Kenn: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{| | |||
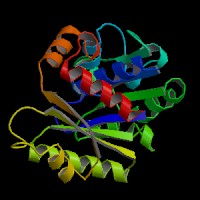
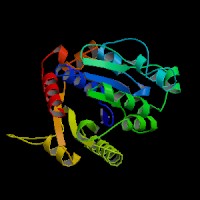
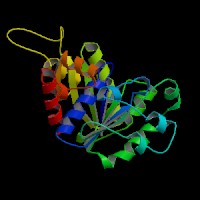
|[[Image:NUBP2BACK.jpg|left|frame|NUBP2 Front view]] | |||
|[[Image:nubp2front.jpg|right|frame|NUBP2 Top view]] | |||
|[[Image:nubp2top.jpg|none|frame|NUBP2 Back view]] | |||
|} | |||
== '''Structure Summary'''== | == '''Structure Summary'''== | ||
| Line 18: | Line 23: | ||
---- | ---- | ||
''EMI result'' | |||
[[Image:EMIi.jpg|center]] | |||
---- | |||
'''DaliLite: Structural Neighbours''' | '''DaliLite: Structural Neighbours''' | ||
| Line 78: | Line 92: | ||
---- | ---- | ||
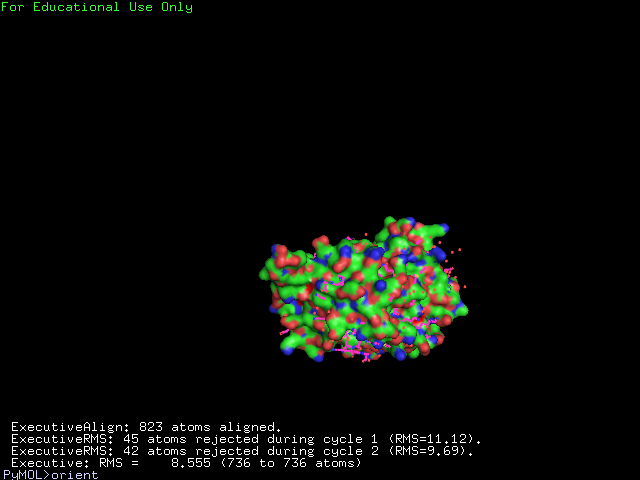
[[Image:alignment1g3p.jpg|400px|center|frame|The alignment of 2ph1 and 1g3q]] | |||
The structure of 1g3q is the most similar to 2ph1 according to the result from Dali.This figure shows the alignment of 2ph1 and 1g3q with RMS 0.55. | |||
[[Image: | |||
---- | ---- | ||
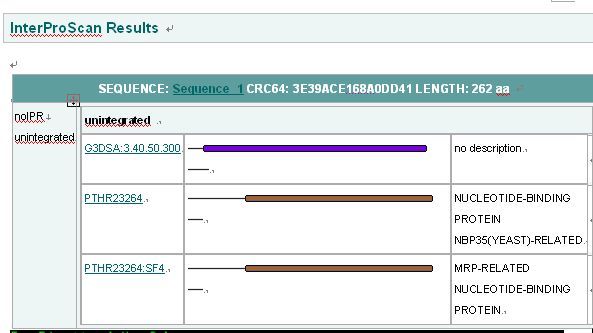
''Interpro Result'' | ''Interpro Result'' | ||
[[Image:INTERPRO.jpg|center]] | [[Image:INTERPRO.jpg|center]] | ||
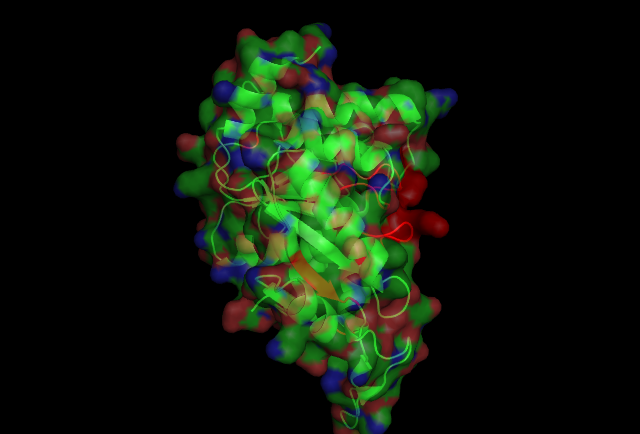
[[Image:2ph1 surface.jpg|400px|frame|The P-loop connecting | [[Image:2ph1 surface.jpg|center|400px|frame|The P-loop connecting | ||
b1 and a1, has well conserved sequences, and its amino | b1 and a1, has well conserved sequences, and its amino | ||
acid side chains exhibit the same conformations among all | acid side chains exhibit the same conformations among all | ||
| Line 95: | Line 108: | ||
---- | ---- | ||
PUBSUM: | |||
[[Image:PUBSUM.jpg]] | |||
[[Image: | [[Image:SECONDARY.jpg]] | ||
[[Image:SURFACE.jpg]] | |||
[[Image: | |||
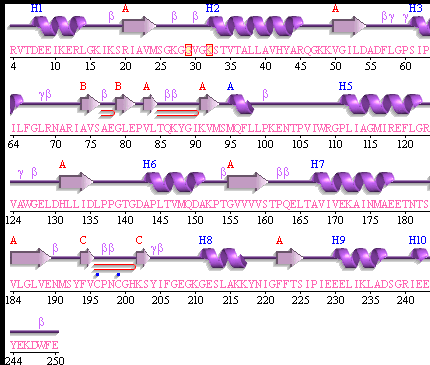
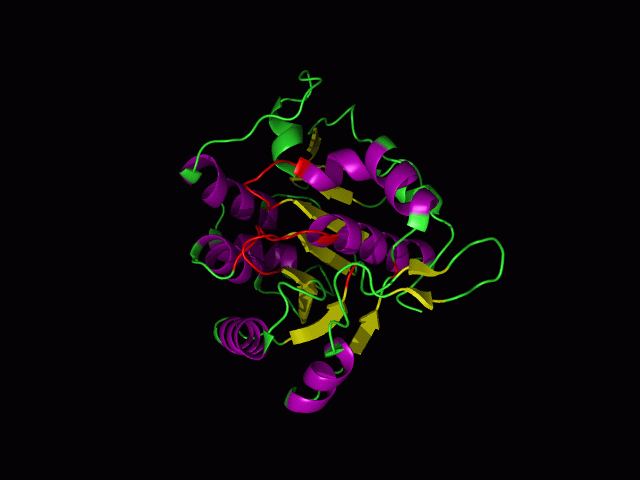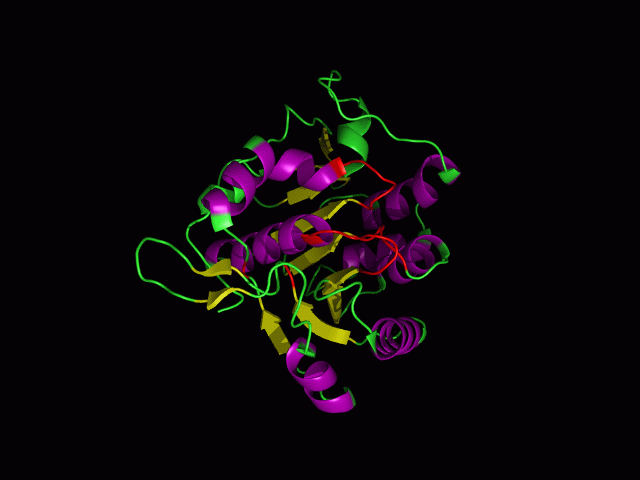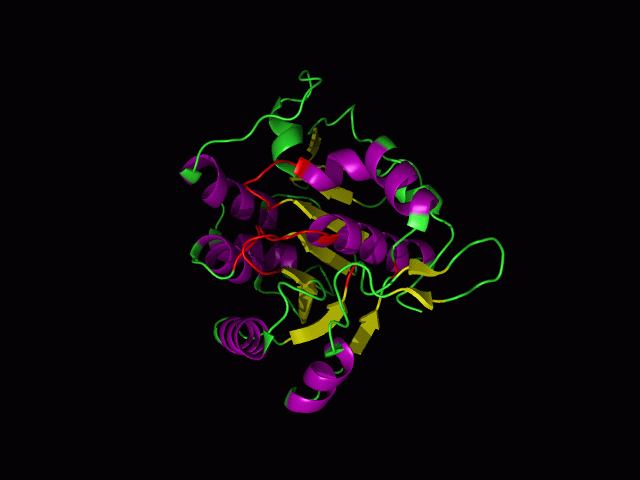
