Structure Protein: Difference between revisions
| Line 38: | Line 38: | ||
Dark colors: 2ifu:A | Dark colors: 2ifu:A | ||
Light colors: 1qqe:A | Light colors: 1qqe:A | ||
=== SCOP === | |||
multiple alignment results, [http://www.ebi.ac.uk/msd-srv/ssm/cgi-bin/ssmserver click here] | |||
== Protein families == | == Protein families == | ||
Revision as of 08:30, 4 June 2007
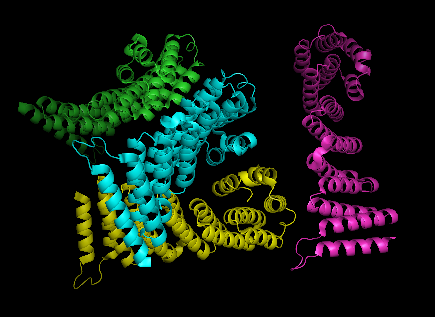
2ifu has 4 chains (A, B, C and D) integrated together to form the whole molecular structure as shown below:
(chain A= green, chain B= cyan, chain c= purple, and chain d= yellow)
ligands: sulfate ion (SO4) and selenomethionine (MSE)
Ten sulfate ions were observed. Each ion differs in arrangement as well as orientation related with each chains.
protein sequence (FASTA)
>gi|118138356|pdb|2IFU|A Chain A, Crystal Structure Of A Gamma-Snap From Danio Rerio AIAAQKISEAHEHIAKAEKYLKTSFXKWKPDYDSAASEYAKAAVAFKNAKQLEQAKDAYLQEAEAHANNR SLFHAAKAFEQAGXXLKDLQRXPEAVQYIEKASVXYVENGTPDTAAXALDRAGKLXEPLDLSKAVHLYQQ AAAVFENEERLRQAAELIGKASRLLVRQQKFDEAAASLQKEKSXYKEXENYPTCYKKCIAQVLVQLHRAD YVAAQKCVRESYSIPGFSGSEDCAALEDLLQAYDEQDEEQLLRVCRSPLVTYXDNDYAKLAISLKVPGGG GGKKKPSASASAQPQEEEDDEYAGGLC
Strutural comparisons
Based on Dali search, 6 proteins with highest Z-value were chosen: 1qqe-A (protein transport), 2fi7-A (protein binding), 1a17 (hydrolase), 2ak6-A, 1fch-A (signalling protein), 1haz4-A (transcription factor),
snap-gamma: multiple structure alignment
Snap-gamma: sequence alignment with 1QQE:A
Dark colors: 2ifu:A
Light colors: 1qqe:A
SCOP
multiple alignment results, click here
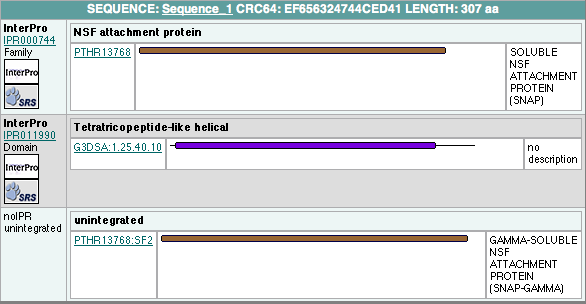
Protein families
has 2 domains: Pfam at Sanger
alignment with Pfam-B_15198
highly conserved residue :residue 104, 109, 121, 168
scorecons result indicated a Diversity of position scores: 89.4% where 100%=high diversity