Pyridoxal Phosphatase Results
Evolution
Basic Local Alignment Search Tool (BLAST)
The BLAST search resulted in a list of organisms to which the target sequence (2cfsA) is related. Complete BLAST results for human pyridoxal phosphatase
Multiple Sequence Alignment
Multiple sequence alignment of the sequences of pyridoxal phosphatase from sample organisms extracted from the blast search resulted in several conserved residues identified as well as some conservative and semi-conservative substitutions.
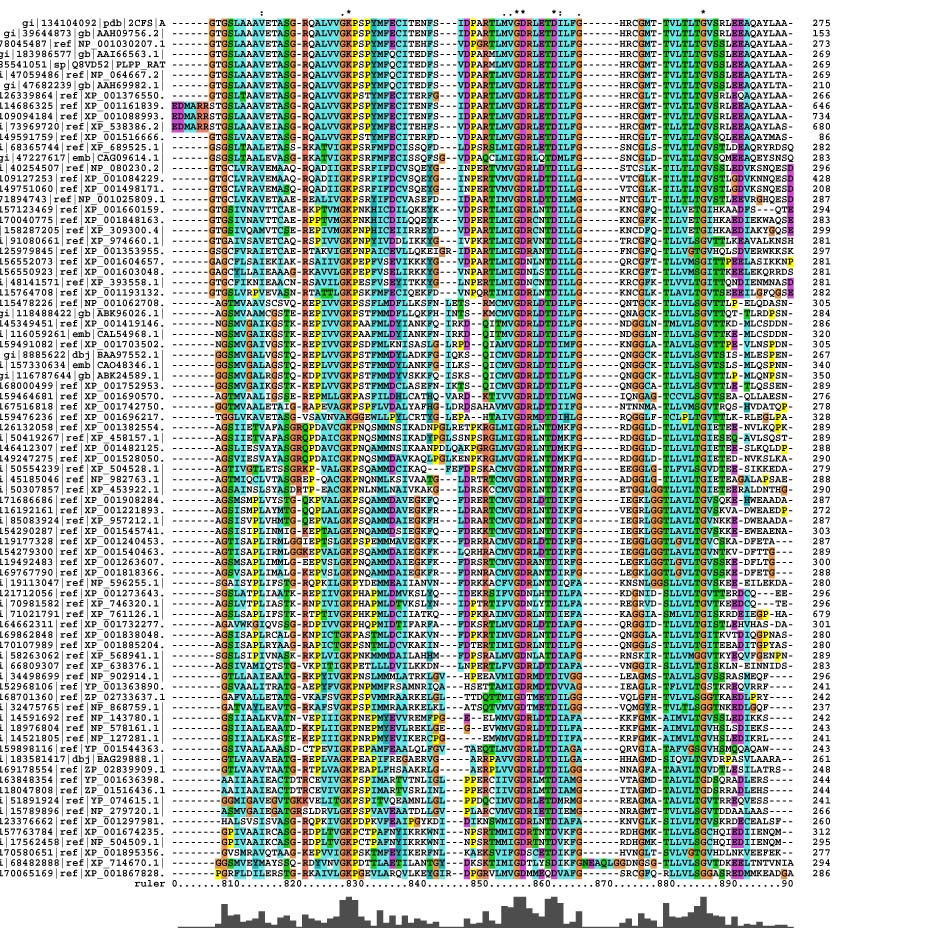
Pyridoxal Phosphatase Multiple Sequence Alignment
It was found that within the multiple sequence alignment, there was only conservation within the 810 to 890 nucleotide region. Outside of that region, there are blocks of similar sequences for some organisms but not for all.
The conserved residues found in the multiple sequence alignment (MSA) correspond to Lys-213, Asp-238, Arg-239 and Gly-260 in the human 2cfsA protein. Two other residues, Val-200 and Leu-245 have conservative substitutions, while Gly-212, Met-235 and Val-236 have semi-conservative substitutions.
