Structure for haloacid dehalogenase-like hydrolase domain containing 2
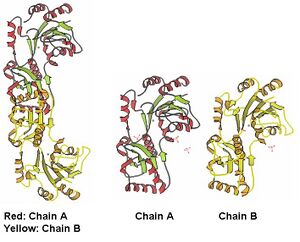
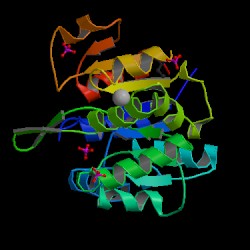
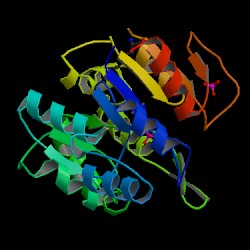
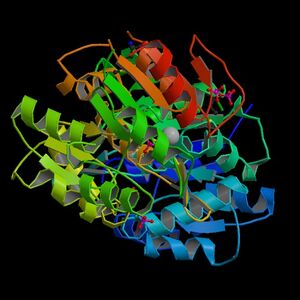
The information on the PDB website suggested that 2HO4 is a hydrolase belonging to the haloacid dehalogenase (HAD) superfamily, consisting of 2 domains. It consists of 259 residues and has a molecular weight of 29 kDa. Moreover, the PDB structure viewers demonstrated that 2HO4 is a monomer but forms a dimer with itself to give Chain A and chain B (Fig 1).
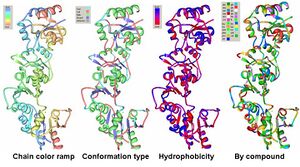
The PDB workshop enabled us to examine the various properties of 2HO4 including its conformation type and hydrophobocity. Its secondary structure mainly consists of alpha helices and beta sheets with 44% helical (14 helices; 123 residues) 17% beta sheet (12 strands; 48 residues) as shown in Fig 2. The two chains bind near their hydrophobic regions to produces dimeric structures. The outer surfaces of the dimeric protein is hydrophilic whereas the more hydrophobic regions lie near the site of binding (Fig 2).
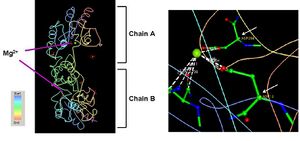
2HO4 has three different ligands, namely, selenium, a magnesium ion, and three phosphate ions (Fig 1). The PDB structures of 2HO4 indicate the presence of one Mg2+ ion, which binds between the coordinates Asp 13 and Asp 204 on each of the chains as shown in (Fig 3).
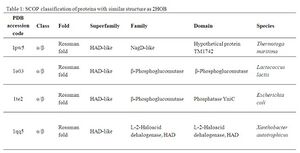
A Dali search with a strict threshold of Z > 12 for 2HO4 found 10 matches. Of these 10 protein matches, most were found to be hydrolases belonging to various different families. We reduced our structural matches of 10 proteins to only the ones that had been previously annotated. These included a putative NagD-like protein, an isomerase β-phosphoglucomutase, a putative phosphatase, and a L-2-Haloacid dehalogenase. The results are summarized in Table 1.
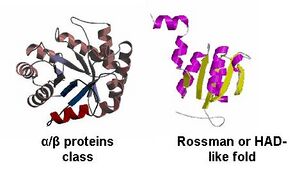
By looking at the architectural classification (using SCOP) of these proteins, it was noted that all of these proteins structures consisted of the same class, fold, and superfamily. The class was alpha and beta (α/β), which contains many parallel beta sheets (beta-alpha-beta units). Likewise, all the proteins had a HAD-like fold consisting of three layers of α/β/α parallel beta-sheet, which amy also be called a Rossman fold (Fig 4). And the superfamily they belonged to was also HAD-like, which usually contains an insertion (sub)domain after strand 1.
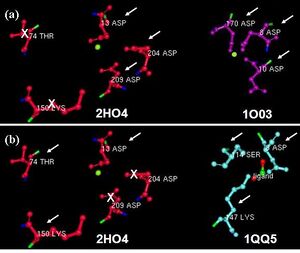
Multiple sequence analysis of H2O4 using the sequence homologs from BLAST as well as the structural homologs from DALI shows that it contains the three highly-conserved sequence motifs of the HAD superfamily, indicating that 2HO4 is a member of this superfamily (Fig 5a). The three motifs are: motif I: DXDX[T/V][L/N]; motif II: [S/T]XX; and motif III: K-[G/S][D/S]XXX[D/N]. These motifs lie next to each other and next the Mg2+ ion, together defining the active site of 2HO4 (Fig 5b) | (Rangarajan et al, 2006).
