Chromosome 1 open reading frame 41 Structure: Difference between revisions
From MDWiki
Jump to navigationJump to search
Hana Hamzah (talk | contribs) No edit summary |
Hana Hamzah (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
'''Protein Structure''' | '''Protein Structure''' | ||
Secondary structure | |||
[[Image:PDB sum 2.PNG|centre|framed|'''Figure 1:''' Representation of c1orf41 secondary structure as presented in PDBsum.]] | |||
[[Image:PDB sum 2.PNG | |||
Figure 1: | |||
Tertiary structure | Tertiary structure | ||
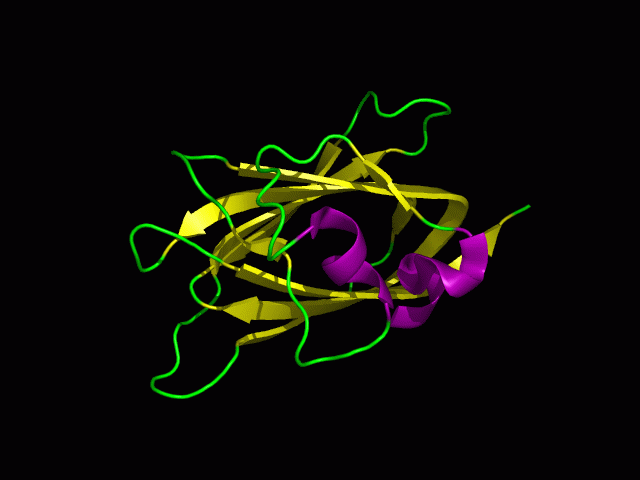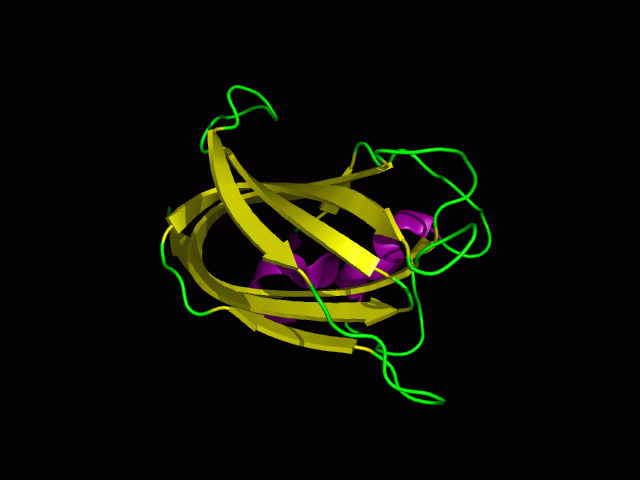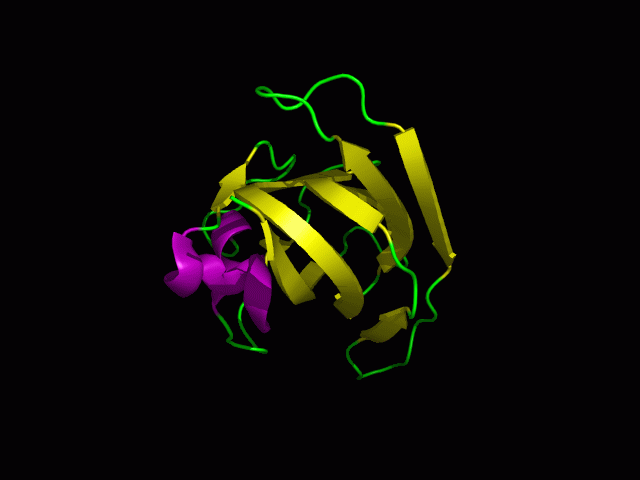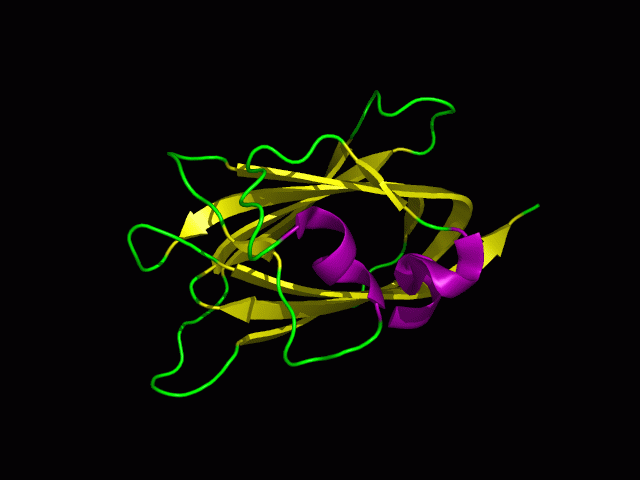
[[Image:1tvg ani.gif]] | [[Image:1tvg ani.gif|centre|framed|'''Figure 2:''' This protein is monomeric, consisting of a single domain. There are 8 beta strands (yellow) and 2 alpha helices (purple). The secondary structures are folded into jelly roll barrel where four pairs of anti-parallel beta strands are organised to form a barrel-like structure.]] | ||
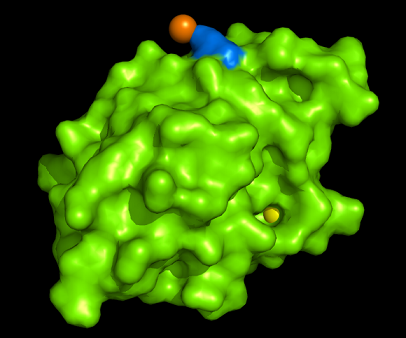
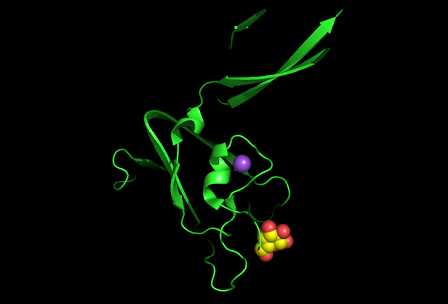
[[Image:1tvg surface.png|centre|framed|'''Figure 3:''' Surface representation of c1orf41. The Ca (II) ion is located within a pocket of the protein. Sm (III) interacts with Asp92 on the protein but its presence was probably due to the method used in solving the phase problem during structure solution using x-ray crystallogrophy.]] | |||