[[Image:movie.jpg|center|frame|The secondary structure of 2ph1]] | [[Image:movie.jpg|center|frame|The secondary structure of 2ph1]] | ||
This picture shows the secondary structure of 2ph1, the alpha helixes were shown in purple, and the beta sheets are shown in yellow. | This picture shows the secondary structure of 2ph1, the alpha helixes were shown in purple, and the beta sheets are shown in yellow. | ||
| Line 112: | Line 118: | ||
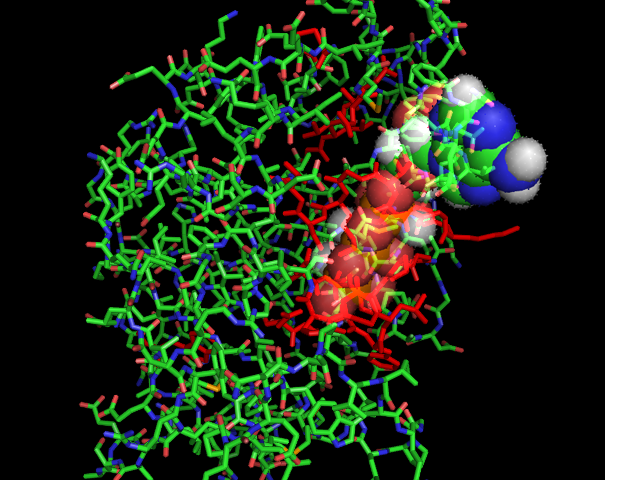
[[Image:ATPbinding.jpg|center|frame|ATP binding to 2ph1]] | [[Image:ATPbinding.jpg|center|frame|ATP binding to 2ph1]] | ||
The ATP molocule was binding to the p-loop of 2ph1. | The ATP molocule was binding to the p-loop of 2ph1. | ||
---- | ---- | ||
Revision as of 00:55, 10 June 2008
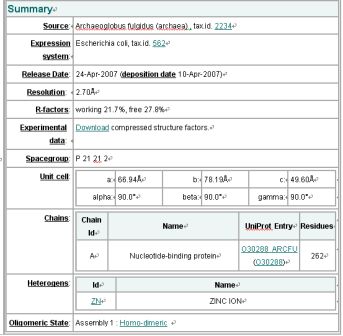
Structure Summary
NESG ID: GR165 PDB ID: 2ph1 Deposition date: 2007-04-10 Common Name: Nucleotide-binding protein Length (a.a.): 240 Organism: Archaeoglobus fulgidus SwissProt / TrEMBL ID: O30288_ARCFU Oligomerization: monomer Molecular weight: 26087
Secondary Structure Elements: alpha helices: 7-14, 32-45, 61-65, 111-122, 143-151, 167-178, 211-217, 230-237 beta strands: 83-84, 91-93, 50-54, 131-135, 20-24, 155-160, 184-189, 222-225, 74-76, 79-81, 194-195, 202-203
Resolution: 2.70 Å R-factor: 0.217 R-free: 0.278
EMI result
DaliLite: Structural Neighbours
Query: mol1A MOLECULE: NUCLEOTIDE-BINDING PROTEIN;
Matches are sorted by Z-score. Similarities with a Z-score lower than 2 are spurious.
Summary
No: Chain Z rmsd lali nres %id Description
1: 2ph1-A 47.3 0.0 247 247 100 MOLECULE: NUCLEOTIDE-BINDING PROTEIN;
2: 1g3q-A 20.0 2.7 207 237 21 MOLECULE: CELL DIVISION INHIBITOR;
3: 1ion-A 19.8 2.6 206 243 20 MOLECULE: PROBABLE CELL DIVISION INHIBITOR MIND;