Phylogeny Tree
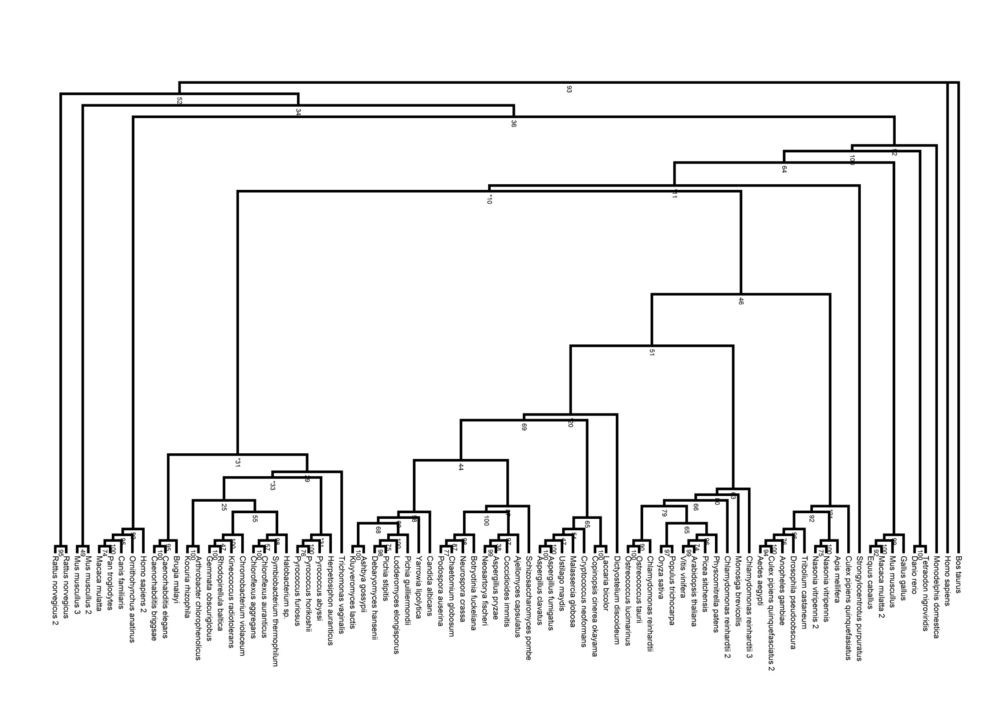
Calculation of relationships between the different pyridoxal phosphatase proteins from the organisms allowed the generation of phylogeny trees. The trees show the divergence of the sequences of the pyridoxal phosphatase-related proteins in various organisms.
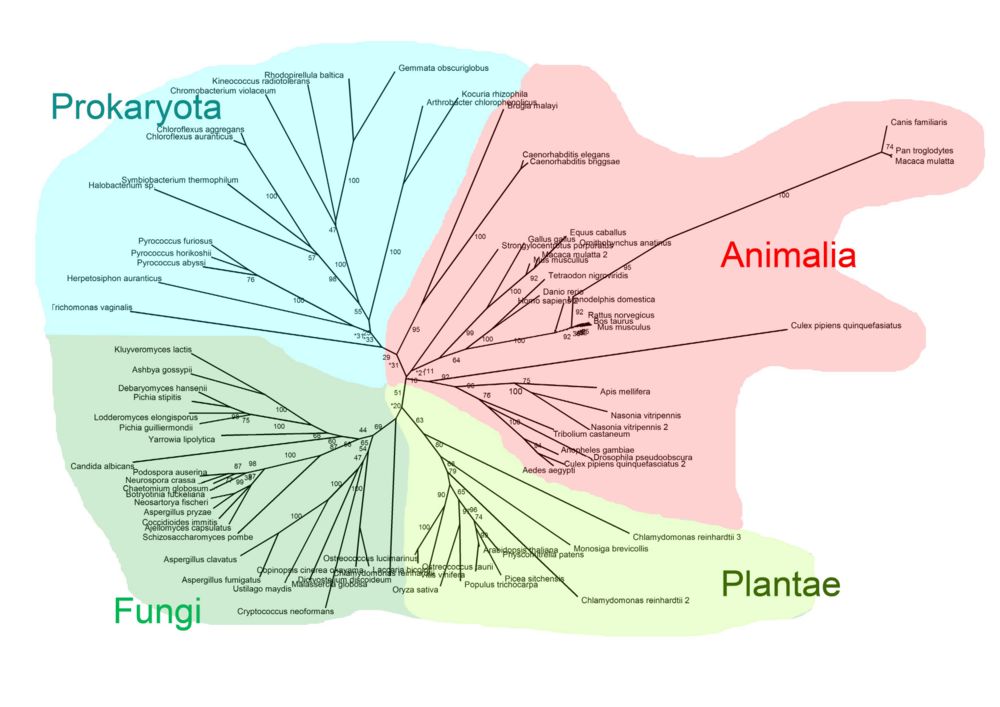
Pyridoxal Phosphatase Phylogeny Diagram
The radial cladogram above, showed that organisms from the same Kingdom tended to clump together in one branch of the tree. Four Kingdoms of organisms with related pyridoxal phosphatase proteins were found; Prokaryota, Fungi, Plantae and Animalia.
Structure
General Structure of Pyridoxal Phosphatase & 2cfsA

PDB
Based on the information provided in the website, Pyridoxal Phosphatase has the following characteristics:
- Isolated from Homo Sapiens and expressed in Escherichia Coli.
- Structure is similar to the Pyridoxal Phosphate Phospatase protein
- 1 (A) Chain
- Magnesium ions (x2)
- Resolution of 2.40 angstroms. This means that the number of sidechains in the wrong rotamer is smaller as compared to proteins of a higher resolution (>2.5 angstroms). Other characteristics of proteins of similar resolutions are: (1) many small detectable errors, (2) correct folding, (3) fewer number of errors in the surface loops and (4) visible water molecules and small ligands.
