Chromosome 1 open reading frame 41 Structure: Difference between revisions
Hana Hamzah (talk | contribs) No edit summary |
Hana Hamzah (talk | contribs) No edit summary |
||
| Line 12: | Line 12: | ||
Structure analysis | Structure analysis | ||
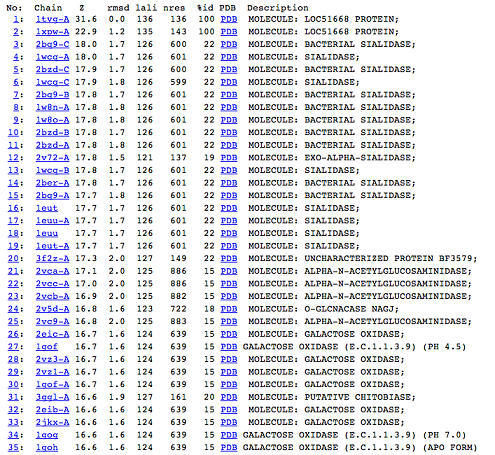
DALI was used to find protein that are similar in structure to 1TVG. The results showed that sialidases, alpha-N-acetylglucosaminidases and galactose oxidases have similar structure to our protein as presented below. | |||
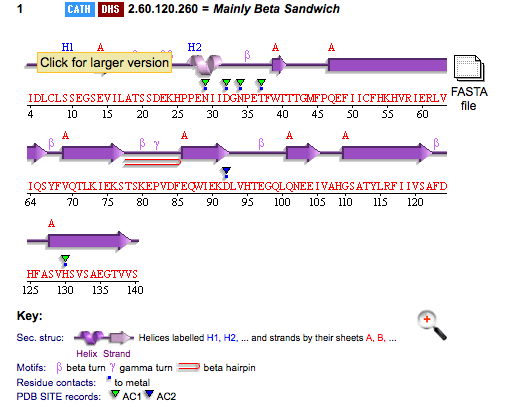
[[Image:Picture 1.png]] | |||
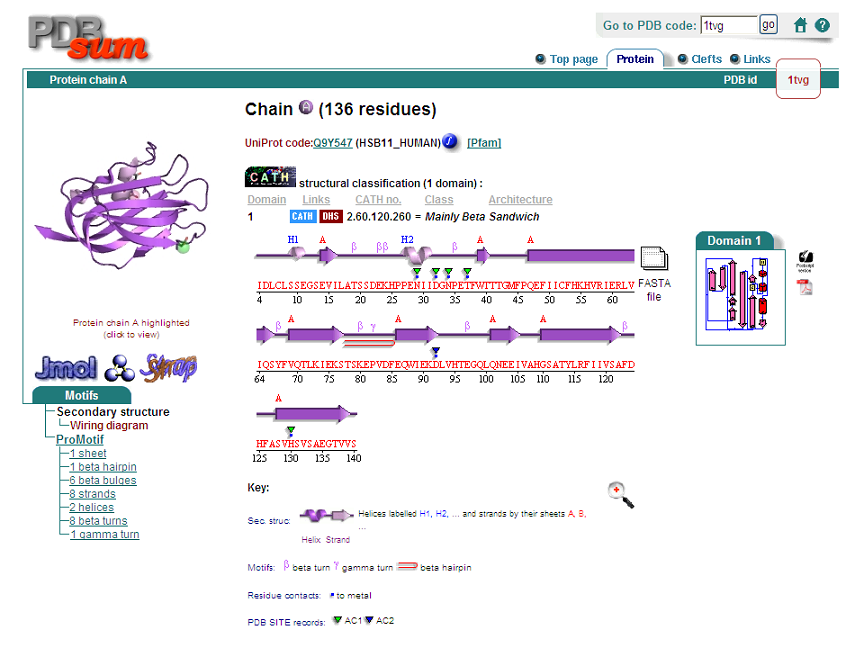
[[Image:Pdb sum.png]] | [[Image:Pdb sum.png]] | ||
Revision as of 01:42, 9 June 2009
Protein Structure
PDB ID: 1TVG Information from the PDB stated that x-ray diffraction was sued to solve the structure of this protein with an R value of 0.215 and at 1.6Å. Two ligands were present in the crystal structure, a calcium (II) ion and a samarium (III) ion. This protein has 153 residues and its secondary structure consists of two helices and nine beta sheet strands.
Above is the representation of 1TVG secondary structure using PDBsum.
Structure analysis
DALI was used to find protein that are similar in structure to 1TVG. The results showed that sialidases, alpha-N-acetylglucosaminidases and galactose oxidases have similar structure to our protein as presented below.
This protein's secondary structure consists of 2 alpha helices and 9 beta sheets.
Pymol structure displaying the folding of the secondary structure. Red is alpha helices, yellow is beta sheets and green is random coils connecting the the sheets and helices.