DHRS1 Results
Phylogenetic Tree
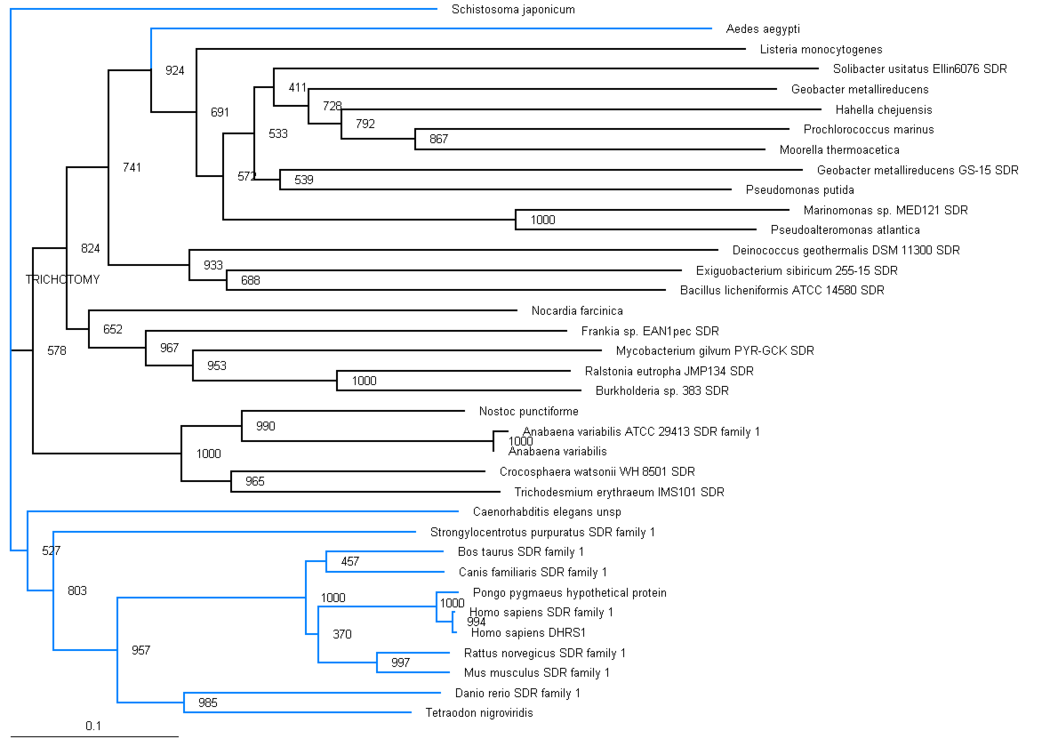
The phylogenic tree shows a small section of the family is contained in land based higher organisms (Bos taurus through to Mus Muluscus), a smaller section is found in sea based eukaryotes (Danio rerio and Tetraodon nigroviridis) and the majority of the family is round in bacteria. The majority of bacteria found to possess this family are proteobacteria, although there are also gram positive rod (eg. bacillus) and cyanobacteria (eg. anabena). This suggests that this particular family has been present for a long time and has evolved from bacteria, most likely proteobacteria. Blue branches show organisms of the anamalia kingdom, Black branches show organisms of the bacteria kingdom.
Aedes aegypti and Schistosoma japonicum are both found in the anamalia kingdom, although are seperated on the graph. Both of these organisms are parasitic by nature. Aedes aegypti is commonly known as the yellow fever mosquito. Schistosoma japonicum is a parasite and one of the major infectious agents of schistosomiasis.
The distribution of the branches in the tree is consistent with full organism genome taxonomic distribution, for example the prokaryotes are grouped together, and the eukaryotes are grouped together, with one distinct exception, Ades aegypti.
