Chromosome 1 open reading frame 41 Structure: Difference between revisions
From MDWiki
Jump to navigationJump to search
Hana Hamzah (talk | contribs) No edit summary |
Hana Hamzah (talk | contribs) No edit summary |
||
| Line 10: | Line 10: | ||
This protein's secondary structure consists of 2 alpha helices and 9 beta sheets. | This protein's secondary structure consists of 2 alpha helices and 9 beta sheets. | ||
[[Image:Pymol ss1.PNG]] | |||
Pymol structure displaying the folding of the secondary structure. Red is alpha helices, yellow is beta sheets and green is random coils connecting the the sheets and helices. | |||
Revision as of 04:58, 7 June 2009
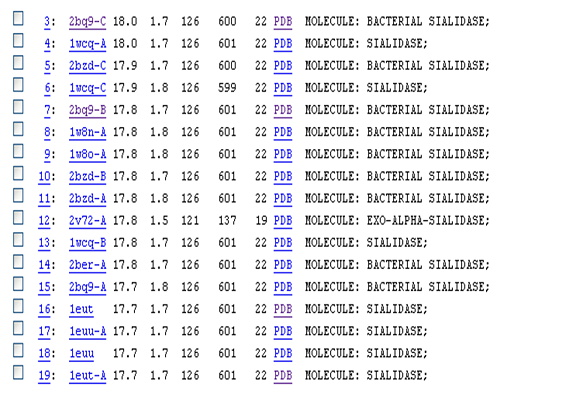
Based on the DALI result, sialidases have similar structures to our protein. However, the sequence identity between our protein and sialidase proteins are small (~22). Sialidases are enzymes that cleave sialic acids from various glyco-conjugates (Newsted et al., 2005).
Another enzyme that our protein is similar in structure to is galactose oxidase. This enzyme catalyzes the oxidation of D-galactose to D-galacto-hexodialdose and H2O2 (Ito et al., 1991).
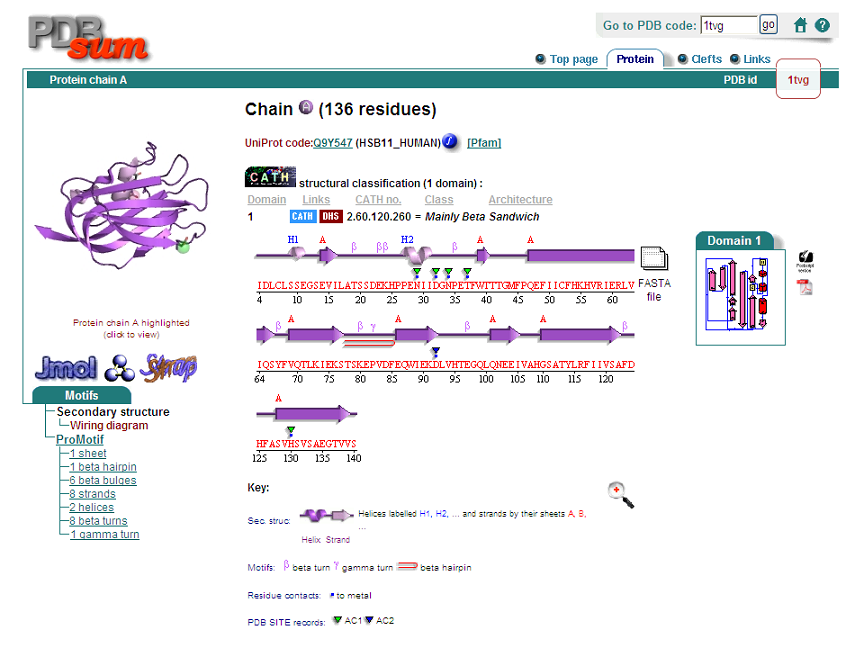
This protein's secondary structure consists of 2 alpha helices and 9 beta sheets.
Pymol structure displaying the folding of the secondary structure. Red is alpha helices, yellow is beta sheets and green is random coils connecting the the sheets and helices.