Chromosome 1 open reading frame 41 Structure
Protein Structure
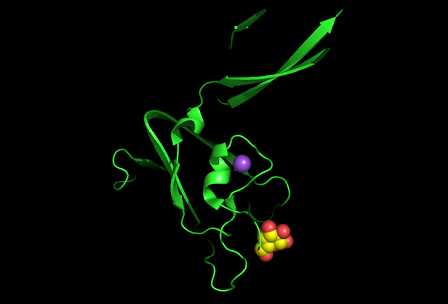
PDB ID: 1TVG Information from the PDB stated that x-ray diffraction was sued to solve the structure of this protein with an R value of 0.215 and at 1.6Å. Two ligands were present in the crystal structure, a calcium (II) ion and a samarium (III) ion. This protein has 153 residues and its secondary structure consists of two helices and nine beta sheet strands.
Scop result Root: scop Class: All beta proteins [48724] Fold: Galactose-binding domain-like [49784] sandwich; 9 strands in 2 sheets; jelly-roll Superfamily: Galactose-binding domain-like [49785] uperfamily Family: APC10-like [69210] Pfam 03256 Protein: Placental protein 25, pp25 [117104] Species: Human (Homo sapiens) [TaxId: 9606] [117105] SQ Q9Y547
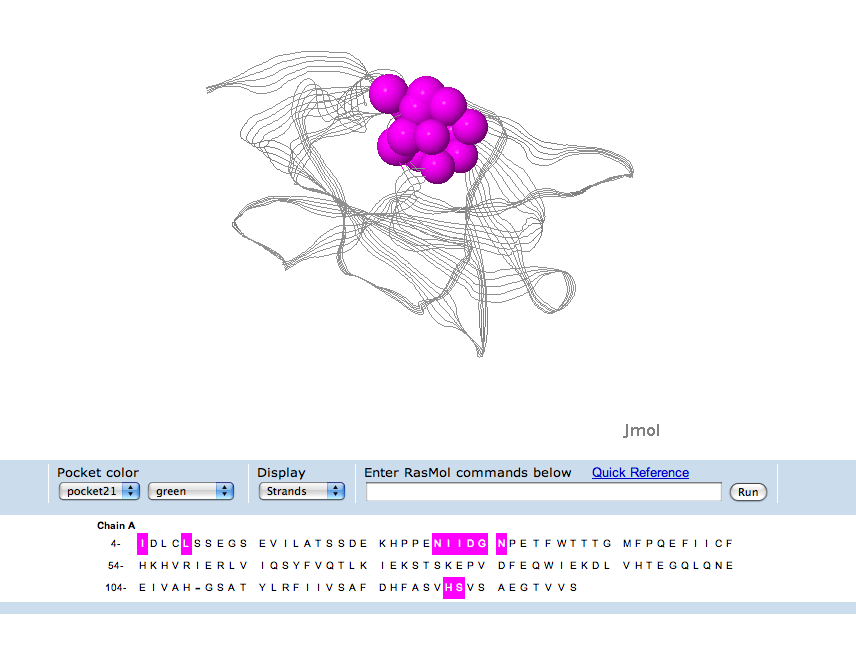
Figure 1: Above is the representation of c1orf41 secondary structure as presented in PDBsum.
Tertiary structure
Figure 2: This cartoon representation of c1orf41 shows that this protein is monomeric, consisting of a single domain. the fold of the protein is a jelly roll barrel where four pairs of anti-parallel beta strands are organised to form a barrel-like structure.
Structure analysis
1)Structure similarities
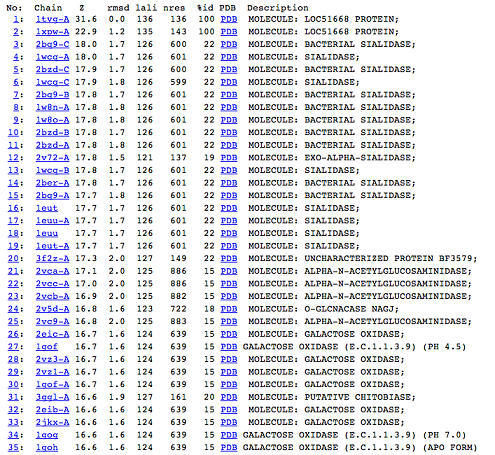
DALI was used to find proteins that are similar in structure to c1orf41. The results showed that sialidases, alpha-N-acetylglucosaminidases and galactose oxidases have similar structure to this protein.