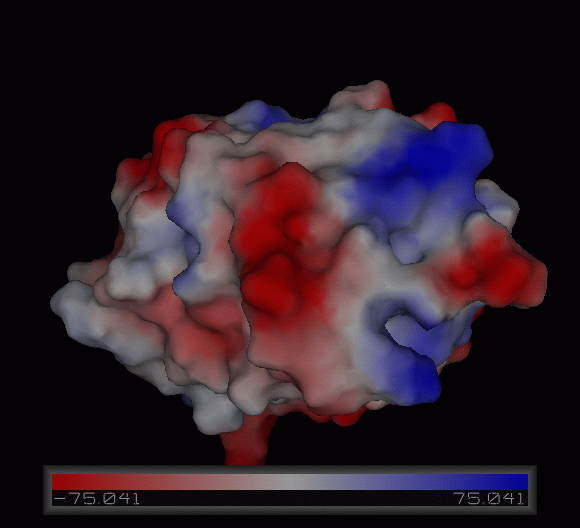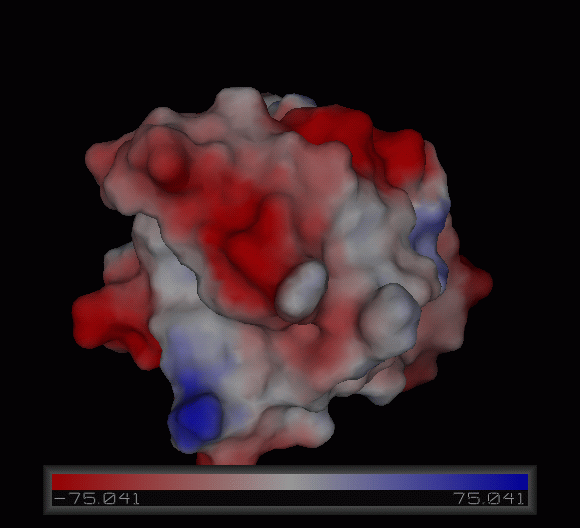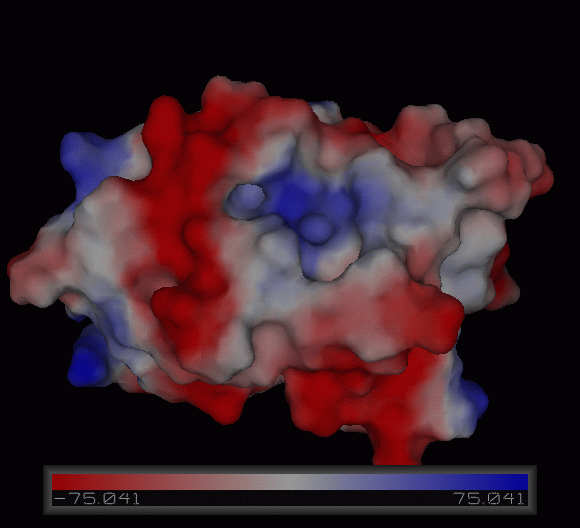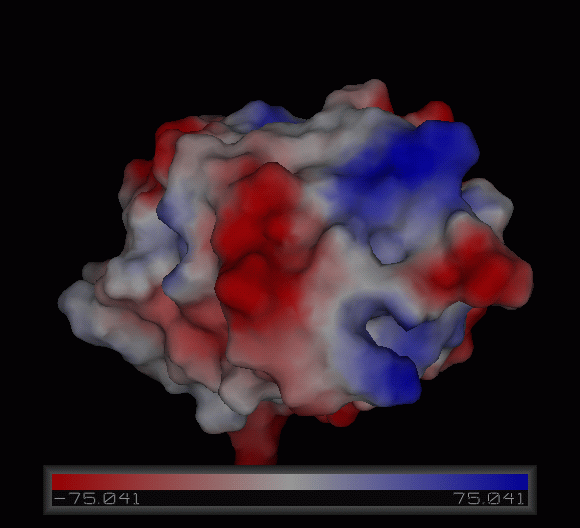
Figure | [[Image:Charge surface ani.gif|centre|framed|'''Figure 4:''' Electrostatic charge distribution on the surface of c1orf41.]] | ||
---- | ---- | ||
''' | '''Structural analysis''' | ||
'''1)Structural similarities''' | |||
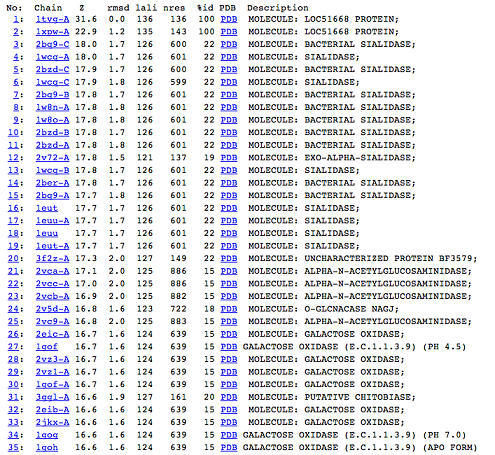
[[Image:Picture 1.png|centre|framed|'''Figure 5:''' DALI result showing the first 35 proteins that have similar structures to c1orf41. Even though the proteins are similar in structure they have very little sequence identity to c1orf41. 1xpw is c1orf41 but the structure was solved by NMR.]] | |||
[[Image:Picture 1.png | |||
Figure 5: DALI result showing the first 35 proteins that have similar structures to c1orf41. Even though the proteins are similar in structure they have very little sequence identity to c1orf41. 1xpw is c1orf41 but the structure was solved by NMR. | |||
'''2)Domain Classification''' | '''2)Domain Classification''' | ||
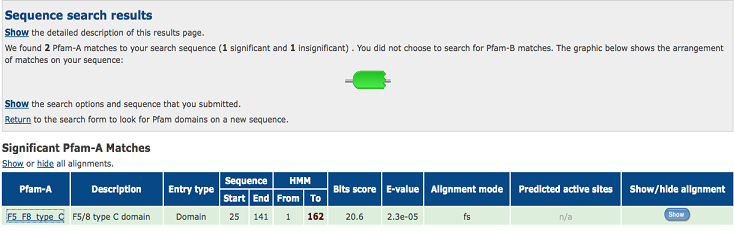
[[Image:1TVG pfam.png | [[Image:1TVG pfam.png|centre|framed|'''Figure 6:''' Pfam result indicated that our protein has a F5/8 type C domain, also known as the discoidin domain which is apart of galactose binding domain super family.]] | ||
Figure 6:Pfam result indicated that our protein has a F5/8 type C domain, also known as the discoidin domain which is apart of galactose binding domain super family. | |||
