Kenn: Difference between revisions
(PH1) |
No edit summary |
||
| Line 117: | Line 117: | ||
---- | ---- | ||
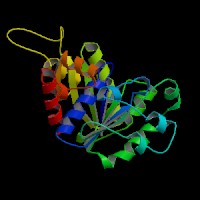
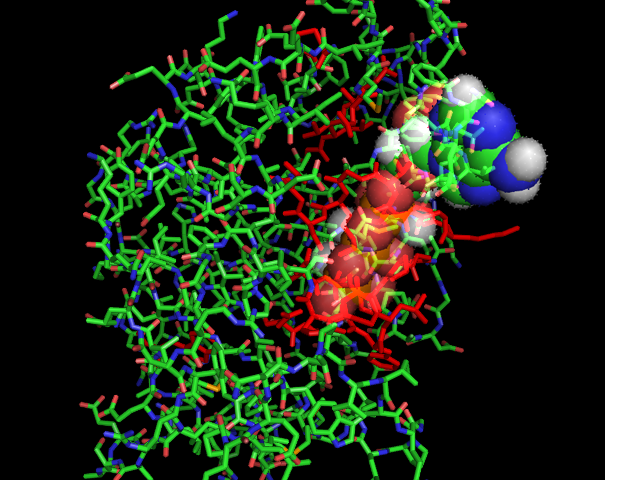
[[Image:ATPbinding.jpg|center|frame|ATP binding to 2ph1]] | [[Image:ATPbinding.jpg|center|frame|ATP binding to 2ph1]] | ||
The 2ph1 | The 2ph1 molecule with ATP binding | ||
---- | ---- | ||
Revision as of 01:22, 10 June 2008
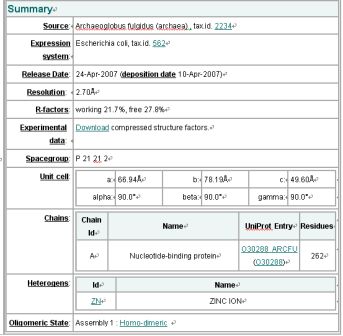
Structure Summary
NESG ID: GR165 PDB ID: 2ph1 Deposition date: 2007-04-10 Common Name: Nucleotide-binding protein Length (a.a.): 240 Organism: Archaeoglobus fulgidus SwissProt / TrEMBL ID: O30288_ARCFU Oligomerization: monomer Molecular weight: 26087
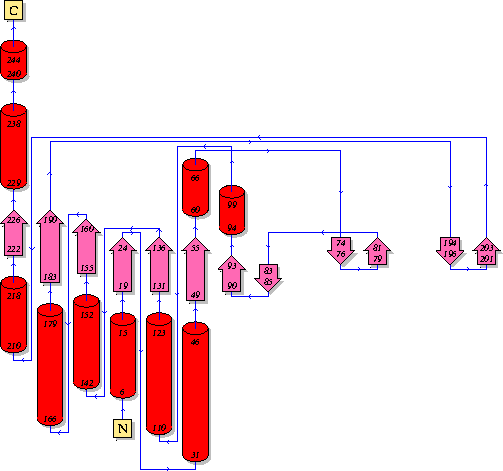
Secondary Structure Elements: alpha helices: 7-14, 32-45, 61-65, 111-122, 143-151, 167-178, 211-217, 230-237 beta strands: 83-84, 91-93, 50-54, 131-135, 20-24, 155-160, 184-189, 222-225, 74-76, 79-81, 194-195, 202-203
Resolution: 2.70 Å R-factor: 0.217 R-free: 0.278
EMI result
DaliLite: Structural Neighbours
Query: mol1A MOLECULE: NUCLEOTIDE-BINDING PROTEIN;
Matches are sorted by Z-score. Similarities with a Z-score lower than 2 are spurious.
