ATP binding domain 4 Structures: Difference between revisions
Anharmustafa (talk | contribs) |
Anharmustafa (talk | contribs) |
||
| Line 81: | Line 81: | ||
'''ATP binding Domain 4 (1RU8) and Queuosine Biosynthesis (3BL5)''' | '''ATP binding Domain 4 (1RU8) and Queuosine Biosynthesis (3BL5)''' | ||
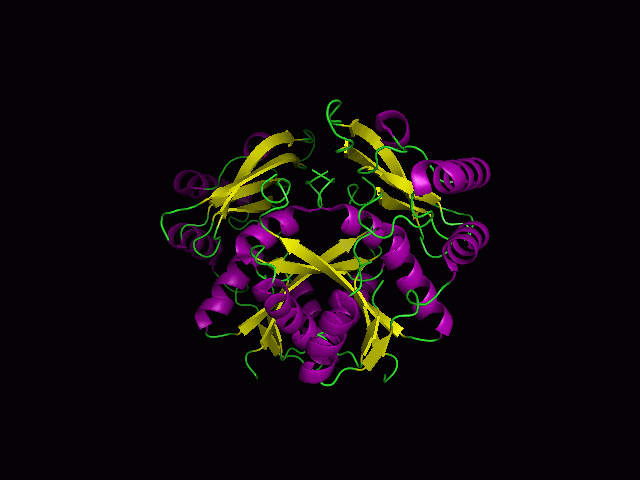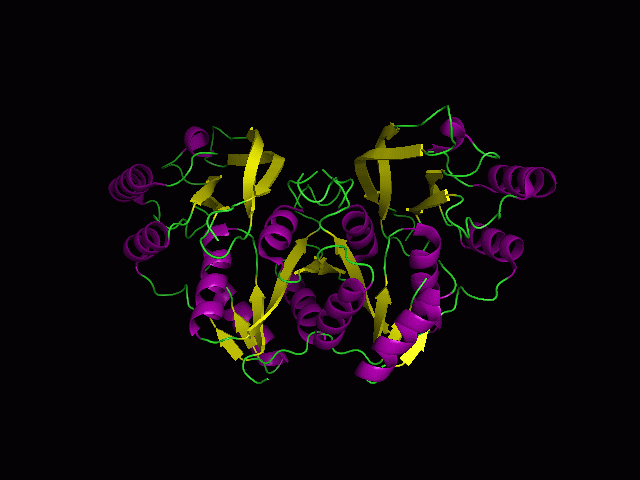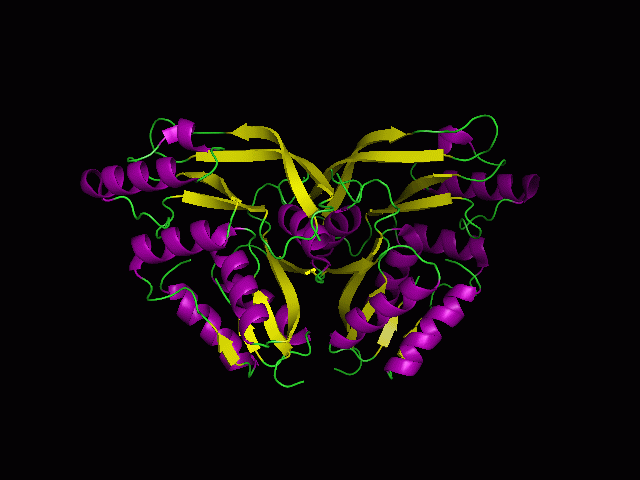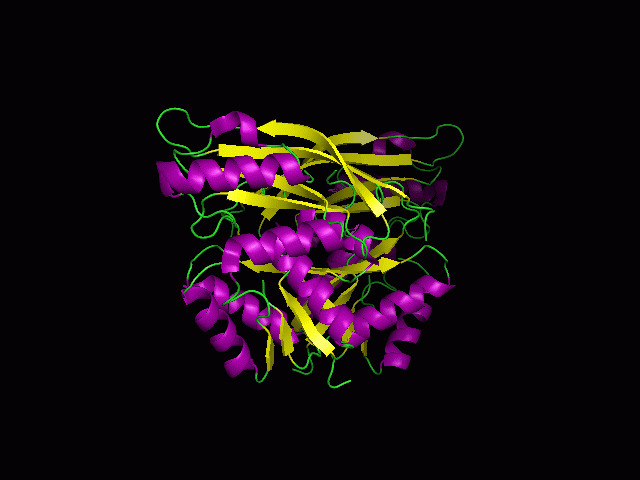
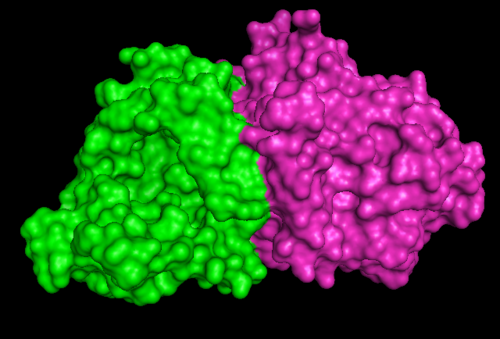
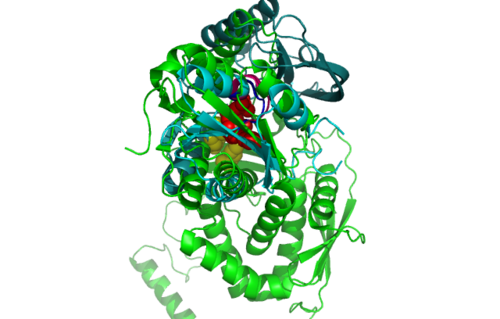
[[Image:1RU8 and 3BL5.png|left|thumb|500px|'''Figure 6.0'''. Structure alignment between 1RU8 and 3BL5 generated via PyMol]] | [[Image:1RU8 and 3BL5.png|left|thumb|500px|'''Figure 6.0'''. Structure alignment between 1RU8 (magenta) and 3BL5 (green) generated via PyMol. Highlighted colour of blue and red represent the conserved region of PP-loop.]] | ||
<BR><BR><BR><BR><BR><BR><BR><BR><BR><BR><BR><BR> | <BR><BR><BR><BR><BR><BR><BR><BR><BR><BR><BR><BR> | ||