4: 1hyq-A 19.6 3.1 197 232 25 MOLECULE: CELL DIVISION INHIBITOR (MIND-1);
5: 1g3r-A 19.6 2.7 206 237 20 MOLECULE: CELL DIVISION INHIBITOR;
6: 1cp2-A 19.0 2.8 204 269 16 MOLECULE: NITROGENASE IRON PROTEIN;
7: 1g1m-B 18.9 2.8 202 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
8: 1xd9-A 18.8 2.8 203 289 17 MOLECULE: NITROGENASE IRON PROTEIN 1;
9: 1cp2-B 18.8 2.7 203 269 16 MOLECULE: NITROGENASE IRON PROTEIN;
10: 1nip-B 18.4 2.8 202 287 18 NITROGENASE IRON PROTEIN
11: 1de0-A 18.4 2.8 203 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
12: 1xdb-A 18.3 2.9 204 289 18 MOLECULE: NITROGENASE IRON PROTEIN 1;
13: 1de0-B 18.3 2.8 203 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
14: 2bej-A 18.1 3.0 202 245 26 MOLECULE: SEGREGATION PROTEIN;
15: 1n2c-F 18.1 2.9 203 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
16: 1n2c-E 18.1 3.0 205 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
17: 1g5p-B 18.1 2.7 202 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
18: 1g5p-A 18.1 2.8 201 286 17 MOLECULE: NITROGENASE IRON PROTEIN;
19: 1g1m-A 18.1 2.8 202 287 18 MOLECULE: NITROGENASE IRON PROTEIN;
20: 2nip-B 18.0 2.7 201 289 18 MOLECULE: NITROGENASE IRON PROTEIN;
21: 2nip-A 18.0 2.8 201 286 17 MOLECULE: NITROGENASE IRON PROTEIN;
22: 1nip-A 18.0 2.8 203 283 18 NITROGENASE IRON PROTEIN
23: 1n2c-G 18.0 2.9 203 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN;
24: 1m34-F 18.0 2.8 201 274 18 MOLECULE: NITROGENASE MOLYBDENUM-IRON PROTEIN ALPHA CHAIN;
The structure of 1g3q is the most similar to 2ph1 according to the result from Dali.This figure shows the alignment of 2ph1 and 1g3q with RMS 0.55.
Interpro Result
This picture shows the secondary structure of 2ph1, the alpha helixes were shown in purple, and the beta sheets are shown in yellow.
The ATP molocule was binding to the p-loop of 2ph1.