Summary
No: Chain Z rmsd lali nres %id Description
1: 2ph1-A 47.3 0.0 247 247 100 MOLECULE: NUCLEOTIDE-BINDING PROTEIN;
2: 1g3q-A 20.0 2.7 207 237 21 MOLECULE: CELL DIVISION INHIBITOR;
3: 1ion-A 19.8 2.6 206 243 20 MOLECULE: PROBABLE CELL DIVISION INHIBITOR MIND;
4: 1hyq-A 19.6 3.1 197 232 25 MOLECULE: CELL DIVISION INHIBITOR (MIND-1);
5: 1g3r-A 19.6 2.7 206 237 20 MOLECULE: CELL DIVISION INHIBITOR;
6: 1cp2-A 19.0 2.8 204 269 16 MOLECULE: NITROGENASE IRON PROTEIN;
7: 1g1m-B 18.9 2.8 202 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
8: 1xd9-A 18.8 2.8 203 289 17 MOLECULE: NITROGENASE IRON PROTEIN 1;
9: 1cp2-B 18.8 2.7 203 269 16 MOLECULE: NITROGENASE IRON PROTEIN;
10: 1nip-B 18.4 2.8 202 287 18 NITROGENASE IRON PROTEIN
11: 1de0-A 18.4 2.8 203 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
12: 1xdb-A 18.3 2.9 204 289 18 MOLECULE: NITROGENASE IRON PROTEIN 1;
13: 1de0-B 18.3 2.8 203 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
14: 2bej-A 18.1 3.0 202 245 26 MOLECULE: SEGREGATION PROTEIN;
15: 1n2c-F 18.1 2.9 203 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
16: 1n2c-E 18.1 3.0 205 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
17: 1g5p-B 18.1 2.7 202 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
18: 1g5p-A 18.1 2.8 201 286 17 MOLECULE: NITROGENASE IRON PROTEIN;
19: 1g1m-A 18.1 2.8 202 287 18 MOLECULE: NITROGENASE IRON PROTEIN;
20: 2nip-B 18.0 2.7 201 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
21: 2nip-A 18.0 2.8 201 286 17 MOLECULE: NITROGENASE IRON PROTEIN;
22: 1nip-A 18.0 2.8 203 283 18 NITROGENASE IRON PROTEIN
23: 1n2c-G 18.0 2.9 203 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
24: 1m34-F 18.0 2.8 201 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN ALPHA CHAIN;
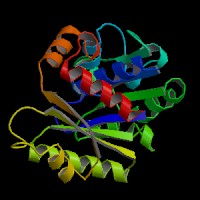
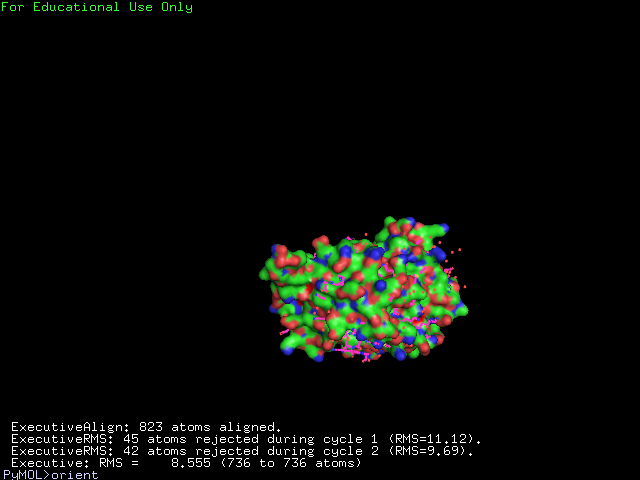
The structure of 1g3q is the most similar to 2ph1 according to the result from Dali.This figure shows the alignment of 2ph1 and 1g3q with RMS 0.55.