| Line 91: | Line 91: | ||
<BR><BR> | <BR><BR> | ||
== Surface Cleft and Topography == | == Surface Cleft and Topography == | ||
Revision as of 16:56, 10 June 2009
General Properties
General information from PDB indicates that:
(a) 1RU8 is a putative n-type ATP pyrophosphatase isolated from Pyrococcus furiosus, expressed in Escherichia Coli.
(b) Is a member of clan PP-loop
(c) Resolution of 2.7 angstroms, with an r-value of 0.218.
(d) Ligand chemical component identified as TRS (2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL).
Information from SCOP when searched for 1RU8:
1. Root: scop
2. Class: Alpha and beta proteins (a/b) [51349]
Mainly parallel beta sheets (beta-alpha-beta units)
3. Fold: Adenine nucleotide alpha hydrolase-like [52373]
core: 3 layers, a/b/a ; parallel beta-sheet of 5 strands, order 32145
4. Superfamily: Adenine nucleotide alpha hydrolases-like [52402]
share similar mode of ligand (Adenosine group) binding
can be subdivided into two group with closer relationships within each group than between the groups; the first three families
form one group whereas the last two families form the other group
5. Family: N-type ATP pyrophosphatases [52403]
6. Protein: Putative N-type ATP pyrophosphatase PF0828 [102264]
7. Species: Archaeon Pyrococcus furiosus [TaxId: 2261] [102265]
Surface Structure
Pymol Visualization
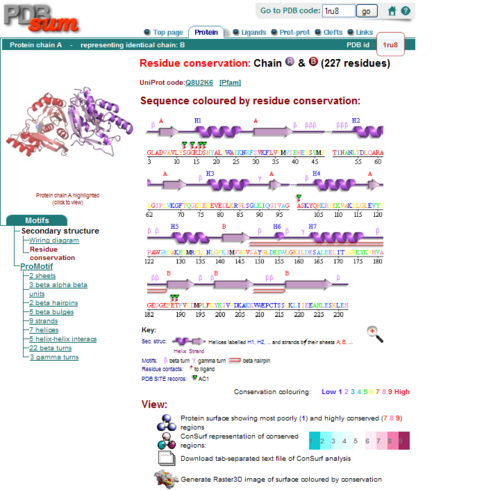
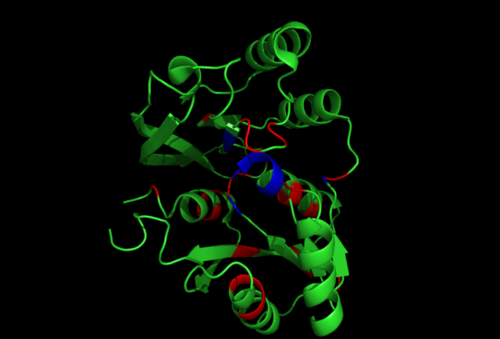
Secondary Structure and Location of PP-loop
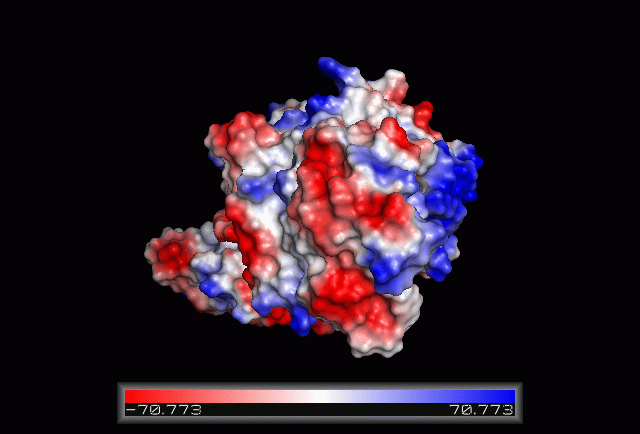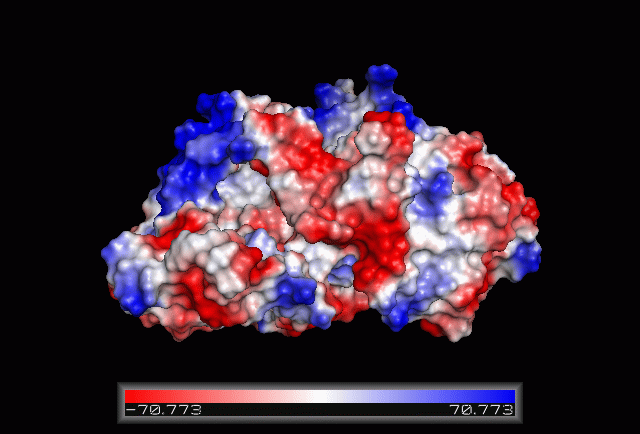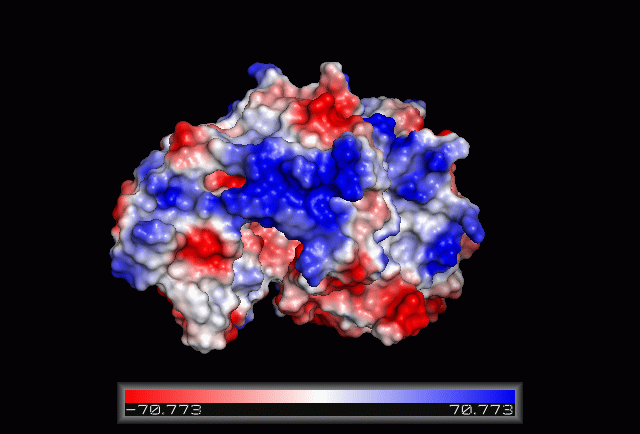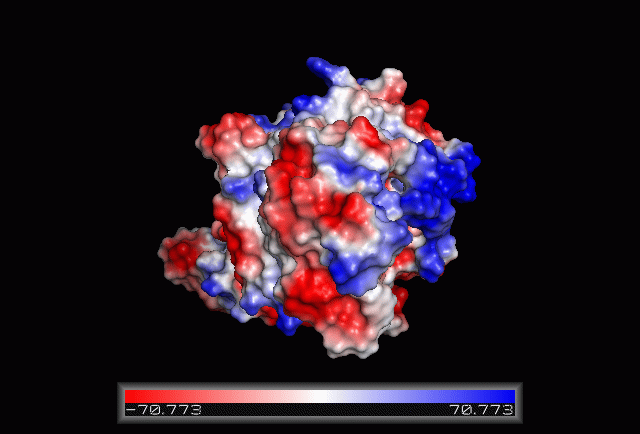
Electrostatic Surface Potential
Structure Similarities
Z score , the statistical significance of the similarity between protein-of-interest and other neighbourhood proteins. The program optimizes a weighted sum of similarities of intramolecular distances.