http://www.rcsb.org/pdb/explore/explore.do?structureId=2cfs
DALI
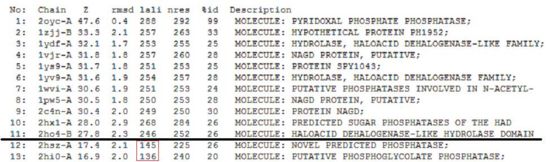
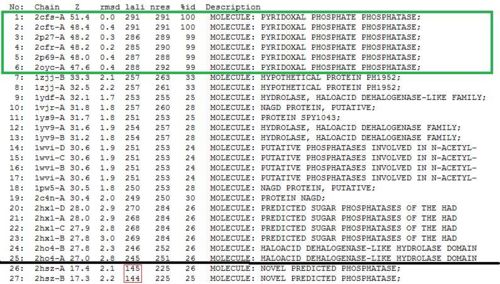
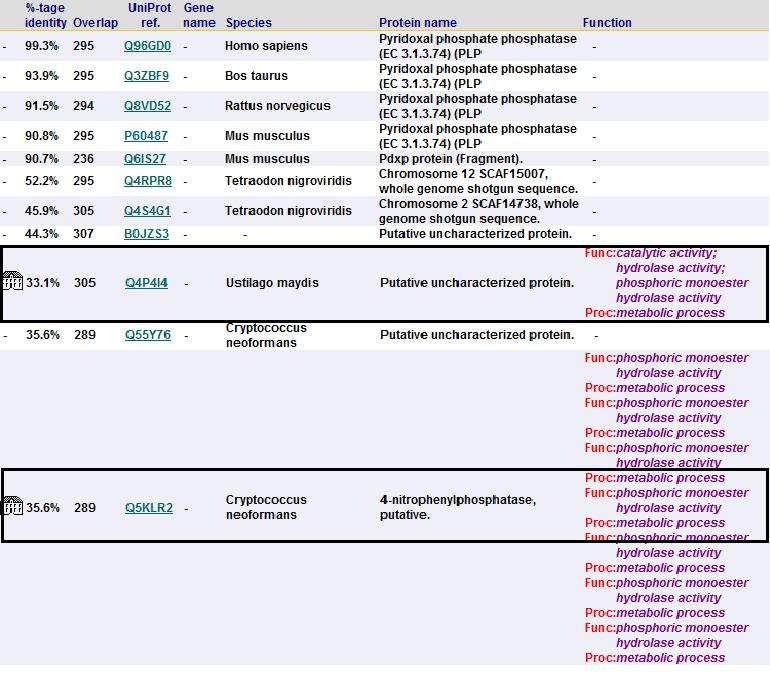
A total of 176 hits were generated, of which only the first 11 (as shown in Fig. 6) were predicted to be significant. The others were rejected on account that their respective lali scores (refer to the red, boxed section) were less than half of Pyridoxal Phosphatase's (nres: 296). The lali value is important as this refers to the number of structurally equivalent residues between the hit protein and the protein-of-interest. If two proteins are not structurally similar/related, chances are they may not have related functions.
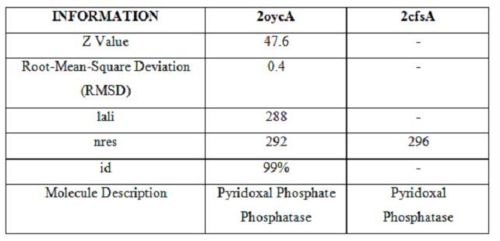
It was noted that none of the hits actually matched 2cfsA. The closest was a protein belonging to the Pyridoxal Phosphate Phosphatase family (PDB ID 2oycA) which based on the results, was predicted to be highly similar to 2cfsA. Table 1 highlights the similarities between 2cfsA and 2oycA.
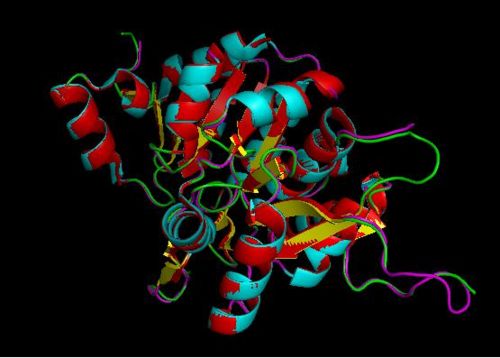
Using the PyMOL software, 2oycA was superimposed against 2cfsA and both structures, as shown in Fig. 7, are structurally similar.
Towards the end of the study, 2cfsA was re-run against the DALI database. The rationale behind doing so was that the initial results did not yield 2cfsA as a protein hit, much less the query protein.
As opposed to the first run, 515 hits were generated. The top 25 protein hits were deemed significant, with the 26th protein hit onwards rejected for the same reason as that of the first batch - that their lali values were too low (less than half of 2cfsA's) for them to be considered as structurally significant to 2cfsA. More importantly was the observation that the DALI database identified 2cfsA as the query protein (unlike the first run). In addition, protein hit numbers 2 (2cftA) - 6 (2oycA) were ALL observed to belong to the Pyridoxal Phosphate Phosphatase family, as was 2cfsA. Based on this, it was easier to narrow down the potential research candidates in terms of structural similarities. Due to time constraint however, not all the protein hits were researched on.