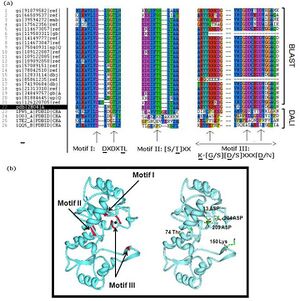
.
Dali revealed the β-Phosphoglucomutase of Lactococcus lactis (PDB accession code 1O03) to be structurally similar to 2HO4 with 24% sequence identity. We found that they had the three residues in common, which were conserved in the three HAD motifs. Figure 6 shows a comparison of the common residues in 2HO4 and 1O03 present near their active sites (Fig 6). Besides the architectural similarity between 2HO4 and 1O03, 1O03 also consists of a Mg2+ ion bound to its catalytic site. It is possible that the two proties share similar reaction mechanisms.
ROUGH
Pfam description (Accession number: PF00702)
This family are structurally different from the alpha/ beta hydrolase family (Abhydrolase_1). This family includes L-2-haloacid dehalogenase, epoxide hydrolases and phosphatases. The structure of the family consists of two domains. One is an inserted four helix bundle, which is the least well conserved region of the alignment, between residues 16 and 96 of HAD1_PSEUC. The rest of the fold is composed of the core alpha/beta domain.
N terminus = blue C terminus = red so the first 100 residues (aa's) should be in the blue region.
InterPro description (entry IPR005834)
This group of hydrolase enzymes is structurally different from the alpha/beta hydrolase family (abhydrolase). This group includes L-2-haloacid dehalogenase, epoxide hydrolases and phosphatases. The structure consists of two domains. One is an inserted four helix bundle, which is the least well conserved region of the alignment, between residues 16 and 96 of HAD1_PSESP. The rest of the fold is composed of the core alpha/beta domain
Structure similarity Table (results from dali search)
| number | PDB code | structural similarity (RMSD) (better if zero) | seuqence identity (%IDE) (better if 100%) | protein name | comments |
|---|---|---|---|---|---|
| 2 | 1pw5 | 2.9 | 25 | STRUCTURAL GENOMICS, UNKNOWN FUNCTION, nagd protein, pu | Could be a hydrolase, not sure |
| 3 | 2hx1 | 4.7 | 23 | HYDROLASE: predicted sugar phosphatases of the had superfamily | The E. coli HAD phosphatases show high catalytic efficiency and affinity to a wide range of phosphorylated metabolites that are intermediates of various metabolic reactions. (pubmed) |
| 5 | 1o03 | 2.5 | 24 | ISOMERASE: beta-phosphoglucomutase (in lactococcus lactis) | Phosphoglucomutases catalyze the interconversion of D-glucose 1-phosphate and D-glucose 6-phosphate, a reaction central to energy metabolism in all cells and to the synthesis of cell wall polysaccharides in bacterial cells. Two classes of phosphoglucomutases (alpha-PGM and beta-PGM) are distinguished on the basis of their specificity for alpha- and beta-glucose-1-phosphate. beta-PGM is a member of the haloacid dehalogenase (HAD) superfamily, which includes the sarcoplasmic Ca(2+)-ATPase, phosphomannomutase, and phosphoserine phosphatase. beta-PGM is unusual among family members in that the common phosphoenzyme intermediate exists as a stable ground-state complex in this enzyme. Herein we report, for the first time, the three-dimensional structure of a beta-PGM and the first view of the true phosphoenzyme intermediate in the HAD superfamily. (Pubmed) |
| 22 | 2fpx | 3.3 | 20 | HYDROLASE: histidine biosynthesis bifunctional protein | HisB from Escherichia coli is a bifunctional enzyme catalyzing the sixth and eighth steps of l-histidine biosynthesis. The N-terminal domain (HisB-N) possesses histidinol phosphate phosphatase activity, and its crystal structure shows a single domain with fold similarity to the haloacid dehalogenase (HAD) enzyme family. (Pubmed) |
