Results - NANP
Results
Query Sequence
The amino acid query sequence of 2gfh protein (Figure 3) from Mus musculus is obtained from Genbank.
| <1 mgsdkihhhh hhmglsrvra vffdldntli dtagasrrgm levikllqsk yhykeeaeii
61 cdkvqvklsk ecfhpystci tdvrtshwee aiqetkggad nrklaeecyf lwkstrlqhm 121 iladdvkaml telrkevrll lltngdrqtq rekieacacq syfdaivigg eqkeekpaps 181 ifyhccdllg vqpgdcvmvg dtletdiqgg lnaglkatvw inksgrvplt sspmphymvs 241 svlelpallq sidckvsmsv> |
Figure 3. The 260 amino acid sequence of 2gfh protein.
Sequence Homology
From the BlastP similarity was used for comparison as these had shown higher homology to the query sequence sequence search, a total of 500
proteins were yielded.Only a total of 38 proteins, in contrast with the remainder of the search results.These proteins were chosen according to
their bit scores and E-values. Two more outlier partial sequences contributing to poor overall alignment (huge deletion gaps) were subsequently
removed. The remaining 36 sequences were used for the generation of the phylogenetic tree (and bootstrapped tree as well).
Multiple Sequence Alignment
The following multiple sequence alignment (MSA) was obtained (Figure 4). From the alignments, gi|10888xy and
gi|10888yz are representative of gi|108881764 and gi|108881765 respectively. Both these
hypothetical proteins belong to the mosquito Aedes aegypti.
The identifier numbers for these two proteins were initially changed to an alpha-numeric one, due to the inability of Phylip to generate a tree