PDBsum
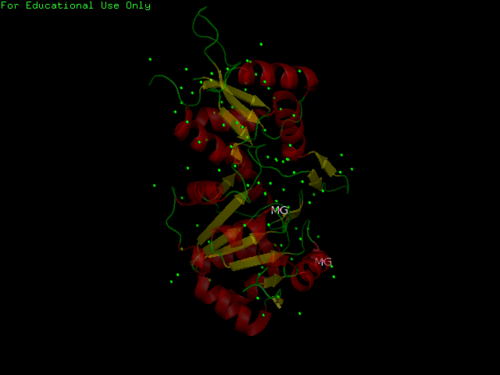
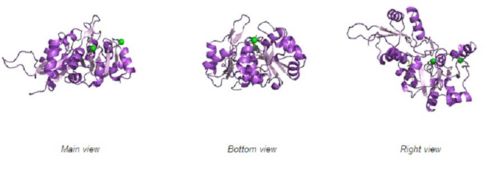
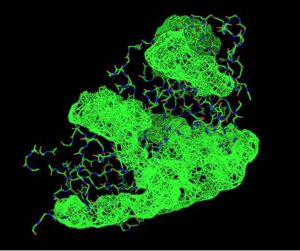
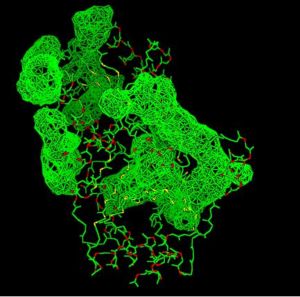
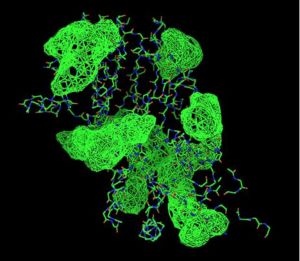
Fig. 8 offers three different views of 2cfsA. The purple chains represent the amino acid chain of 293 amino acids while the green spheres represent the magnesium ions (x2).
By clicking the "Protein chain" link, the user was re-directed to a website containing information pertaining to the secondary structures of both 2cfsA and 2oycA.
Secondary Structures of 2cfsA (L, http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/GetPage.pl) and 2oycA (R).
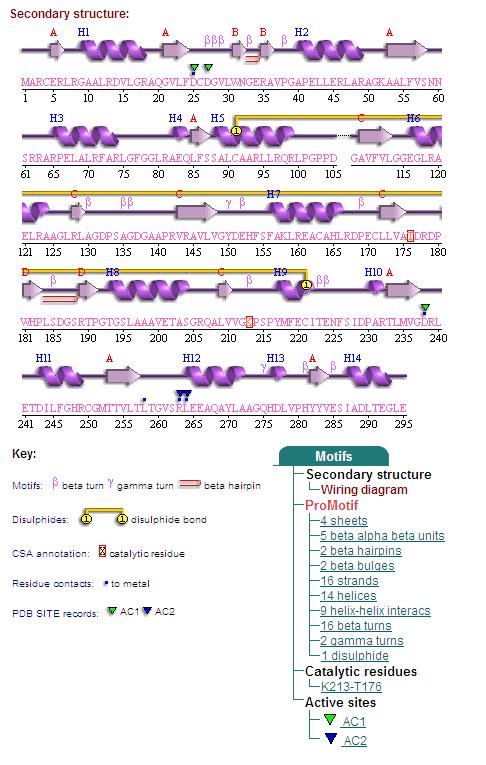
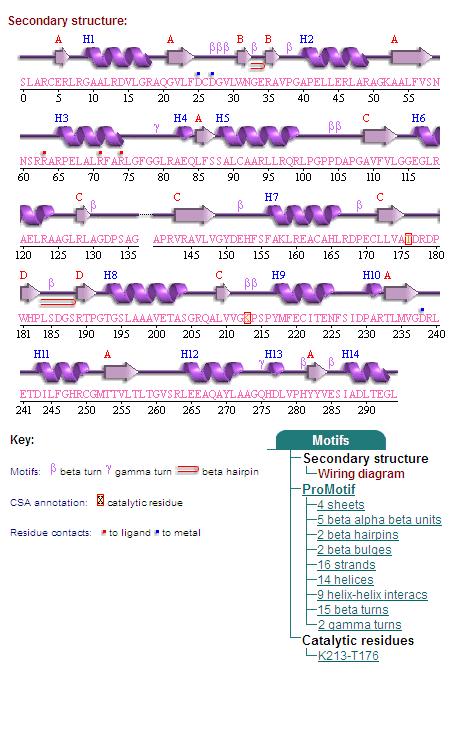
Topology diagrams of 2cfsA (Fig 9.1. http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/GetPage.pl?pdbcode=2cfs&template=protein.html&r=wiring&l=1&chain=A) and 2oycA (Fig 9.2). The topology diagram is a simplified version of the secondary structure information as provided above, and is an indication of the location of the alpha helices (represented by the red cylinders) and beta strands (represented by the pink arrows).
