2i2O Discussion
Evolution of the MIF4G Domain Containing Protein
From the research conducted on the MIF4G domain containing protein, it was found to be part of the eIF4G domain which function is to initiate the translation of RNA in eukaryotic organisms. With the evolution of more complex organisms, the innovation of mew components of the machinery used in intiating translation was needed (Aravind & Koonin 2000). Most of these new domains that evolved contain a number of alpha-helicases within the structure of the new proteins formed (Aravind & Koonin 2000). The early ancestor of eukaryotes was thought to contain only one major eIF4G domain. Over the course of evolution, divergence of this one domain occured (Marintcher & Wagner 2005). Earlier studies indicate that this divergence within the eIF4G domain occur early in the evolution of the eukaryotes. The three domains that make up the larger eIF4G translation initiation factor are all present in the vertebrate animals (Martincher & Wagner 2005). To gain a better understanding in the evolutionary history of eIF4G, a focus on the middle domain protein was undertaken.
The multiple alignment sequence conducted on the 55 sequences indicate several areas within the protein where the sequences are conserved. This suggests that these parts of the protein has an important function. This would be the reason why the conservation between the sequences are observed across the species. All theses sequences were from the Metazoa clade.
There could be several reasons why plant and fungi sequences were not incorporated into the constructed phylogenetic tree. It is quite possible that there were several plant and fungi sequences, but their e-values were not significant enough and these sequences were discarded before the next step in constructing the phylogenetic tree. It is quite likely that plant and fungi do have MIF4G domain containing protein but the sequences are not homologous with the protein found in humans. Also, research does indicate that several eukaryotic families that ancestrally contained the three protein domains of eIF4G have undergone major gene deletions. This may have resulted in the loss of the MIF4G domain containing protein from plant and fungi species (Martincher & Wagner 2005).
The two smaller trees located within the bootstrapped phylogenetic tree may indicate a divergence of the ancestral MIF4G domain containing protein. This divergence may indicate two or more MIF4G proteins present within the same species. The reason for this is that they perform slightly different functions in different cells. Analysis on the different protein sequences with the different species should be carried out. This would allow the proteins' structure and function to be determined within the different species. This would enable the relationship to be observed between the MIF4G protein found in humans and the MIF4G found in different species.
Function of the MIF4G Domain Containing Protein
ProFunc search was performed to identify the potential function of the protein. Its collaborative approach integrating many databases was extremely useful in linking all known information about the potential structure and function (see Figure 2.0 and 2.1).
The query submitted to Pfam using MIF4G domain identified the domain to be occurring in the eukaryotic translation initiation factor IV (eIF4) as well as in NMD2p and CBP80. Literature available on these proteins showed them to be structurally similar to MIF4G (Marintchev & Wagner 2005) and also as being related to domains from within eIF4 domain known as HEAT domains. A paper by Perry and Kleckner (2003) described HEAT domain as a superhelical forming scaffolding matrice that is comprised of single HEAT repeat units made of a pair of anti-parallel helices linked by an elastic loop. These can also occur in series. Three consecutive HEAT domains are present within eIF4. As found by (Marintchev & Wagner, 2005) MIF4G is congruent with HEAT-1 and as such CBP80 (which also contains three HEAT domains and is structurally similar to eIF) has been hypothesised to be similar to the middle unit of CBP80. Such that MIF4G corresponds to HEAT-1 in the same way that the middle unit of CBP80 corresponds to its own HEAT-1. Marintchev & Wagner (2005) showed that the 2 proteins were derived from similar or the same origin and are highly conserved. Furthermore it is suggested MIF4G binds to eIF4A, an RNA helicase and RNA and is speculated to play a role in the binding of eIF4A to RNA.
The NEST analysis (Figure 2.6, Table 2.1) produced 3 hits with scores of 2.28, 3.46 and 4.96. Since all scores above 2 are considered to be functionally significant and residue conservation for the most part is high we can infer that these structural motifs are important sites for the function of the protein (Pal et al. 2002). Futher analysis is required before a more specific conclusion can be made as to the significance of identified NESTS and their functional properties.
Results from ProKnow (Figure 2.4) show that the likely function of MIF4G Domain containing protein is RNA binding. This was found using the frequency of ontology from 3D folds and the score of ontologies from 3D motifs based on conservation. Thus by combining the information gathered about the function and functional sites we can hypothesise that the structural motifs identified by NEST are important for RNA binding.
