Evolutionary analysis of 2ece: Difference between revisions
No edit summary |
No edit summary |
||
| Line 13: | Line 13: | ||
[[Image:gitable.png]] | [[Image:gitable.png]] | ||
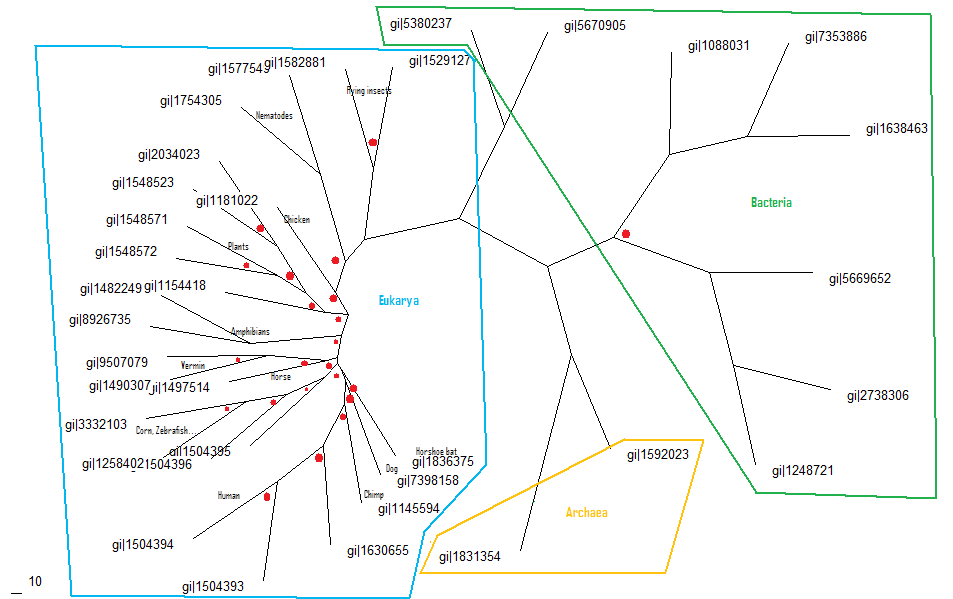
The shown alignment was prepared into a phylogenic tree, shown below, using the nieghbourhood joining method, then creating a consensus tree with bootstrap values. The three kingdoms are circled and labelled and nodes with bootstrap values of less than 75% shown with red dots. | |||
[[Image:boottree.png]] | |||
Revision as of 13:59, 9 June 2008
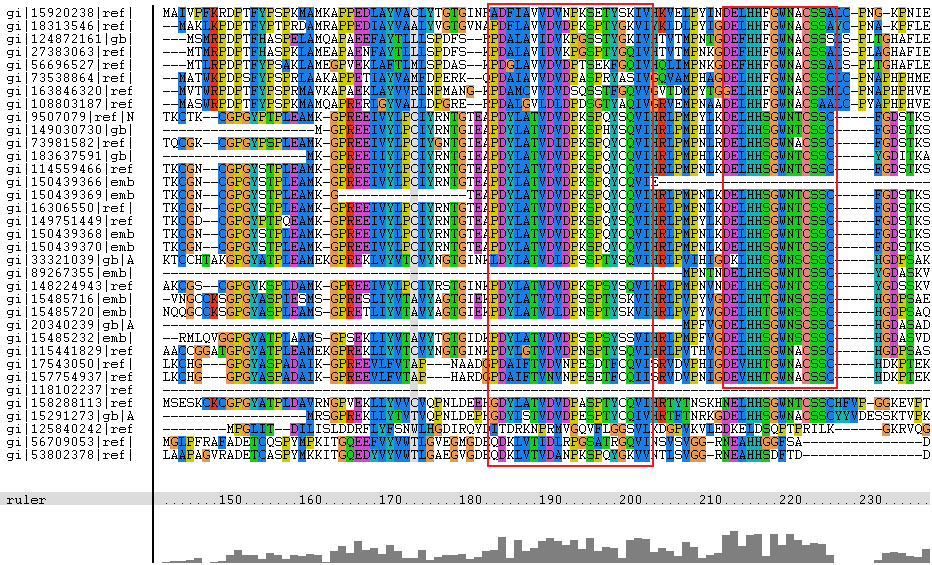
The amino acid sequence of selenium binding protein 2ece, sourced from the organism Sulfolobus tokodaii str. 7, was compared with other proteins using a psi blast(Media:psiblast.txt), with hits from the results hand selected (to show biological diversity) to go through a multiple sequence alignment in clustalx with portions of the alignment shown below
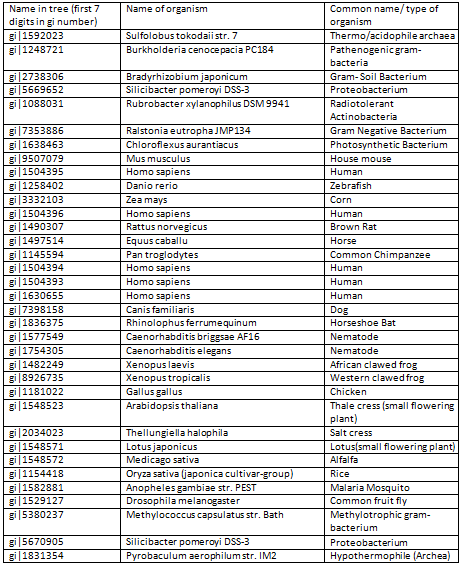
The segments in red boxes are reasonably highly conserved amongst the organism, and the segment in a brown box apperas to be a common insertion shared between some organisms. The protein sequences are labelled with gi numbers, the table below associates organisms to specific gi numbers.
The shown alignment was prepared into a phylogenic tree, shown below, using the nieghbourhood joining method, then creating a consensus tree with bootstrap values. The three kingdoms are circled and labelled and nodes with bootstrap values of less than 75% shown with red dots.