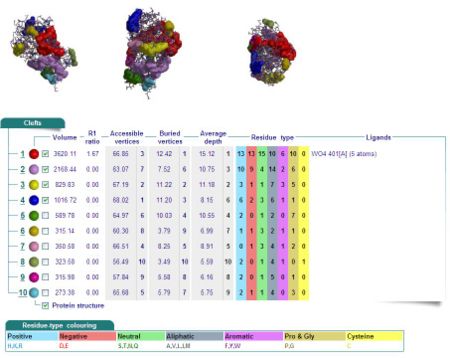
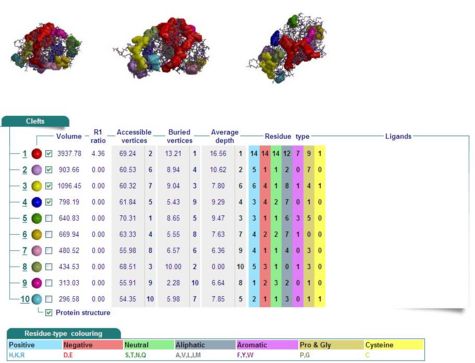
Cleft Analyses of 2cfsA (Fig 10.1. http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/GetPage.pl?pdbcode=2cfs&template=clefts.html&pdbcode=2cfs&r=speedfill) and 2oycA (Fig 10.2).
Results of 2cfsA cleft analysis show that there are no ligands at potential active sites, whereas the same analysis of 2oycA indicates the presence of a WO4 ligand. Another name for 2oycA is chronophin, which has a PLP ligand. It is thus predicted that WO4 is in fact PLP.
Cleft Analysis via PyMol
PROFUNC (2cfsA)
Related Protein Sequences in the PDB (SAS)

Matches to existing PDB Structures
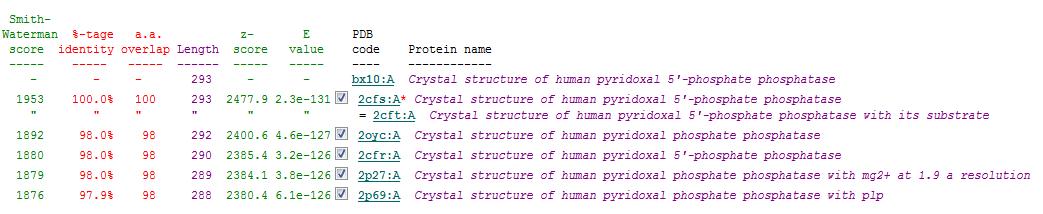
Secondary Structure Matching (SSM)
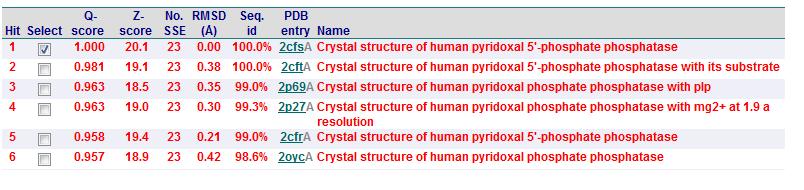
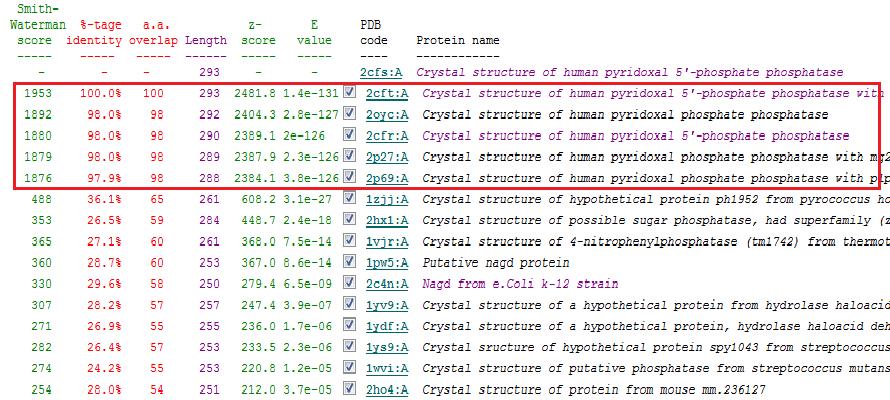
Based on the results obtained via the local alignment method (Smith-Waterman method), it was noted that 2fcsA and 2oycA were indeed similar in terms of sequence and structure. However, contrary to the implications of the DALI-generated results, 2oycA was no longer the most closely-related protein to 2cfsA. That role now belonged to 2cftA. Based on the results generated by Profunc, 2cftA's sequence and structure, as provided by PDB, were identical to that of 2cfsA. The secondary structure matching, however, showed that 2cfsA and 2cftA had differing (albeit minimal) Q scores (1.000 and 0.981 respectively). 2cfsA was described as a "Crystal structure of human pyridoxal 5'-phosphate phosphatase", while 2cftA was described as a "Crystal structure of human pyridoxal 5'-phosphate phosphatase with its substrate". 2oycA, originally the most closely-related protein to 2cfsA, was rated 4 tiers below 2cftA (second in the scoring table).