'''3)Possible ligand binding sites''' | '''3)Possible ligand binding sites''' | ||
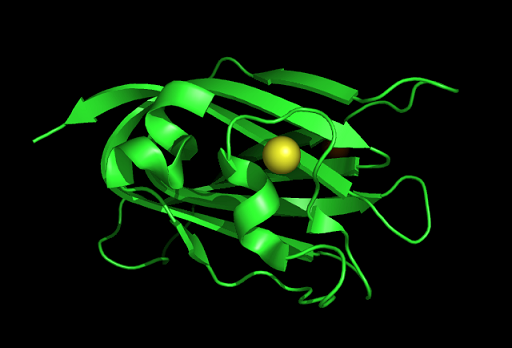
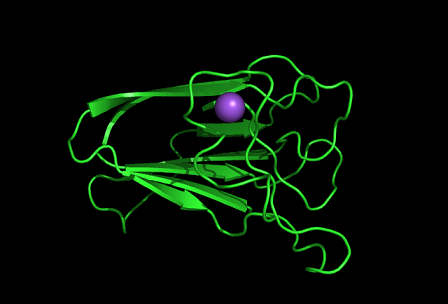
[[Image:1tvg | [[Image:1tvg ligand1.png|centre|framed|'''Figure 7:''' This figure shows the position where calcium (II) ion (yellow) is located in within a loop of c1orf41 structure. This loop where metal ion is coordinated to was also observed in several proteins from the DALI result.]] | ||
[[Image:1gog Na.png|centre|framed|]] | |||
[[Image: | [[Image:2bzd discoidin2.png|centre|framed|'''Figure 8:'''Shown at the top is the discoidin domain of galactose oxidase with a sodium ion (purple) located within a loop. The bottom figure is bacterial sialidase discoidin domain with a sodium ion (purple) and a beta-D-galactose molecule. The sodium ion is also located i within a loop.]] | ||
[[Image:1tvg castp3.PNG|centre|framed|'''Figure 9:'''CastP identified a surface cleft that is located at the loop position.]] | |||
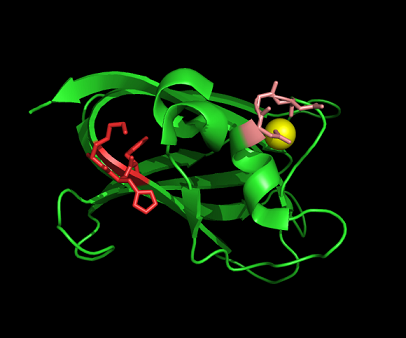
[[Image: | [[Image:1tvg ca res2.PNG|centre|framed|'''Figure 10:'''Pymol figure displaying possible metal binding position on c1orf41. Pink are the residues that may be involved in coordinating the metal and yellow is the calcium ion. The residues are based on CastP result. The aspartate and glutamate residues are likely to be important in binding positively charged metal ion]] | ||