Clustal Alignment
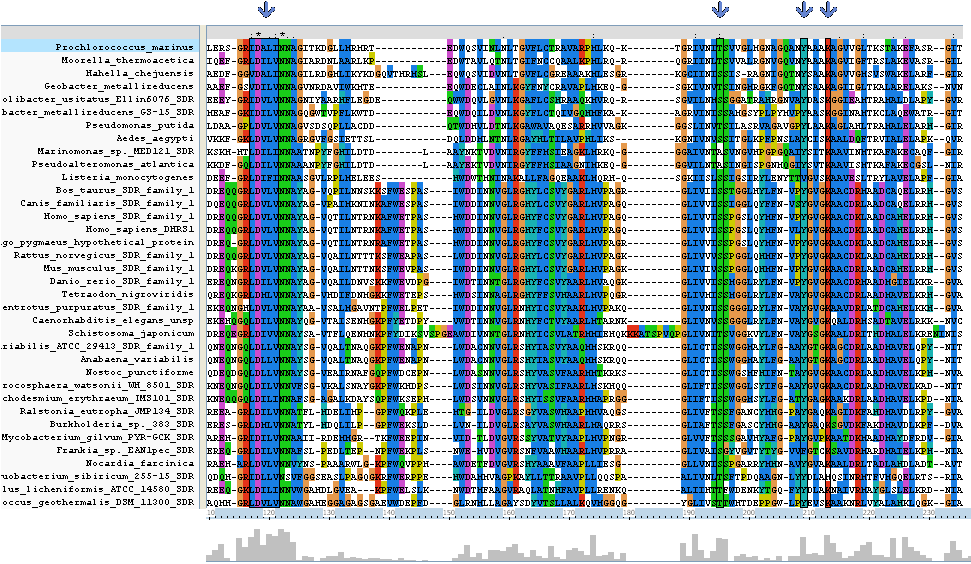
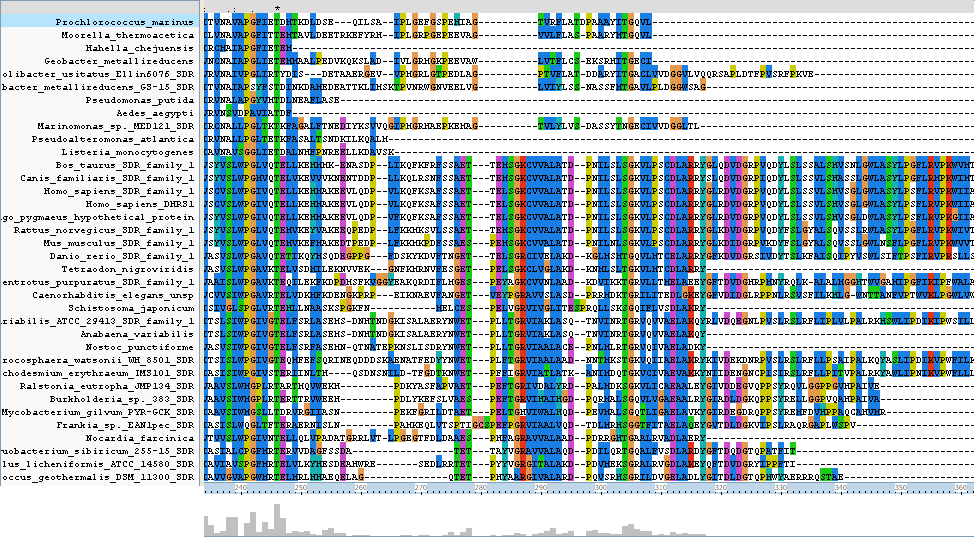
Clustal alignment shows that Homo_sapiens_SDR_family_1 (an exact match for our target protein) with regards to protein sequence, contains all eight key residues (locations 117 - 121, 195,209 and 213). It also contains a sequence K-[A,S]-F-W-E-x-P-A-S at location 138 - 146 which is conserved only between land based animals.
The N-terminus region of the clustal alignment shows the main difference between classical SDR's and extended SDR's. There is a highly conserved tail (locatoin 280 - 360) which appears only in some of the results aligned. As our target sequence is in the extended SDR family, most of the sequences aligned are also in the extended SDR family.
