Talk:Structural analysis
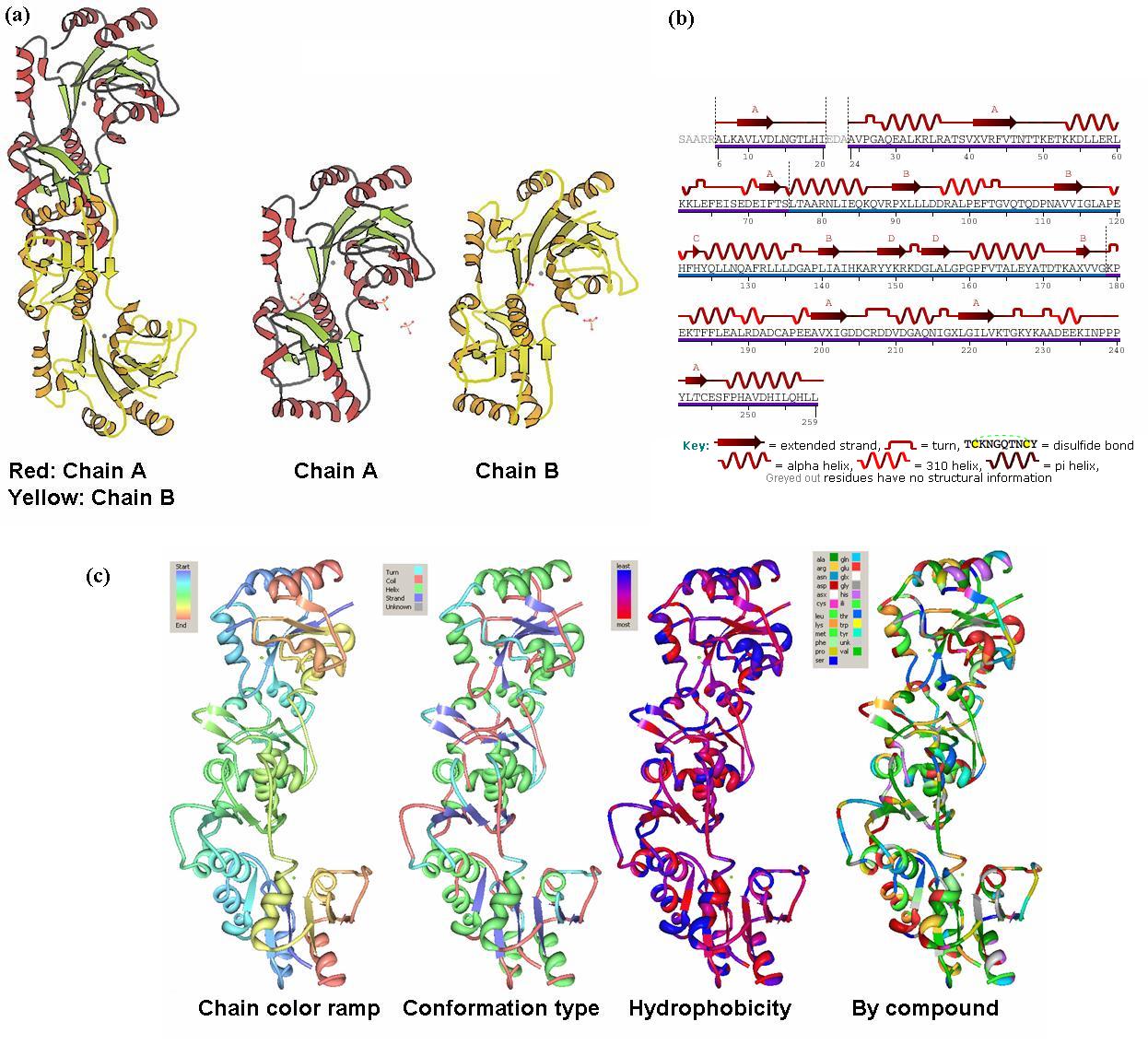
Structural Description
Residues = 259 Mol. Wt = 29KDa Domains = 2 Conformation = mainly aplha helices and beta-sheets Exists as dimers (monomers bind near hydrophobic region)
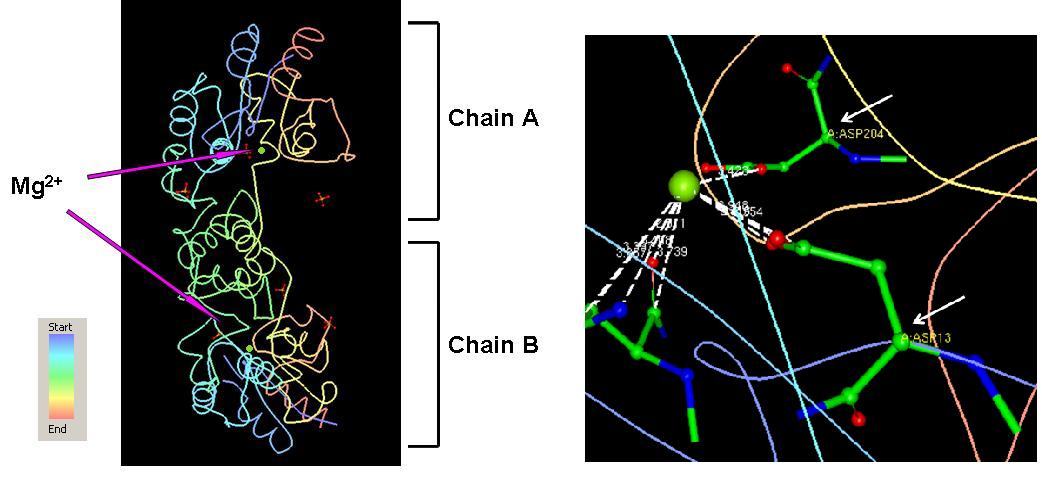
Major Ligands = Mg2+ Coordinates = Asp 13, Asn 15, Asp 204, and Asp 305 on each of the chains as shown in (Fig 2).