from the original identifiers. This was due to the fact that the programme only took the first five numeric digits (10888), thereby resulting
in a programme error prompt which listed both proteins as duplicates (from the identifier numbers). Both these identifiers were subsequently
renamed for the final phylogenetic tree.
Figure 4. MSA of query (top-most sequence – No.1) and related sequences.
From the MSA, it can be observed that there are generally slight domain conservations throughout the protein sequences. Small insertion and
deletion gaps were noticeable along the alignment as well. A particularly large insertion gap was observed between amino acids 91 to 114.
The organisms with the large insertion gaps were as identified below:
Bacillus licheniformis
Bacillus subtilis
Bacillus halodurans
Bacillus clausii
Symbiobacterium thermophilum
A highly conserved (with invariant) section of amino acids (LV)–(LVA)–(LIV)–(LIV)-T-N-G was observed in all the sequences from amino acid 211
to 217 in the alignment. Downstream of this conserved portion of genes are 5 more invariant positions (1 or 2 amino acids in length).From these
short conservation regions, the functions or even structure of the encoded proteins could have significance in its evolutionary pattern.
Phylogenetic Tree
The tree was plotted to obtain the phylogenetic lineage (Figure 5).
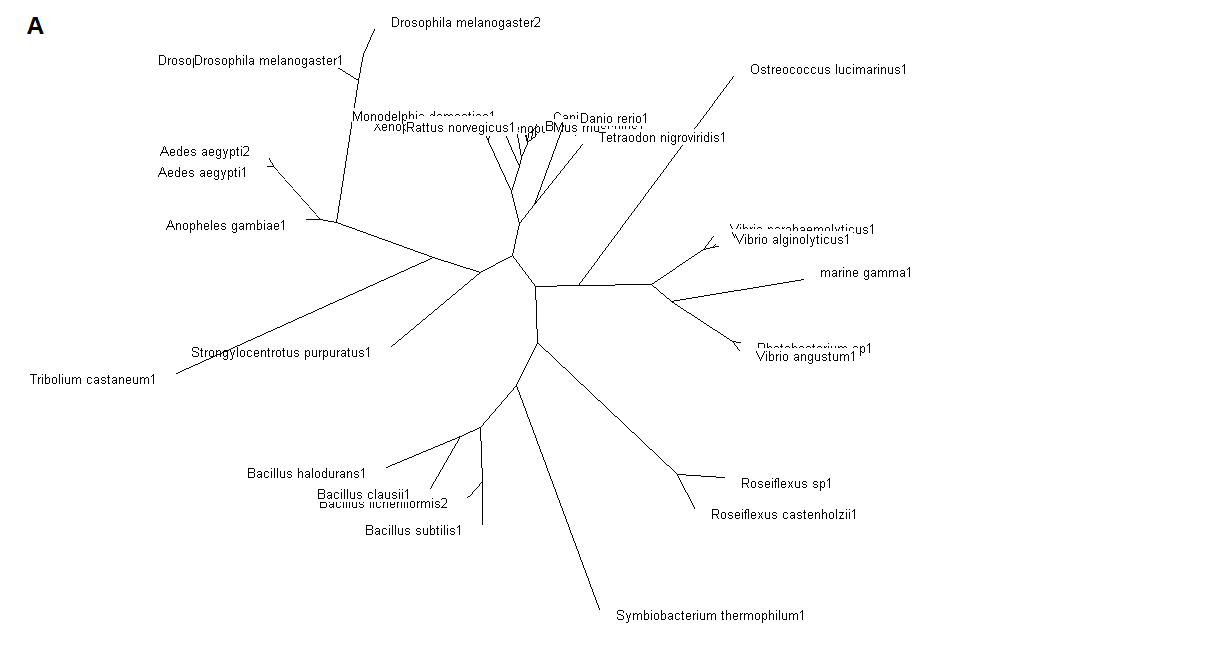
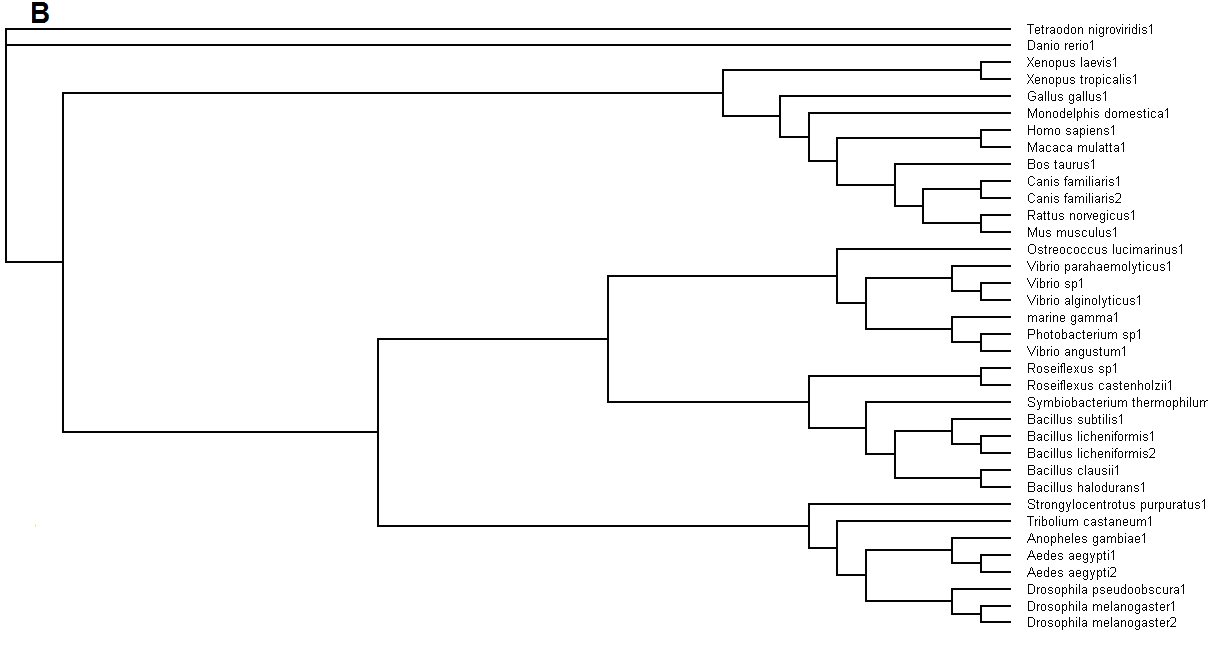
Figure 5. (A) Phylogenetic tree showing organisms with related protein sequence homology in Radial Tree view. (B) Rectangular
Cladogram view with related protein sequence homology.
