0.0 246 246 100 0 0 1 S HYDROLASE haloacid dehalogenase-like hydrolase domain 2: 3033-A 1fez-A 18.1 3.5 178 256 22 0 0 13 S HYDROLASE phosphonoacetaldehyde hydrolase (bacillus c 3: 3033-A 2hsz-A 17.9 3.3 168 222 23 0 0 13 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION novel predicted 4: 3033-A 1qq5-A 17.3 3.1 198 245 19 0 0 12 S HYDROLASE l-2-haloacid dehalogenase (xanthobacter aut 5: 3033-A 1o03-A 17.0 5.0 188 221 20 0 0 11 S ISOMERASE beta-phosphoglucomutase (lactococcus lactis 6: 3033-A 2b0c-A 16.4 2.6 184 199 20 0 0 13 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION putative phospha 7: 3033-A 2fdr-A 15.8 4.4 190 214 19 0 0 15 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION conserved hypoth 8: 3033-A 2p11-A 15.7 2.9 194 211 16 0 0 20 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 9: 3033-A 1te2-A 15.7 3.6 170 211 19 0 0 15 S HYDROLASE putative phosphatase (escherichia coli o157 10: 3033-A 1yns-A 15.3 4.0 169 254 11 0 0 13 S HYDROLASE e-1 enzyme (enolase-phosphatase e1) (homo s 11: 3033-A 1qyi-A 15.0 3.5 198 375 19 0 0 17 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 12: 3033-A 2i6x-A 14.9 3.1 176 199 19 0 0 18 S HYDROLASE hydrolase, haloacid dehalogenase-like family 13: 3033-A 1u7p-A 14.3 2.9 144 164 18 0 0 14 S HYDROLASE magnesium-dependent phosphatase-1 (mdp-1) ( 14: 3033-A 1ymq-A 14.1 2.3 130 260 16 0 0 14 S TRANSFERASE sugar-phosphate phosphatase bt4131 (bacte 15: 3033-A 1j8d-A 13.1 2.5 141 180 11 0 0 12 S HYDROLASE deoxy-d-mannose-octulosonate 8-phosphate ph 16: 3033-A 2ho4-A 12.9 2.4 131 246 19 0 0 14 S HYDROLASE haloacid dehalogenase-like hydrolase domain 17: 3033-A 1pw5-A 12.7 2.3 136 246 21 0 0 12 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION nagd protein, pu 18: 3033-A 1nf2-A 12.7 2.6 127 267 13 0 0 11 S STRUCTURAL GENOMICS/UNKNOWN FUNCTION phosphatase (the 19: 3033-A 1rlm-A 12.4 2.8 131 269 13 0 0 14 S HYDROLASE phosphatase Mutant (escherichia coli) bacte 20: 3033-A 1f5s-A 12.1 3.5 159 210 14 0 0 15 S HYDROLASE phosphoserine phosphatase (psp) (methanoco 21: 3033-A 1cr6-B 12.0 3.8 177 541 18 0 0 18 S HYDROLASE epoxide hydrolase (mus musculus) mouse expr 22: 3033-A 1rku-A 11.9 3.6 172 206 11 0 0 18 S TRANSFERASE homoserine kinase (pseudomonas aeruginosa 23: 3033-A 2b30-A 11.8 2.7 134 284 16 0 0 12 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION pvivax hypotheti 24: 3033-A 1kyt-A 10.5 2.5 122 216 13 0 0 15 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 25: 3033-A 2o2x-A 10.3 3.6 139 204 17 0 0 14 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 26: 3033-A 1u02-A 10.1 2.7 128 222 16 0 0 12 S STRUCTURAL GENOMICS trehalose-6-phosphate phosphatase 27: 3033-A 2fea-A 10.0 3.5 167 219 7 0 0 21 S HYDROLASE 2-hydroxy-3-keto-5-methylthiopentenyl-1- pho 28: 3033-A 2hx1-A 9.6 3.2 130 275 24 0 0 19 S HYDROLASE predicted sugar phosphatases of the had supe 29: 3033-A 1mh9-A 9.2 3.2 146 194 15 0 0 15 S HYDROLASE deoxyribonucleotidase (mitochondrial 5'(3')-