Root Mean Square Distance (RMSD), root-mean-square deviation of C-alpha atoms in the least-squares superimposition of the structurally equivalent C-alpha atoms. RMSD is not optimized and is only reported for information.
lali, the number of structurally equivalent residues.
nres, or the total number of amino acids in the hit protein.
%id, percentage of identical amino acids over structurally equivalent residues.
Structure Alignment
ATP binding Domain 4(1RU8) and Arginosuccinate Synthase (2NZ2)
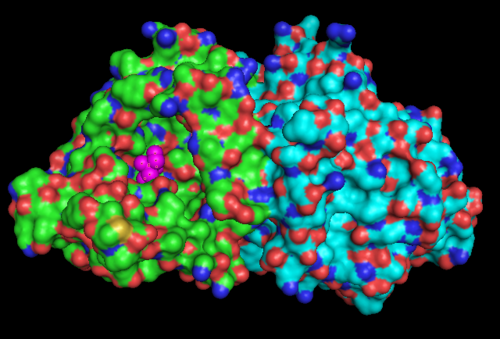
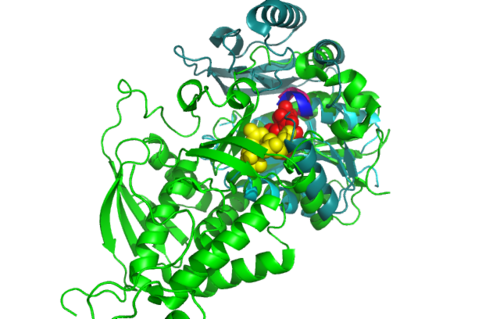
Highlighted in green is the 2NZ2 while in magenta is the 1RU8. Since 1RU8 has 2 domain, domain B is coloured darker in magenta for differentiation. There are 2 ligand indicated by the yellow (Citrulline) and red (ATP) spheres from 2NZ2. Highlighted also is the conserved region of PP-loop in pink and blue for 1RU8 and 2NZ2 respectively.
ATP binding Domain 4 (1RU8) and Queuosine Biosynthesis (3BL5)
Surface Cleft and Topography
The volume and size of the cleft of between protein indicated similarity between 1RU8 and 2NZ2. While 3BL5 has less similarity when compared to both 1RU8 and 2NZ2
Discussion
IRU8 is a protein with an unknown function. The crystallization of 1RU8 was done from Pyrococcus furiosus that is expressed in Escherichia Coli with resolution of 2.7 Angstroms and an R-value of 0.218. Artificial chemical ligand, TRS(2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL) was used in the crystallization in order to maintain the integrity of the structure when crystallized. There is no actual ligand of this protein based on the the literature paper and from the Protein Data Base (PDB). This is probably because the function of the protein is still unknown. Determination of the ligand can also leads to the possible function of the protein. Hence, deduction of the ligand of this protein can only be done, based on other protein's ligand that has similar function or structure.
From the structure obtained from PyMOL, visualization of the secondary structure provide information on the surface properties covering positively charged regions and negatively charged regions (figure 3.0), its domains (figure 2.0) as well as ligand binding sites and surface clefts (figure 7.0), and also conservation of residues across different species(figure 2.0). It was observed that detailed secondary structure can be determined using PDBsum where conserved residues, protein motif and possible ligand binding site was highlighted (Figure 2.0). Information from the PDB and SCOP has indicated that 1RU8 is from the family of N-type ATP pyrophosphatases and also under the clan of PP-loop which is the strongly conserved motif Ser(12),Gly(13),Gly(14),Lys(15) and Asp(16) at the N terminus (Bork & Koonin 1994). This PP-loop was significant as it is the most conserved and is involved in ligand bindings and thus is likely to give function to our protein. Besides that, from CATH domain database, two main domains were found in 1RU8. The first domain ranges from residue 3-97 while the second domain residues from 98-232. Topology of Domain 1 is known to be of Rossmann A-B-A fold, and the superfamily of PP-loop is presence in this domain. Domain 2 is A-B complex classified in CATH.