From the Rectangular Cladogram view, it could be observed that there are four distinct separate groups involving fishes, mammals (where the
query protein is also mapped), bacteria and insects.
Bootstrapping
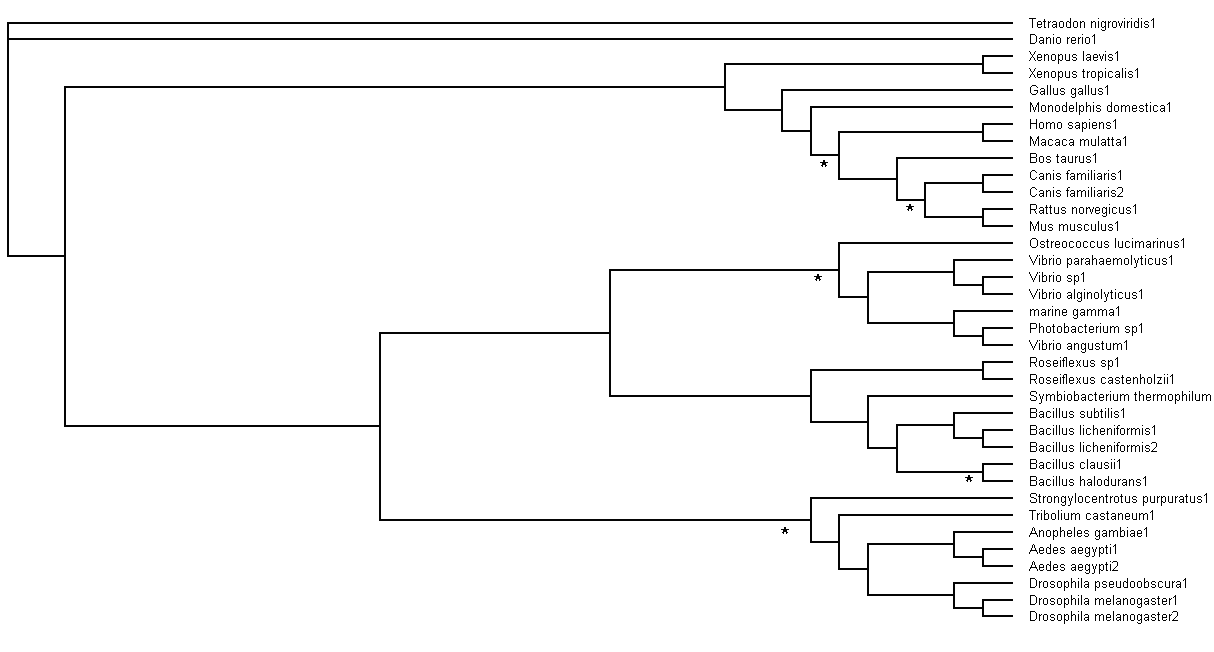
Bootstrapping values obtained were analysed. Branch values occurring below 75% (<75%) would be indicated by an asterisk (*),
as shown in Figure 6.