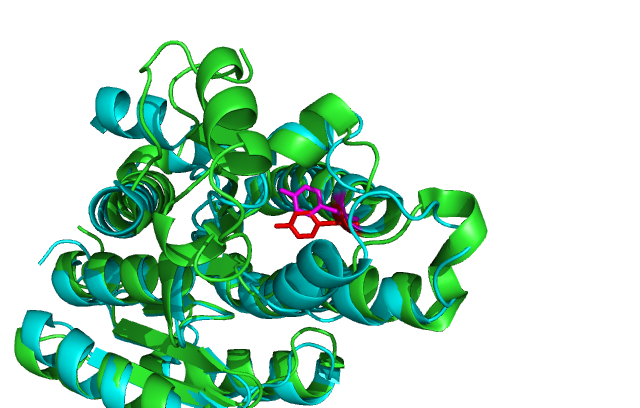
Structure
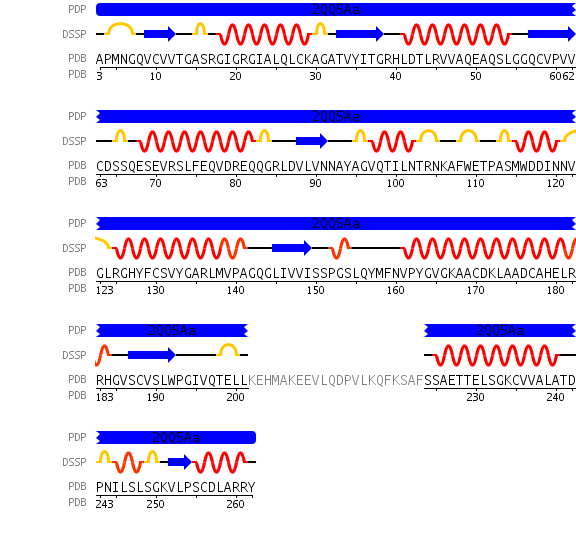
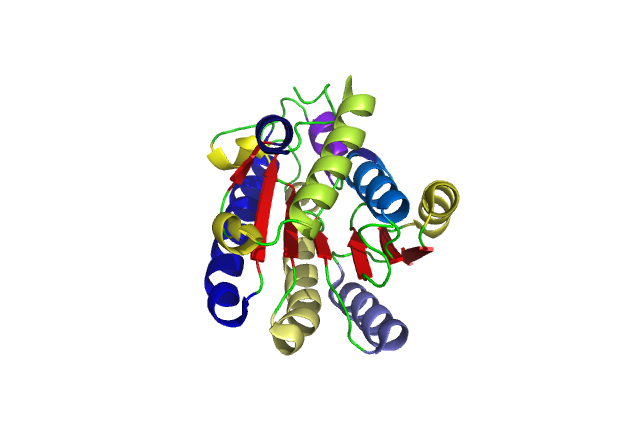
A single monomeric unit of DHRS1 contains 10 α helices 1 central β-sheet region consisting of 7 strands. It is thought to exist biologically as a dimer.
DHRS1 has highly conserved structure compared across the SDR family.
The SDR family is part of the Super Family; NAD(P)-binding Rossmann-fold domain proteins all of which have the Rossmann-fold domain, which is characterised by a central β-sheet surrounded by α-helicies. This puts them in the Alpha and Beta proteins (α/β) class.
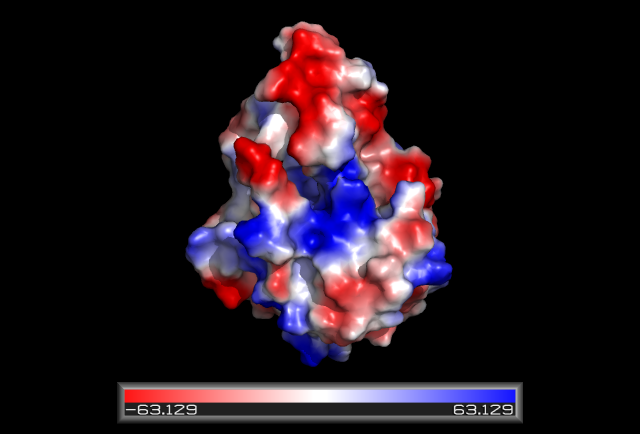
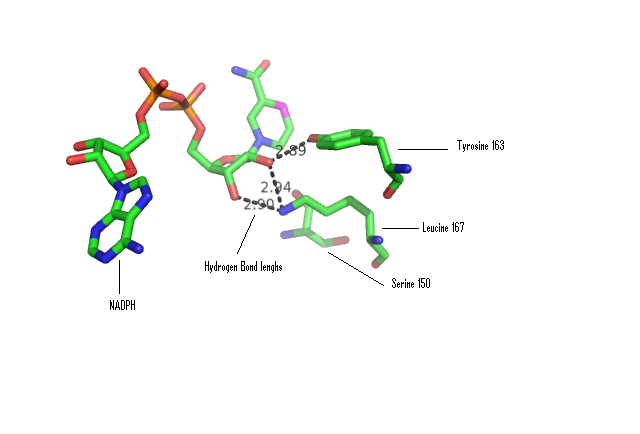
Some key residues are conserved across the entire family. Notable a Tyrosine that binds NAD(P), residue 163 in the sequence above. It is part of a larger motif [S-x(12)-Y-x(3)-K] (residues 150-167)that includes two other residues (Serine, Lysine) involved in binding NAD(P) (Wu Q, et al 2001) This catalytic triad is located in a deep hydrophobic pocket (fig 7) and is responsible for binding NADP (fig 10).
