Hypothetical protein Results
Abstract | Introduction | Method |
Results | Discussion | Conclusion | References
Evolution
Sequences
Sequences (FASTA format) gathered for MSA (including query), available via the following link: [1]
Multiple Sequence Alignment
 Figure 3: Multiple sequence alignment generated using Clustalx 1.83 [1].
Figure 3: Multiple sequence alignment generated using Clustalx 1.83 [1].
Original Curved Bootstrapped Tree
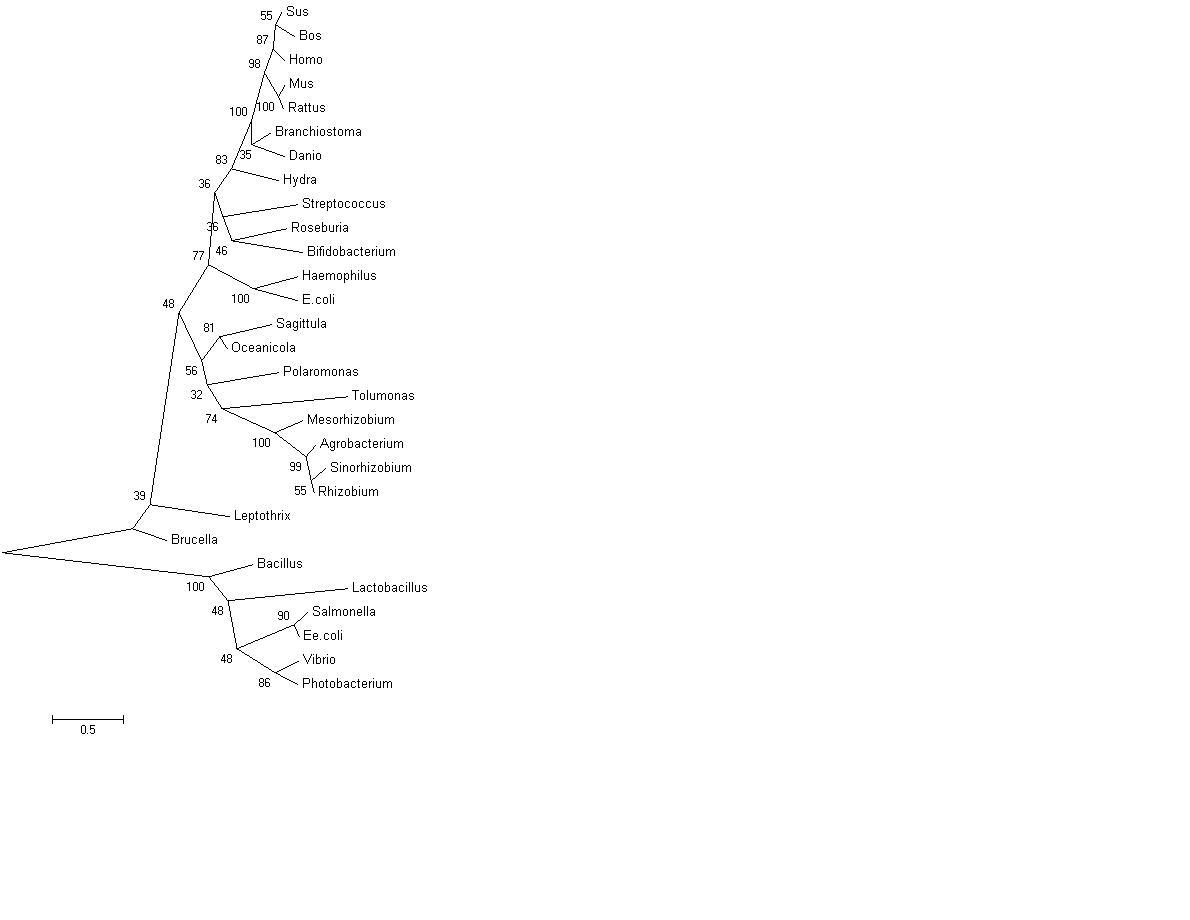 Figure 4: Curved phylogenetic tree with bootstrap values, obtained using MEGA 4.0 [2]
Figure 4: Curved phylogenetic tree with bootstrap values, obtained using MEGA 4.0 [2]
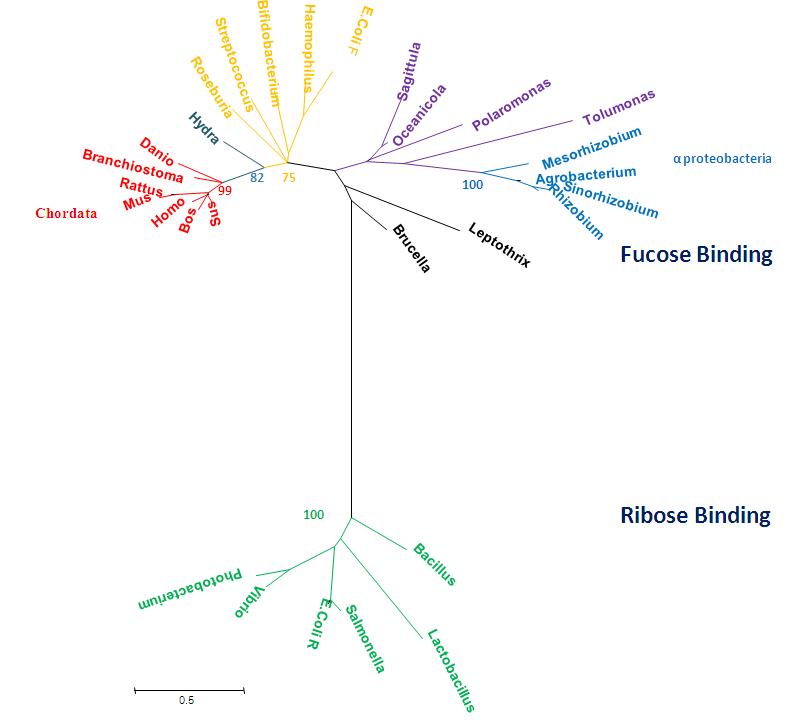
Radiation Tree showing relevant bootstrap values
Figure 5: Phylogenetic tree representing radiation, Obtained using MEGA 4.0 [2] and manually edited in Microsoft Power Point 2007.
Structure
Function
Blast Search
The results from the blast search showed a conserved FucU/RbsD domain.
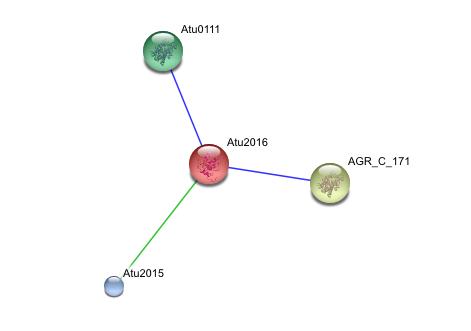
String Analysis
This string analysis shows possible protein - protein interactions. The red protein in the middle represents the target protein and the three surrounding proteins represent interacting proteins.