Structural alignment via DALI enabled us to obtained proteins that is similar in terms of structure relatedness to 1RU8. Based on the DALI output, 2d13 is rejected since it is a hypothetical protein ph1257 that has no known function to compare to. Thus, 2NZ2 and 3BL5 was chosen since it is the most similar based on the z-score (Figure 4.0). Information on 2NZ2 and 3BL5 by InterPro indicated that both protein has conserved motif of PP-loop at the N terminus which is similar to our protein. Hence, emphasizing again the importance of PP-loop to 1RU8's function. These 2 protein were analyzed based on the structural alignment and cleft size and volume. We inferred that the protein that has the most similar features to 1RU8 will probably gives similar function to 1RU8.
Structural alignment of 1RU8 with 2NZ2 and 3BL5 (Figure 5.0 and 6.0) show that 1RU8 and 2NZ2 has the most similar alignment compared to 1RU8 and 3BL5. Since 2NZ2 has a known function which is to catalyzes the citrulline and aspartate into argininosuccinate and pyrophosphate via hydrolysis of ATP, we inferred that the function of our protein may have similar mechanisms to 2NZ2 particularly the hydrolysis of ATP mechanisms. Based on Figure 5.0, we can observed that the ATP is located near the PP-loop highlighted in blue and Citrulline(yellow) is quite far from the loop which is supporting the fact that ATP is hydolysed for citrulline to activate. Hence, similar mechanism may be imply to 1RU8. Besides that, the active site or the binding site structure of both protein is quite similar(based on the structural alignment), and thus we also inferred that the substrate for 1RU8 must then have similar properties and organization of 2nz2 substrate (ATP and Citrulline) in a certain extent. However, the alignment is not convincingly similar due to domain B of 1RU8 (Figure 5.0) that probably is the distinction of differences in function to 2NZ2.
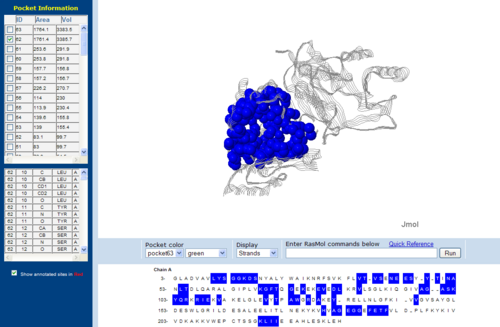
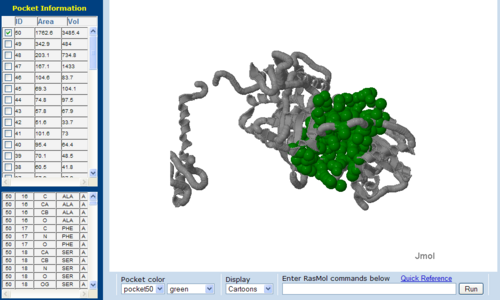
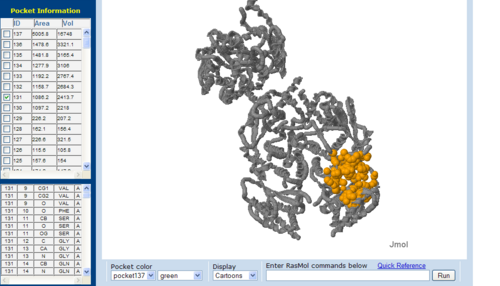
Pockets and cavities in the structure is often associated with binding sites and active sites of proteins. Moreover, it is also believed that there is high possibility that the largest cavity is the active site with some exceptions (Liang et al. 1998). Shape and size parameters of protein pockets and cavities are thus are important for active site analysis. Identification and measurements of surface accessible pockets as well as interior inaccessible cavities of 1RU8, 2NZ2 and 3BL5 were obtained from CASTp (Figure 7.0 to 7.2). It is inferred that the cleft volumes in proteins are related to their molecular interactions and functions (Laskowski et al. 1996). It was observed from the result, cleft of 1RU8 is quite similar in size and volume to 2NZ2 which suggesting that 1RU8 ligand may have similar size to 2NZ2 ligand. Thus, supporting the hypothesis that 1RU8 substrate or ligand may have similar properties to 2NZ2 substrate.
Despite the comparison of the two proteins to 1RU8, further structural comparison is needed in order for us to deduces other possible functions. Besides that determination of actual ligand based on experimental data may also help in finding the function of 1RU8.
Abstract|
Introductions|
Methods|
Structural Analysis|
Functional Analysis|
Evolutionary Analysis|
Discussions|
Conclusion |
References