There are more highly conserved motifs including a [G-x(3)-G-x-G] (residues 14-20) that is characteristic of a possible co-enzyme binding site and [LDVLV] (residues 86-90) involved in the initial folding of the protein (Wu Q, et al 2001). It forms part of the hydrophobic core most notably a strand of the β-sheet that is part of the Rossmann-fold.
Structure comparison
DALI server was used to perform a structural alignment. There where many highly significant hits some from the SDR Family and all hits whre from the Super Family; NAD(P)-binding Rossmann-fold domain proteins (Table 1).
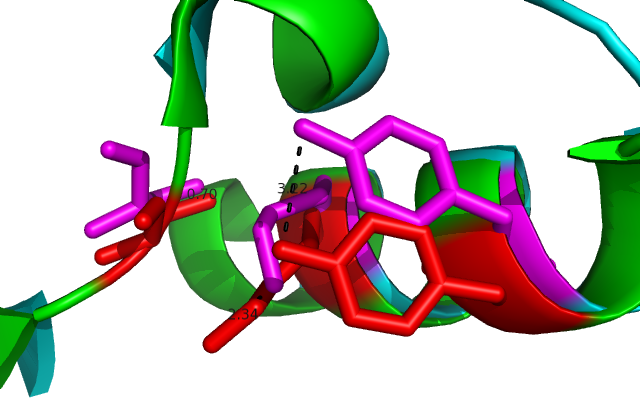
Comparison of DHRS1 to 3-Oxoacyl-(Acyl-Carrier-protien)reductase its closest structurally related protein, finds that whilst it shares only 27% of its sequence, it is structurally very similar (Fig 8). They share many structural features including the central β-sheet region and many α helices, all of which are very closely aligned. Key conserved residues such as the tyrosine shown are also in very similar positions eluding to a similar function. Importantly the key catalytic triad (S-Y-K) is still in the same place with the tips of the residues only moved 0.7-3 Angstroms (Fig 10).
Futher comparison with other related protiens from the DALI structural alignment (Table 1), notably all of the glucose dehydrogenases and the other member of the SDR family SDR4 show this pattern to continue.
Table 1
PDB/chain identifiers and structural alignment statistics from DALI search. Z = standard deviations above that expected. Z < 2.0 means no significant similarity.
No: Chain Z nres %id Description 1: 2qq5-A 48.1 238 100 MOL: DEHYDROGENASE/REDUCTASE SDR1; 2: 2uvd-A 29.6 246 27 MOL: 3-OXOACYL-(ACYL-CARRIER-PROTEIN) REDUCTASE; 3: 1yde-F 29.3 256 29 MOL: RETINAL DEHYDROGENASE/REDUCTASE 3; 4: 1vl8-B 29.1 252 27 MOL: GLUCONATE 5-DEHYDROGENASE; 5: 2bgk-A 29.0 267 25 MOL: RHIZOME SECOISOLARICIRESINOL DEHYDROGENASE; 6: 2q2q-D 28.9 255 26 MOL: BETA-D-HYDROXYBUTYRATEDEHYDROGENASE; 7: 1rwb-F 28.8 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 8: 1rwb-A 28.8 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 9: 1gee-A 28.8 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 10: 1gco-A 28.8 261 24 MOL: GLUCOSE DEHYDROGENASE; 11: 2zat-A 28.7 251 23 MOL: DEHYDROGENASE/REDUCTASE SDR4; 12: 1gee-B 28.7 261 24 MOL: GLUCOSE 1-DEHYDROGENASE; 13: 1gco-E 28.7 261 24 MOL: GLUCOSE DEHYDROGENASE;
Abstract | Introduction | Results | Discussion |
Conclusion | Method | References
Back to Main Page