The larger yellow and green proteins may be releated due to cooccurrence, where as the smaller blue protien may be related due to repeatedly occurring in close proximity to the target protein, however the posible interactions shown in the analysis are inconclusive and unsubstantiated. experimental evidence is needed to prove such relationships
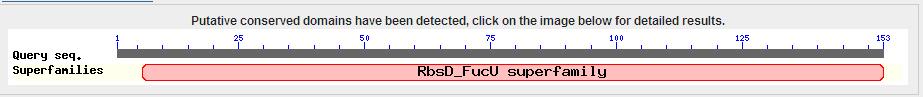
Interpro Scan result
This image shows the results of a InterProScan. The InterProScan search identifies sequence motifs from several databases.These results show that the target protein belongs to the RbsD/FucU superfamily (Purple and red lines) The gray line shows an possible binding histadine site.
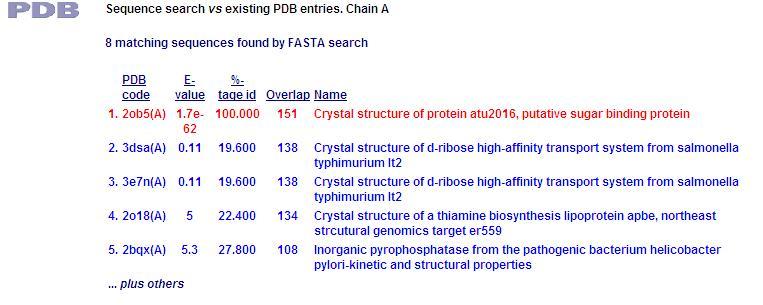
PDB Sequence Match
This PDB scan compares structure alignment. The highest structural match with our sequence is shown to be involved in an d-ribose high affinity transport system. This supports the theory that our hypothetical protein is involved in the ABC transport system of ribose in prokaryote cells.