Figure 6. Branch bootstrap values in Rectangular Cladogram view. Branches with strap values <75% were indicated with
asterisks (*)
DALI Searching
SUMMARY: PDB/chain identifiers and structural alignment statistics NR. STRID1 STRID2 Z RMSD LALI LSEQ2 %IDE REVERS PERMUT NFRAG TOPO PROTEIN
1: 3033-A 2gfh-A 41.1 0.0 246 246 100 0 0 1 S HYDROLASE haloacid dehalogenase-like hydrolase domain 2: 3033-A 1fez-A 18.1 3.5 178 256 22 0 0 13 S HYDROLASE phosphonoacetaldehyde hydrolase (bacillus c 3: 3033-A 2hsz-A 17.9 3.3 168 222 23 0 0 13 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION novel predicted 4: 3033-A 1qq5-A 17.3 3.1 198 245 19 0 0 12 S HYDROLASE l-2-haloacid dehalogenase (xanthobacter aut 5: 3033-A 1o03-A 17.0 5.0 188 221 20 0 0 11 S ISOMERASE beta-phosphoglucomutase (lactococcus lactis 6: 3033-A 2b0c-A 16.4 2.6 184 199 20 0 0 13 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION putative phospha 7: 3033-A 2fdr-A 15.8 4.4 190 214 19 0 0 15 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION conserved hypoth 8: 3033-A 2p11-A 15.7 2.9 194 211 16 0 0 20 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 9: 3033-A 1te2-A 15.7 3.6 170 211 19 0 0 15 S HYDROLASE putative phosphatase (escherichia coli o157 10: 3033-A 1yns-A 15.3 4.0 169 254 11 0 0 13 S HYDROLASE e-1 enzyme (enolase-phosphatase e1) (homo s 11: 3033-A 1qyi-A 15.0 3.5 198 375 19 0 0 17 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 12: 3033-A 2i6x-A 14.9 3.1 176 199 19 0 0 18 S HYDROLASE hydrolase, haloacid dehalogenase-like family 13: 3033-A 1u7p-A 14.3 2.9 144 164 18 0 0 14 S HYDROLASE magnesium-dependent phosphatase-1 (mdp-1) ( 14: 3033-A 1ymq-A 14.1 2.3 130 260 16 0 0 14 S TRANSFERASE sugar-phosphate phosphatase bt4131 (bacte 15: 3033-A 1j8d-A 13.1 2.5 141 180 11 0 0 12 S HYDROLASE deoxy-d-mannose-octulosonate 8-phosphate ph 16: 3033-A 2ho4-A 12.9 2.4 131 246 19 0 0 14 S HYDROLASE haloacid dehalogenase-like hydrolase domain 17: 3033-A 1pw5-A 12.7 2.3 136 246 21 0 0 12 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION nagd protein, pu 18: 3033-A 1nf2-A 12.7 2.6 127 267 13 0 0 11 S STRUCTURAL GENOMICS/UNKNOWN FUNCTION phosphatase (the 19: 3033-A 1rlm-A 12.4 2.8 131 269 13 0 0 14 S HYDROLASE phosphatase Mutant (escherichia coli) bacte 20: 3033-A 1f5s-A 12.1 3.5 159 210 14 0 0 15 S HYDROLASE phosphoserine phosphatase (psp) (methanoco 21: 3033-A 1cr6-B 12.0 3.8 177 541 18 0 0 18 S HYDROLASE epoxide hydrolase (mus musculus) mouse expr 22: 3033-A 1rku-A 11.9 3.6 172 206 11 0 0 18 S TRANSFERASE homoserine kinase (pseudomonas aeruginosa 23: 3033-A 2b30-A 11.8 2.7 134 284 16 0 0 12 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION pvivax hypotheti 24: 3033-A 1kyt-A 10.5 2.5 122 216 13 0 0 15 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 25: 3033-A 2o2x-A 10.3 3.6 139 204 17 0 0 14 S STRUCTURAL GENOMICS, UNKNOWN FUNCTION hypothetical pro 26: 3033-A 1u02-A 10.1 2.7 128 222 16 0 0 12 S STRUCTURAL GENOMICS trehalose-6-phosphate phosphatase 27: 3033-A 2fea-A 10.0 3.5 167 219 7 0 0 21 S HYDROLASE 2-hydroxy-3-keto-5-methylthiopentenyl-1- pho 28: 3033-A 2hx1-A 9.6 3.2 130 275 24 0 0 19 S HYDROLASE predicted sugar phosphatases of the had supe 29: 3033-A 1mh9-A 9.2 3.2 146 194 15 0 0 15 S HYDROLASE deoxyribonucleotidase (mitochondrial 5'(3')-
Figure 7. The DALI search results that were returned through e-mailed. The highlighted (yellow) shows the query protein. With a z value
of 41.1 and a root mean standard deviation of 0.0 and %IDE of 100, shows that it is a HAD family protein. The highlighted (green) shows
significant similarities of query protein as a hydrolase phosphatase as Z values are more then 1, RMSD still of low values and %IDE of more
then 20.Z
From the DALI search (Figure 7), Neu5Ac phosphatase is a haloacid dehalogenase-like hydrolase. This family is structurally different from the
alpha/ beta hydrolase family. It has L-2-haloacid dehalogenase, epoxide hydrolases and phosphatases. This family consists of two domains of
structure. One is an inserted four helix bundle, which is the least well conserved region of the alignment, between residues 16 and 96 of (S)-2-
haloacid dehalogenase I. The remaining of the fold is composed of the core alpha/beta domain. It is classified as a hydrolase found in mouse.
