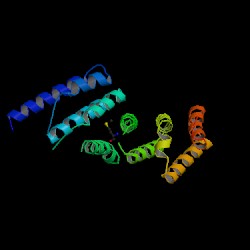
2i2O Protein Structure Presentation
Materials and Methods
1) Protein search on PDB, NCBI Entrez, Pfam and InterPro databases
- PDB code used: 2i2o
2) Identifying other similar structures with DALI server
3) Comparing the identified structures using CE comparison
Results
PDB
- Protein was isolated form Danio rerio and expressed in Escherichia coli.
- Structure has similarities with the “Eukaryotic Translation Initiation Factor 4G (eIF4G)” protein.
- Protein has 2 chains; and consists of nickel (Ni2+) and selenomethionine (C5H11NO2Se) components.
Pfam and InterPro
- Identified as "Middle domain of eIF4G", termed MIF4G.
- Consists essentially of alpha helices and has “multiple alpha-helical repeats” (reference). Within eIF4G, it binds to the RNA helicase eIF4A, eIF3, RNA and DNA.
DALI results
- SUMMARY: PDB/chain identifiers and structural alignment statistics
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3028-A 2i2o-A 37.1 0.0 206 206 100 0 0 1 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION eif4g-like prote 2: 3028-A 1hu3-A 15.2 2.7 164 204 23 0 0 12 S TRANSLATION eif4gii fragment (eukaryotic initiation f 3: 3028-A 1uw4-B 10.9 3.6 164 247 9 0 0 12 S NONSENSE MEDIATED MRNA DECAY PROTEIN regulator of nons 4: 3028-A 1h6k-A 10.6 3.9 175 728 11 0 0 11 S NUCLEAR PROTEIN cbp80 fragment (ncbp 80 kda subunit, 5: 3028-A 2db0-A 8.1 3.3 148 239 14 0 0 13 S PROTEIN BINDING 253aa long hypothetical protein (hypot 6: 3028-A 1b3u-A 8.0 27.7 150 588 13 0 0 13 S SCAFFOLD PROTEIN protein phosphatase pp2a fragment
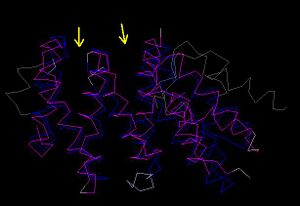
CE Structural comparison
- 1hu3 and 2i2o: Alpha-helices between 2i2o and 1hu3 were folded in same orientation
- 1h6k and 2i2o: N-terminal region of 1h6k displayed similarity with 2i2o; hence a possibility that MIF4G protein is a region, or even an active domain, near the N-terminal of the nuclear cap binding protein complex (CBC).
Discussion
The above results demonstrate a parallel, highly similar structure of eIF4G-like protein to human MIF4G (PDB: 1hu3). While there is a lack of conclusive studies and evidence on our protein of interest, the established protein MIF4G can be utilised as a model to formulate predictions on its structure. According to Wagner et al., MIF4G contains five helical hairpins oriented in a right-handed solenoid, and is similar to the HEAT [Huntingtin, elongation factor 3, a subunit of protein phosphatase 2A (PP2A), and target of rapamycin] domain (Marintchev & Wagner 2005). The overall protein resembles a crescent, with its superhelical axis perpendicular to the cylindrical axes of the alpha helices (Marcotrigiano et al, 2001). The helical hairpins are stacked one on top of the other to confer the protein its overall crescent shape.
MIF4G binds to eIF4A, an RNA helicase, and RNA, hence rendering its role in regulating cell translation. A protease-resistant region identified by proteolysis and mass spectrometry is speculated to be the binding site for eIF4A (Marcotrigiano et al, 2001).
MIF4G is one of the three domains present on eIF4G RNA regulatory protein. eIF4G has three domains, MIF4G, MA3 and W2, which are connected by linkers (Marintchev & Wagner 2005). It was hypothesized that the C’ terminal linker segment of MIF4G interacts with the domain by wrapping around the inter-helical grooves of the domain (Marintchev & Wagner 2005). This will account to an extent the presence and potential function of the numerous grooves observed in the domain structure.
All in all, information regarding the structure of eIF4G-like protein (our protein of interest) is limited to the current knowledge of its MIF4G homologue. Any further hypotheses on structural organization and interactions of the domain within the eIF4G protein necessitate additional analysis.