| 35 | 1f5s | 3.1 | 23 | HYDROLASE: phosphoserine phosphatase (psp) (in Methanococcus jannaschii) | D-Serine is a co-agonist of the N-methyl-D-aspartate subtype of glutamate receptors, a major neurotransmitter receptor family in mammalian nervous systems. D-Serine is converted from L-serine, 90% of which is the product of the enzyme phosphoserine phosphatase (PSP). PSP from M. jannaschii (MJ) shares significant sequence homology with human PSP. PSPs and P-type ATPases are members of the haloacid dehalogenase (HAD)-like hydrolase family, and all members share three conserved sequence motifs. PSP and P-type ATPases utilize a common mechanism that involves Mg(2+)-dependent phosphorylation and autodephosphorylation at an aspartyl side chain in the active site (Abstract from PBD). |
| 38 | 1j8d | 3.0 | 19 | HYDROLASE: deoxy-D-mannose-octulosonate 8-phosphate phosphatase (YrbI) From Haemophilus Influenzae (HI1679) | The crystal structure of the YrbI protein from Haemophilus influenzae (HI1679) was determined at a 1.67-A resolution. The function of the protein had not been assigned previously, and it is annotated as hypothetical in sequence databases. The protein exhibits the alpha/beta-hydrolase fold (also termed the Rossmann fold) and resembles most closely the fold of the L-2-haloacid dehalogenase (HAD) superfamily. Following this observation, a detailed sequence analysis revealed remote homology to two members of the HAD superfamily, the P-domain of Ca(2+) ATPase and phosphoserine phosphatase (Abstract from PDM). |
DALI server results
threshold Z > 9
FSSP FAMILIES OF STRUCTURALLY SIMILAR PROTEINS, VERSION 1.0 (Apr 1 1995) CREATED Tue May 8 05:22:07 BST 2007 for dali on s030-014.ebi.ac.uk METHOD Dali ver. 2.0: Holm, L., Sander, C. (1993) J.Mol.Biol. 233,123-138 DATABASE 8969 protein chains PDBID 3021-A HEADER COMPND SOURCE AUTHOR SEQLENGTH 246 NALIGN 242 WARNING pairs with Z<2.0 are structurally dissimilar
NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN 1: 3021-A 2ho4-A 43.4 0.0 246 246 100 0 0 1 S HYDROLASE haloacid dehalogenase-like hydrolase domain 2: 3021-A 1pw5-A 24.3 2.9 230 246 25 0 0 17 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION nagd protein, pu 5: 3021-A 1o03-A 13.3 2.5 135 221 24 0 0 12 S ISOMERASE beta-phosphoglucomutase (lactococcus lactis 7: 3021-A 1te2-A 12.8 2.4 133 211 17 0 0 15 S HYDROLASE putative phosphatase (escherichia coli o157 9: 3021-A 1qq5-A 12.5 2.1 138 245 20 0 0 11 S HYDROLASE l-2-haloacid dehalogenase (xanthobacter aut 10: 3021-A 2hsz-A 12.2 2.2 125 222 20 0 0 20 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION novel predicted 11: 3021-A 1yns-A 11.6 2.2 127 254 14 0 0 17 S HYDROLASE e-1 enzyme (enolase-phosphatase e1) (homo s 12: 3021-A 2ah5-A 11.3 2.4 125 206 23 0 0 20 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION cog0546 (strept 13: 3021-A 1wdh-A 11.2 2.2 125 248 14 0 0 17 S 14: 3021-A 2p11-A 10.4 2.4 130 211 18 0 0 15 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 15: 3021-A 1rku-A 10.4 2.9 132 206 14 0 0 15 S TRANSFERASE homoserine kinase (pseudomonas aeruginosa 16: 3021-A 2go7-A 10.3 2.5 120 203 25 0 0 15 S HYDROLASE hydrolase, haloacid dehalogenase-like family 17: 3021-A 2fdr-A 10.3 2.3 128 214 20 0 0 17 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION conserved hypoth 18: 3021-A 1ymq-A 10.2 3.7 163 260 17 0 0 22 S TRANSFERASE sugar-phosphate phosphatase bt4131 (bacte 19: 3021-A 2fi1-A 10.0 2.4 112 187 22 0 0 14 S HYDROLASE hydrolase, haloacid dehalogenase-like family 20: 3021-A 1rlm-A 10.0 3.2 146 269 12 0 0 21 S HYDROLASE phosphatase Mutant (escherichia coli) bacte
Literature to read
[| YbiV from Escherichia coli K12 is a HAD phosphatase.]
The protein is comrprised of 2 Biological units.
The Whole protein comprised of both compounds.