Structural Comparison
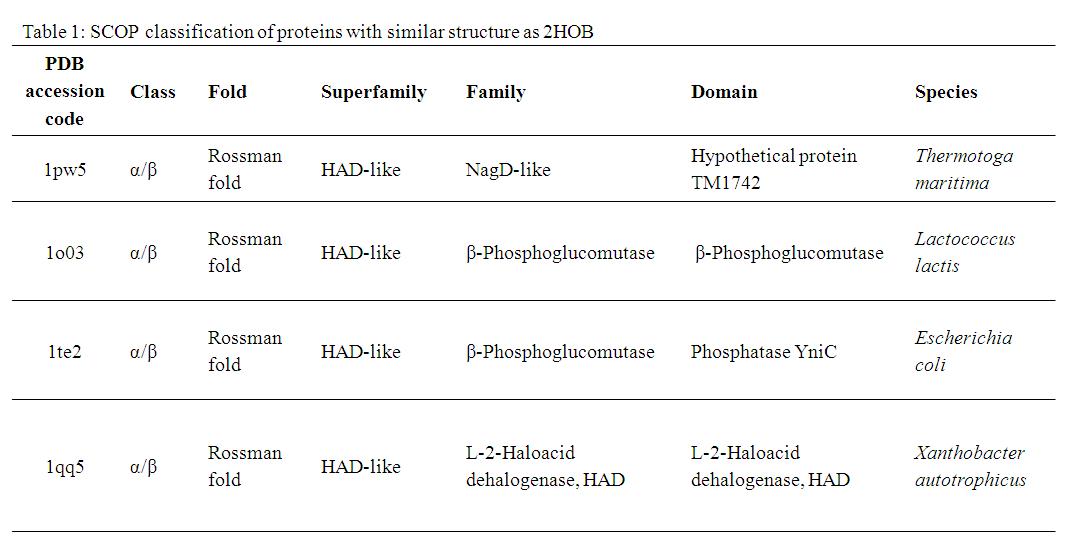
A Dali search with a strict threshold of Z > 12 for 2HO4 found 10 matches. Of these 10 protein matches, we found that most of them were hydrolases belonging to various different families. We reduced our structural matches of 10 proteins to only the ones that had been previously annotated. These proteins had sequence identities within the range of 17% to 25% and they included a putative NagD-like protein of Thermotoga maritima, the β-phosphoglucomutase of Lactococcus lactis , the β-phosphoglucomutase of Escherichia coli, and the L-2-Haloacid dehalogenase of Xanthobacter autotrophicus. The results are summarized in Table 1.
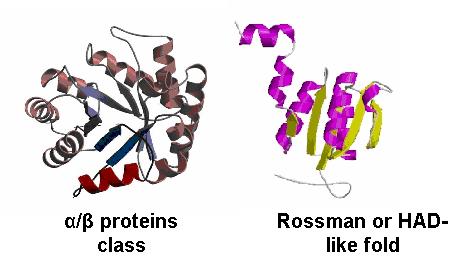
By looking at the architectural classification (using SCOP) of these proteins, it was noted that all of these proteins structures consisted of the same class, fold, and superfamily. The class was alpha and beta (α/β), which contains many parallel beta sheets (beta-alpha-beta units). Likewise, all the proteins had a Rossman fold (or HAD-like fold) consisting of three layers of α/β/α and parallel beta-sheets of 6 strands (Fig 3). And the superfamily they belonged to was HAD-like, which usually contains an insertion (sub)domain after strand 1.