The chemical components would be phosphate ion, sodium ion, 1,2-ethanediol, chloride ion. PO4 and EDO are ligands while
Na and Cl are metals.
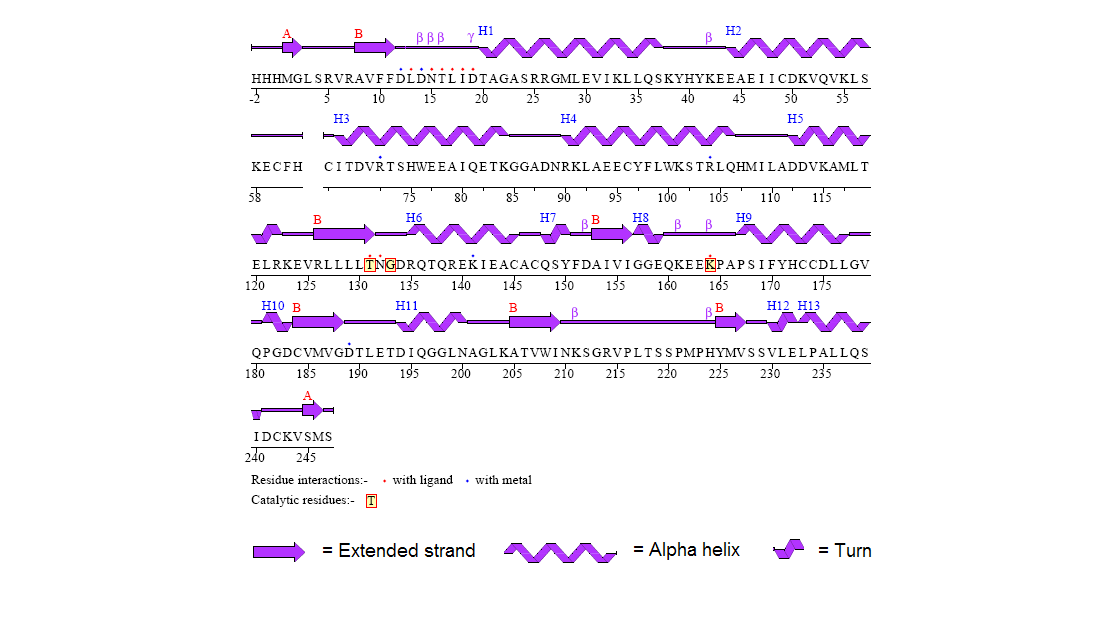
Protein Structure
Figure 8. Secondary structure of 2gfh protein with residue interaction and the catalytic residues marked out in red boxes. (http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/CheckCode.pl)
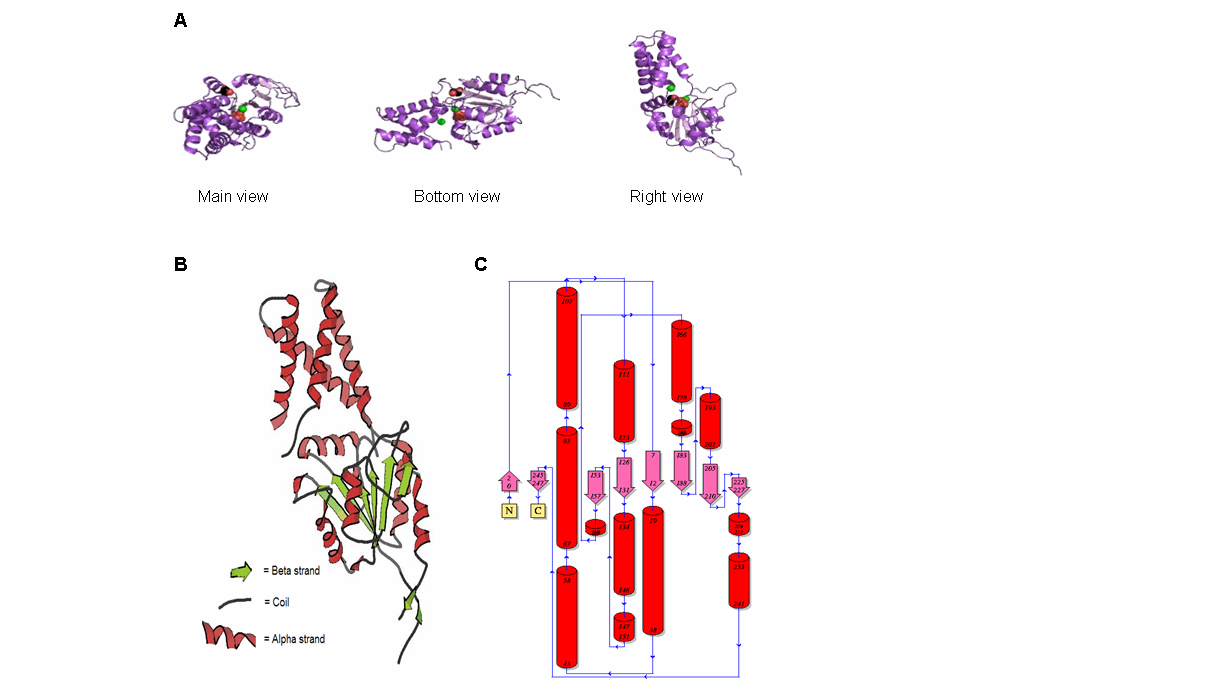
Figure 9. (A) Main, bottom and right view of 2gfh protein, the spheres represent the element/chemical components. (B) 2gfh protein viewed using KiNG. (C) Topology diagram of 2gfh showing the beta and alpha strand. (http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum/CheckCode.pl)
The structure of 2gfh protein was determined to a be polypeptide(L) with 260 residues. Secondary structure (Figure 8) comprises of 56% helical
(13 helicals; 146 residues) and 11% beta sheet (8 strands; 31 residues)
Protein Folding
Table 1. Matching folds detected by SSM and Dali, with scores values between the Neu5Ac-9-P phosphatase and other proteins.(http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/profunc/GetResults.pl?source=profunc&user_id=bb32&code=143144)