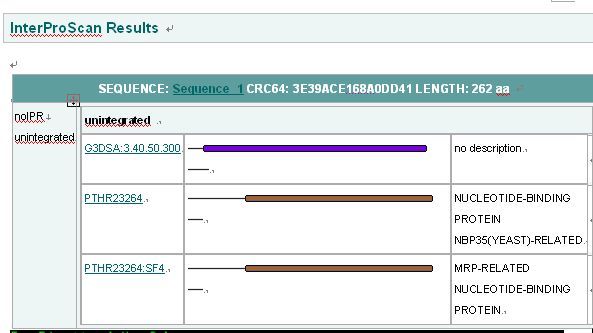
Interpro Result
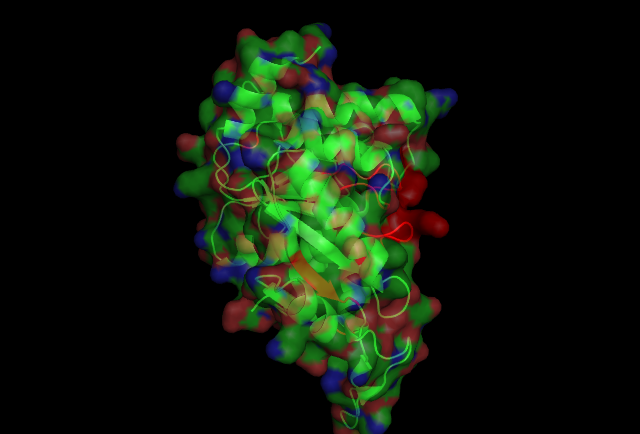
PUBSUM:
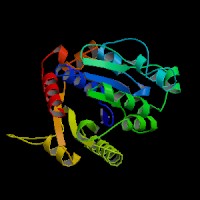
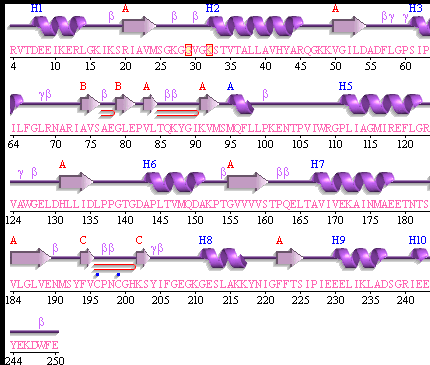
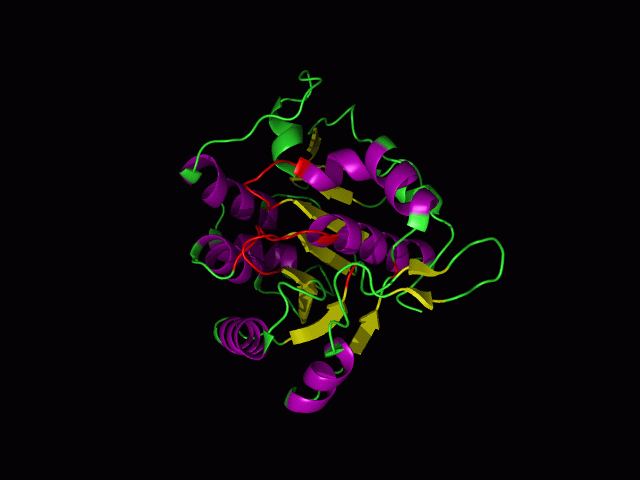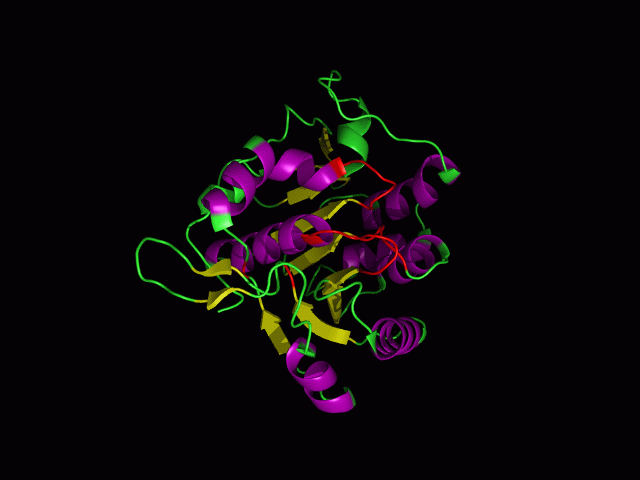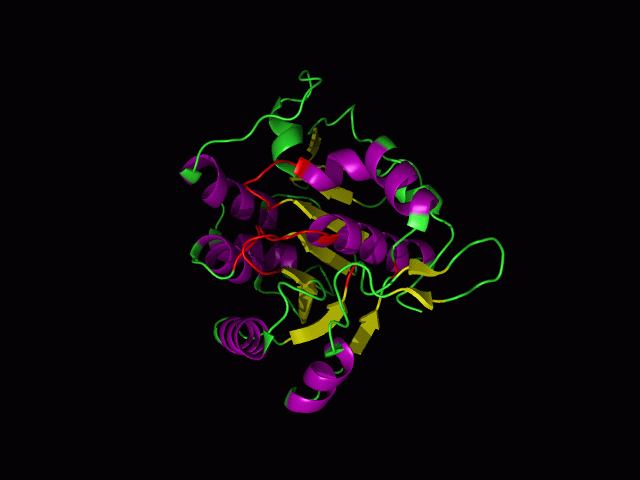
This picture shows the secondary structure of 2ph1, the alpha helixes were shown in purple, and the beta sheets are shown in yellow.
The 2ph1 molecule with ATP binding