PROFUNC (2oycA)
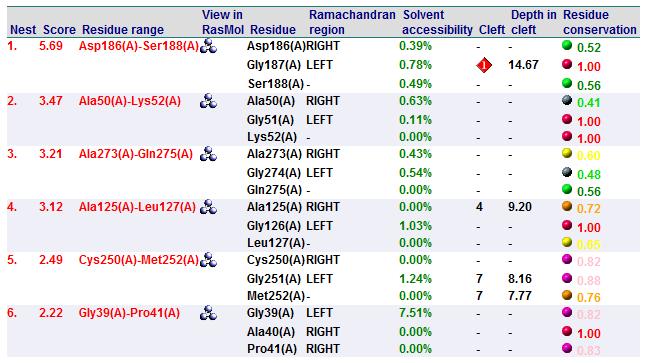
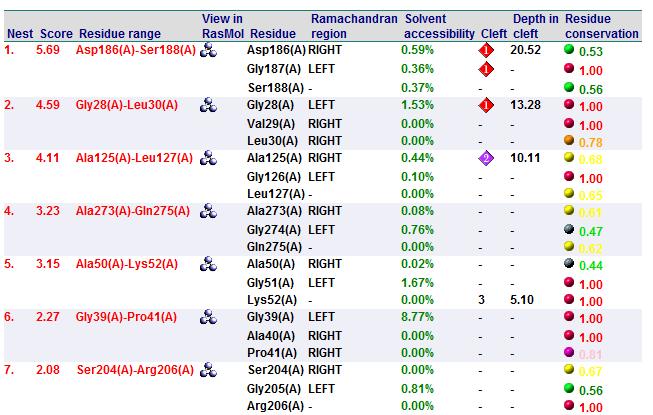
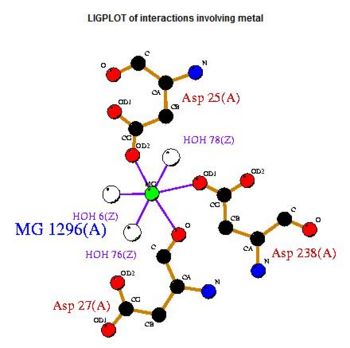
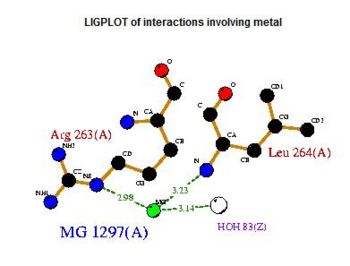
A search on 2oycA was also carried out via PROFUNC, and consistency in the results confirmed that 2cfsA and 2oycA were indeed structurally similar. This lends weight to the original hypothesis that they could be functionally related. In fact, based on the Nest Analysis method, both 2cfsA and 2oycA were observed to be sharing a large number of clefts. The result of the Nest Analysis (2oycA) are as shown below. Notice the similarities between the active sites of both 2oycA and 2cfsA. The reader may need to refer back to the Nest Analysis results for 2cfsA.
In addition, the predicted function of 2oycA was similar to 2cfsA. Based on the Summary of Predicted Function, protein activity is intracellular, involves metabolism on a biological level, and tends to take on the role of a catalyst biochemically. It could also be involved in other biochemical processes such as hydrolase activity, phosphoric monoester hydrolase activity, hydrolase activity (acting on ester bonds) and phosphoric ester hydrolase activity.
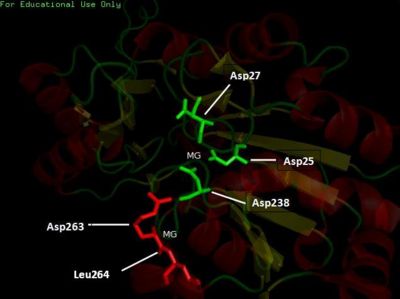
Visualizing the potential catalytic site of 2cfsA
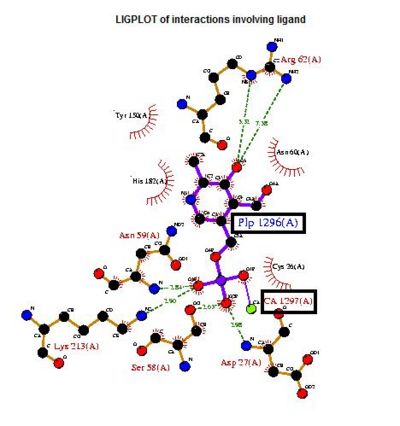
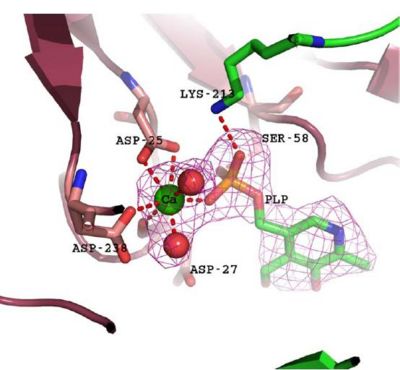
Based on the above-mentioned results, it was concluded that 2cfsA, 2cftA and 2oycA were structurally similar, and that further functional studies could indeed uncover the function of the protein-of-interest: 2cfsA
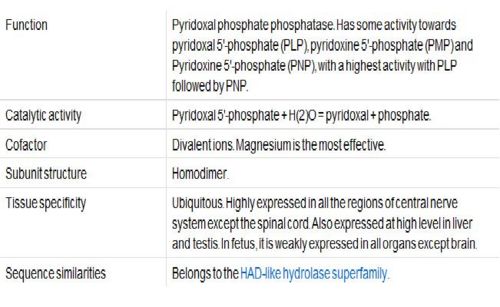
Function
Vitamin B6, Pyridoxal Phosphatase, generally functions as a coenzyme for many reactions and can help facilitate decarboxylation, transamination, racemization, elimination, replacement and beta-group interconversion reactions. The Enzyme Commission (EC; http://www.chem.qmul.ac.uk/iubmb/enzyme/ ) has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. To help predict any unknown functions of Pyridoxal Phosphatase, sequence homology, structural similarities, neighbouring genes, expression level in tissue and protein-protein interactions were analysed.