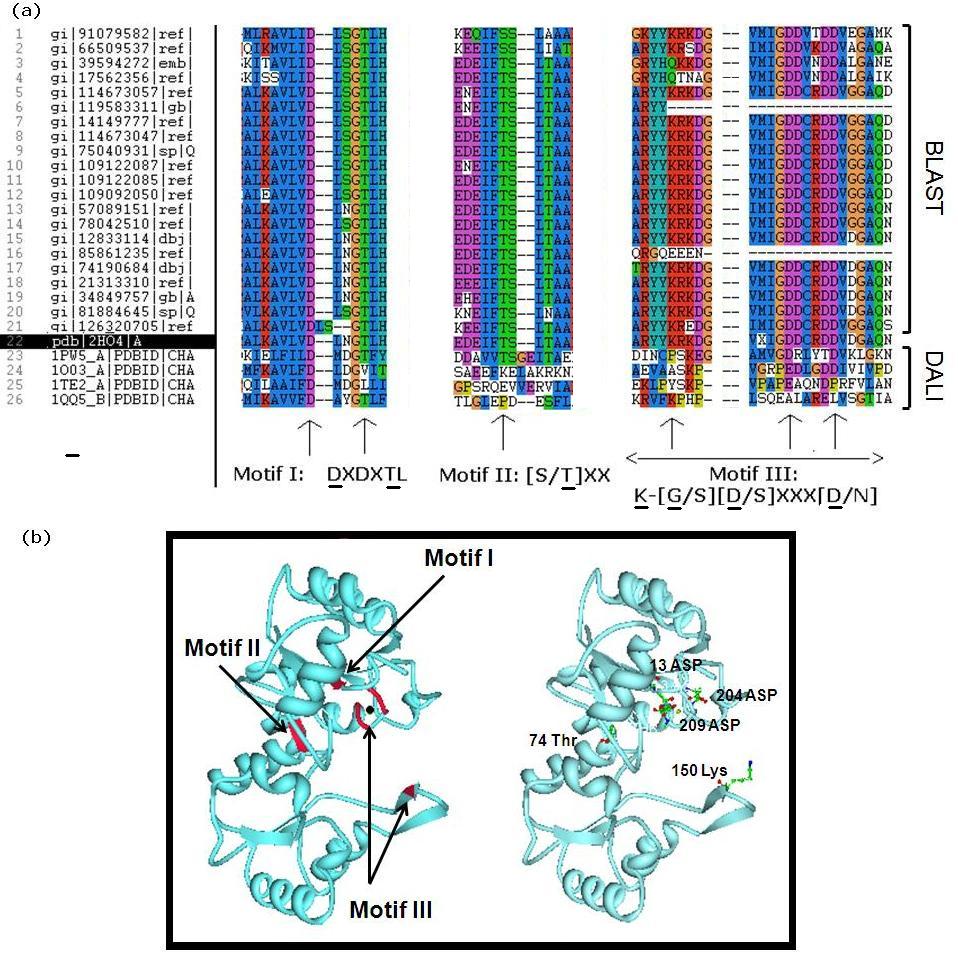
Literature seach indicated that members of the HAD superfamily included L-2-haloacid dehalogenases, sugar-phosphatases, and P-type ATPases, and all members shared three highly-conserved motifs. We did a multiple sequence analysis of H2O4 using the sequence homologs from BLAST as well as the structural homologs from DALI to show that it contains the three highly-conserved sequence motifs of the HAD superfamily. This confirmed that 2HO4 is a member of this superfamily (Fig 4a). The three motifs are: motif I: DXDX[T/V][L/N]; motif II: [S/T]XX; and motif III: K-[G/S][D/S]XXX[D/N]. These motifs lie next to each other and next the Mg2+ ion, together defining the active site of 2HO4 (Fig 4b) | (Rangarajan et al, 2006).
.
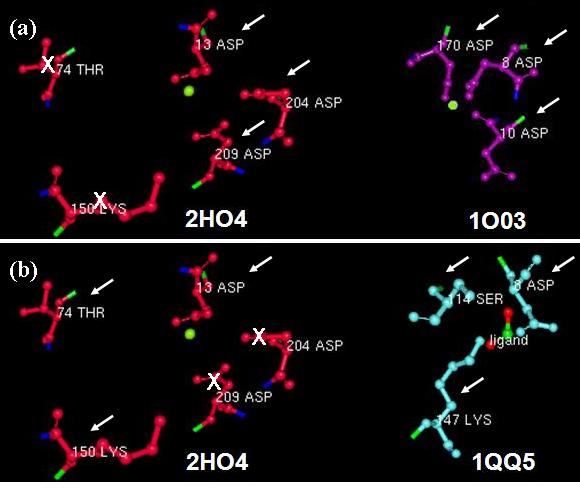
Furthermore, we investigated the alignment of the residues around the active site in space for any similarities. From the DALI results we used two well studied proteins: the β-Phosphoglucomutase of L. lactis (PDB code 1O03), and the L-2-Haloacid dehalogenase of X. autotrophicus (PDB code 1QQ5). We found that 2HO4 and β-Phosphoglucomutase of L. lactis had the three Asp residues from motifs I and III in common (Fig 5a), whereas 2HO4 and the L-2-Haloacid dehalogenase of X. autotrophicus had the Asp from motif I, the Ser/Thr from motif II, and the Lys from motif III in common (Fig 5b). Besides the architectural similarity between 2HO4 and β-Phosphoglucomutase of L. lactis (1O03), the L. lactis protein also consists of a Mg2+ ion bound to its catalytic site. It is possible that the two proteins share similar reaction mechanisms.
In order to determine which family of the HAD superfamily 2HO4 belonged to, we reviewed the literature. We found that HAD members with the conserved Asp residues and Mg2+ dependent reaction mechanisms could be P-type ATPases, which are involved in ATP hydrolysis.