Figure 5: DALI result showing the first 35 proteins that have similar structures to c1orf41. Even though the proteins are similar in structure they have very little sequence identity to c1orf41. 1xpw is c1orf41 but the structure was solved by NMR.
2)Domain Classification
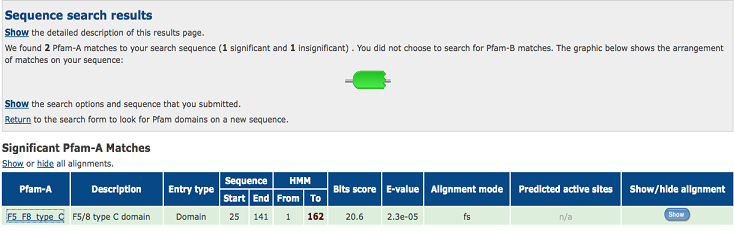
Pfam result indicated that our protein has a F5/8 type C domain, also known as the discoidin domain which is apart of galactose binding domain super family.
Closer inspection of the other proteins from the DALI result showed that they also have discoidin domain in their structures.
3)Possible ligand binding sites
Observations and comparisons of several proteins from the DALI result with our protein, there is a similar position on each proteins where a metal ion is located.
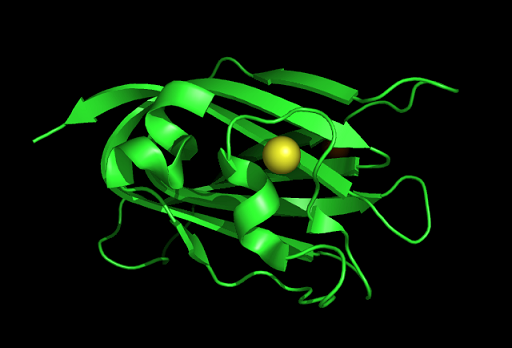
This figure shows the position where calcium (II) ion (yellow) is located in within our protein structure. This loop structure was also observed in several top proteins in the DALI result.
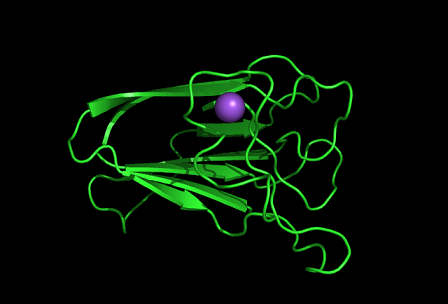
Shown is the discoidin domain of galactose oxidase and a sodium ion (purple).
Shown is the discoidin domain of bacterial sialidase and a sodium ion (purple) and a beta-D-galactose molecule.
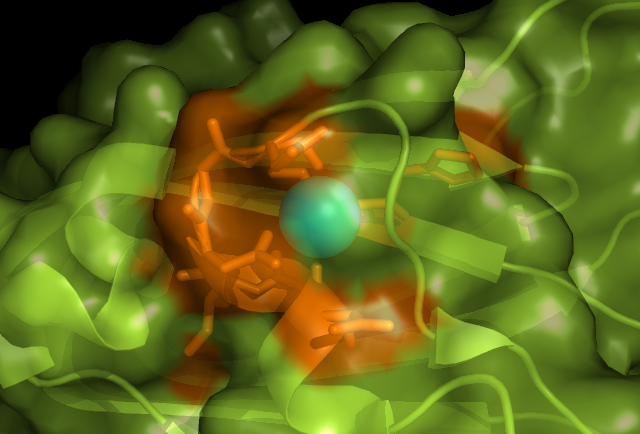
CastP also showed a surface cleft that is includes the loop region as shown below.
Below is a possible metal binding position on c1orf41. Orange is the residues that may be involved in coordinating the metal and blue is the calcium ion.
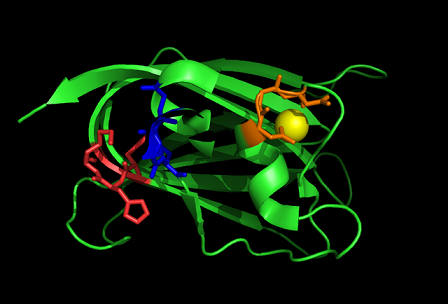
Nest analysis by Profunc suggested two other ligand binding sites.
The locations of the binding sites are shown below. CastP also predicted surface clefts similar to the erea of the nests.