[[Image: | [[Image:Profunc nest.PNG|centre|framed|'''Figure 11:'''Nest 1 and 2 had scores higher than 2. So, they are more likely to be functional. His54 and Lys55 NH atoms are accessible from a large surface cleft of the protein. The cleft is also deep indicating that the nest is functionally important. The residues of Nest 2 are part of the loop that coordinates the Ca (II)ion.]] | ||
Figure 12: Pymol representation of Nest 1 and 2. | [[Image:1tvg nests2.png|centre|framed|'''Figure 12:'''Pymol representation of Nest 1 and 2.]] | ||
[[Image:Surface nest3.PNG]] | [[Image:Surface nest3.PNG|centre|framed|'''Figure 13:'''Arrows indicating the pockets of the nests that may be accessible to ligands.]] | ||
[[Chromosome 1 open reading frame 41|Back to main page]] | [[Chromosome 1 open reading frame 41|Back to main page]] | ||
Revision as of 15:46, 12 June 2009
Protein Structure
Secondary structure
Tertiary structure
Structural analysis
1)Structural similarities
2)Domain Classification
3)Possible ligand binding sites
Figure 10:Pymol figure displaying possible metal binding position on c1orf41. Pink are the residues that may be involved in coordinating the metal and yellow is the calcium ion. The residues are based on CastP result. The aspartate and glutamate residues are likely to be important in binding positively charged metal ion
Figure 11:Nest 1 and 2 had scores higher than 2. So, they are more likely to be functional. His54 and Lys55 NH atoms are accessible from a large surface cleft of the protein. The cleft is also deep indicating that the nest is functionally important. The residues of Nest 2 are part of the loop that coordinates the Ca (II)ion.