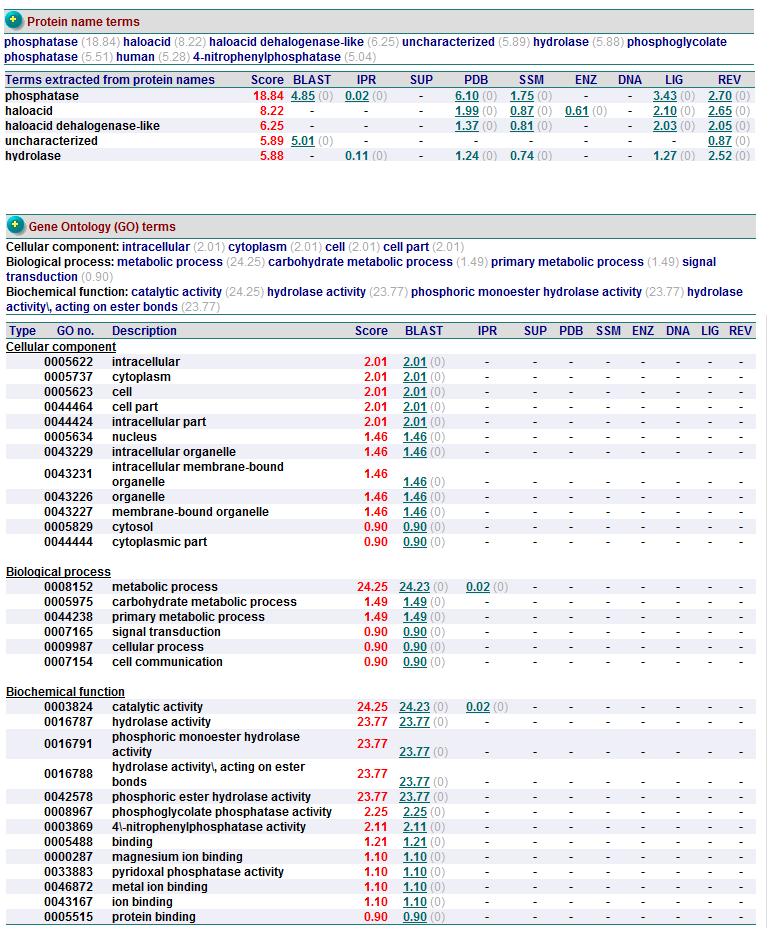
General Annotation
Known Functions
Amino Acid Metabolism
2cfsA is involved in almost all amino acid metabolism. Some of these amino acid metabolism include transamination, transsulfuration, selenoamino acid metabolism and conversion of typtophan to niacin.
Gluconeogenesis
2cfsA can catalyse transamination reactions that are essential for providing amino acids as a substrate for gluconeogenesis. 2cfsA is also required as a coenzyme for glycogen phosphorylase, which in turn catalyse glycogenolysis.
Neurotransmitter Synthesis
Pyridoxal phosphate-dependent enzymes catalyses the biosynthesis of serotonin, epinephrine, norepinephrine and gamma-aminobutyric acid(The four important neurotranmsitters).
Histamine Synthesis
2cfsA is involved in the metabolism of histamine.
Hemoglobin Synthesis and Function
Pyridoxal phosphate aids in the synthesis of heme and can also bind to two sites on hemoglobin to enhance the oxygen binding of hemoglobin.
Gene Expression
Pyridoxal phosphate has been implicated in increasing or decreasing the expression of certain genes. Increased intracellular levels of 2cfsA will lead to a decrease in the transcription of glucocorticoid hormones. Also, vitamin B6 deficiency will lead to the increased expression of albumin mRNA. Also, pyridoxal phosphate will influence gene expression of glycoprotein IIb by interacting with various transcription factors. The result is inhibition of platelet aggregation.
Sequence homology
To check for function among similar homologs, 2cfsA sequence was run against FASTA, Pfam and PSI Blast non-redundant database. A search using Pfam and BLAST revealed that 2cfsA belonged to the superfamily, Haloacid dehalogenase-like hydrolase. It is noted that the top 15 search results from BLAST database all belong to the same superfamily, HAD-like hydrolase. Results from FASTA (below) also showed five results with almost identical sequence, all being HAD-like hydrolases.