The results generated by the LOCATE database show that the potential function is RNA binding. With information gathered from other sites we can use this to infer a more specific function, for instance the type of and position on RNA the protein binds and its purpose for doing so. The method for predicting the location of the protein is MultiLoc prediction. It is based on Amino Acid sequence and the presence of an N-terminal targeting sequence. Based on this prediction method there is a 0.93 probability (93% chance) that the protein is cytoplasmic.
In addition to the location, three proteins were identified as Riken cDNA templates, similar to the location and possible function of MIF4G. In parallel to the Superfamily HMM program (Figure 2.3; Gough et al. 2001), LOCATE showed the protein to contain ARM repeats (armadillo repeats). ARM and HEAT motifs according to Andrade (Andrade et al. 2001) are repeats approximately 50 residues long tandemly repeated throughout many eukaryotic proteins. The function of which is largely unknown except to say both repeats have been implicated as participating in the regulation of protein-protein interactions. Originally it was believed that ARM and HEAT repeats were similar however it has been recently found that the two repeats were divergent and contain significant structural and functional differences. Namely the ARM repeat consists of two helices and the HEAT domain of three. This is consistent with our findings, that the domain is rich in alpha helices. This is significant to our domain containing protein MIF4G as it provides more evidence to the function being the mediation of protein-protein interactions during the initiation of translation. Further research would have to be undertaken to discover if the MIF4G HEAT domain in eIF4G consists of the ten alpha helices as five repeats important in cap dependant and cap-independent translation initiation as previoulsy suggested(Marcotrigiano et al. 2001).
Ten gap regions were analysed in Cleft (Table 2.2). Four of these are highly conserved and hydrophobic. These results allow us to see potential binding sites on the protein and their possible residues. The residue type provides clues as to the type of molecule that binds to the functional site and hence what the function of the protein is. Collaboratively, the results from LOCATE and ProKnow further show the function to be RNA binding during the initiation of translation phase. Unfortunately without experimental analysis there is no way of knowing exactly which residues are important for function and thus the exact function of the protein.
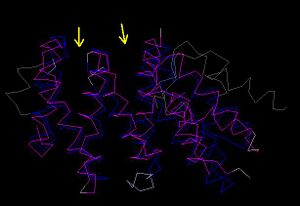
The results for 3D functional templates demonstrate the Reverse template comparison versus PDB structures of 1hu3 and 2i2O show they are significantly both structurally and functionally similar (Figure 2.5). The figure produced in results (Figure 2.5) show that all of the regions of high sequence identity (boxed sequence) correspond to structurally fittbale regions (as denoted by the red and blue arrows). This segment also contains three matched template side chains and several residues equivalenced from within 10Å of the template residues. Therefore it is possible to assume that the region could be a conserved functional region common to both proteins (Laskowski et al. 2005).
There were no lingand binding, DNA binding or enzyme active site templates found which makes sense in light of our other findings that our protein is RNA binding.
Structure of the MIF4G Domain Containing Protein
The above results demonstrate a parallel, highly similar structure of eIF4G-like protein to human MIF4G (PDB: 1hu3). While there is a lack of conclusive studies and evidence on our protein of interest, the established protein MIF4G can be utilised as a model to formulate predictions on its structure. According to Wagner et al., MIF4G contains five helical hairpins oriented in a right-handed solenoid, and is similar to the HEAT [Huntingtin, elongation factor 3, a subunit of protein phosphatase 2A (PP2A), and target of rapamycin] domain (Marintchev & Wagner 2005). The overall protein resembles a crescent, with its superhelical axis perpendicular to the cylindrical axes of the alpha helices (Marcotrigiano et al. 2001). The helical hairpins are stacked one on top of the other to confer the protein its overall crescent shape.
MIF4G binds to eIF4A, an RNA helicase, and RNA, hence rendering its role in regulating cell translation. A protease-resistant region identified by proteolysis and mass spectrometry is speculated to be the binding site for eIF4A (Marcotrigiano et al, 2001).
MIF4G is one of the three domains present on eIF4G RNA regulatory protein. eIF4G has three domains, MIF4G, MA3 and W2, which are connected by linkers (Marintchev & Wagner 2005). It was hypothesized that the C’ terminal linker segment of MIF4G interacts with the domain by wrapping around the inter-helical grooves of the domain (Marintchev & Wagner 2005). This will account to an extent the presence and potential function of the numerous grooves observed in the domain structure.
All in all, information regarding the structure of eIF4G-like protein (our protein of interest) is limited to the current knowledge of its MIF4G homologue. Any further hypotheses on structural organization and interactions of the domain within the eIF4G protein necessitate additional analysis.
Return to Scientific Report