http://blast.ncbi.nlm.nih.gov/Blast.cgi
http://pfam.sanger.ac.uk/search/sequence/results?jobId=A74388CE-361C-11DD-9F74-DFE079E860E8
Structural Similarities
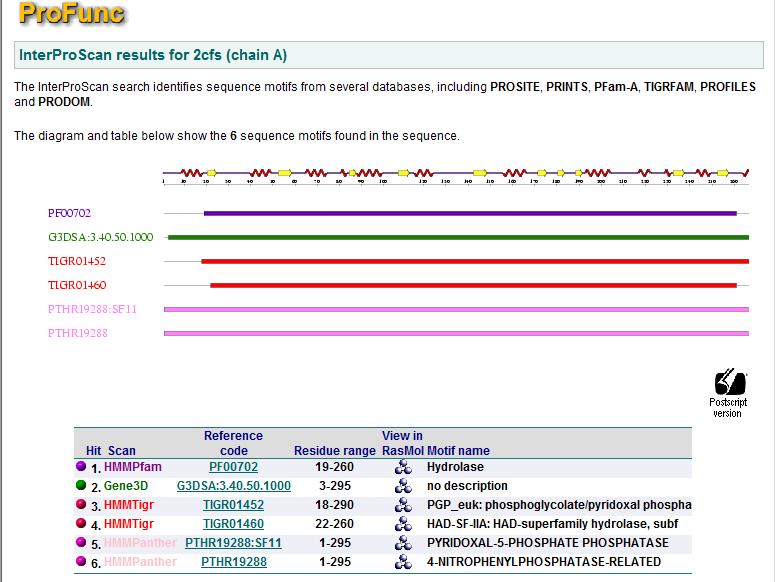
A search based on motifs resulted in six hits. Five of which are hydrolases and one with function unknown. Proteins with similar motifs does not suggest any other function for 2cfsA.
Location of Gene on Genome
It was noted that neighbouring genes also belongs to the same super family, HAD-like hydrolase.
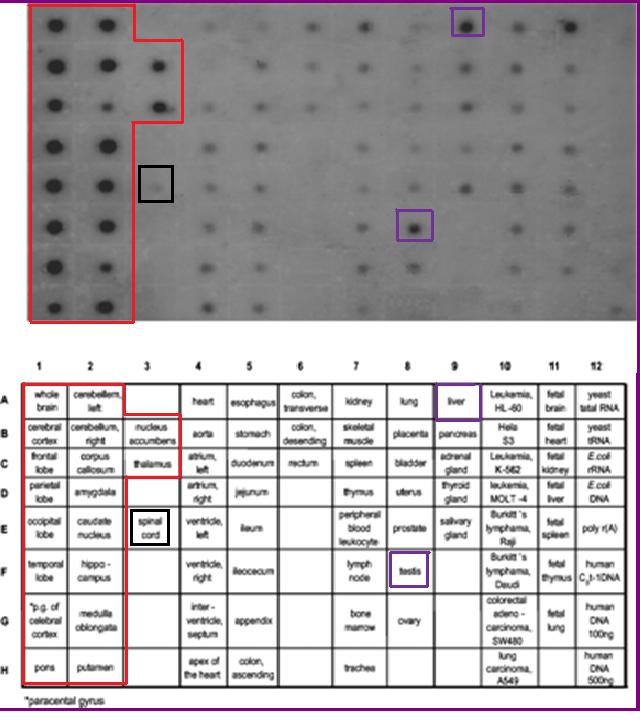
Functional Expression in Tissue
The mRNA expression levels of 2cfsA is experimentally proven to be ubiquitous. This is consistent with its essential role in cellular metabolism. However, 2cfsA may have a more specific function in the brain, where the expression level is substantially higher.(Young M. J. et al, 2003)
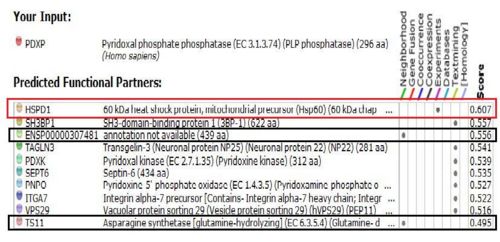
Protein-Protein Interactions
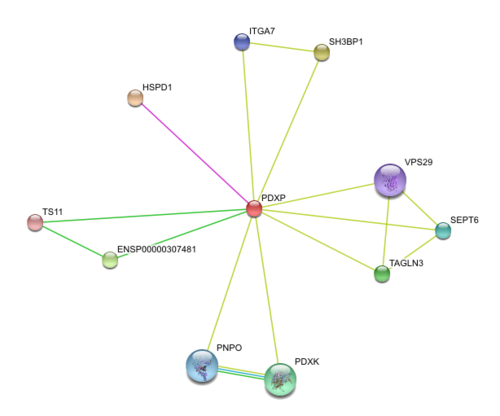
Results from STRING showed 11 predicted functional partners with a medium confidence score of >0.4. Of these 11, putative homologs of HSPD1, a chaperonin, was found to interact with PDXP in E coli K12, two were neighbour genes in 16 and 18 different genomes and eight were found by textmining. The eight were not considered due to lack of evidence. Homolog interactions suggests that 2cfsA maybe a cofactor of HSPD1 and play a role in protein folding. The search also revealed that TS11 is a neighbour gene for 2cfsA for 16 different genomes, hence it is possible that 2cfsA may actually bind to TS11 and function as a coenzyme for asparagine synthetase.
http://string.embl.de/newstring_cgi/show_network_section.pl?taskId=ph89gQxcLucL&allnodes=1
back to Pyridoxal Phosphatase main page