| Hit | Z-score | No. SSE |
RMSD (Å) |
Sequence Id |
PDB entry |
Name |
| 1 | 16.6 | 16 | 0.00 | 100.0% | 2gfhA | Crystal structure of protein c20orf147 homolog (17391249) from Mus musculus at 1.90 a resolution |
| 2 | 9.4 | 16 | 2.34 | 23.0% | 1x42A | Crystal structure of a haloacid dehalogenase family protein (ph0459) from Pyrococcus horikoshii ot3 |
| 3 | 9.2 | 10 | 1.63 | 26.0% | 1swwA | Crystal structure of the phosphonoacetaldehyde hydrolase d12a mutant complexed with magnesium and substrate phosphonoacetaldehyde |
| 4 | 9.3 | 10 | 1.66 | 26.0% | 1swvA | Crystal structure of the d12a mutant of phosphonoacetaldehyde hydrolase complexed with magnesium |
| 5 | 7.1 | 11 | 1.75 | 24.4% | 1fezA | The crystal structure of Bacillus cereus phosphonoacetaldehyde hydrolase complexed with tungstate, a product analog |
| 6 | 6.3 | 12 | 2.34 | 20.5% | 2p11A | Crystal structure of hypothetical protein (yp_553970.1) from Burkholderia xenovorans lb400 at 2.20 a resolution |
| 8 | 7.5 | 11 | 1.96 | 26.8% | 1rqlA | Crystal structure of phosponoacetaldehyde hydrolase complexed with magnesium and the inhibitor vinyl sulfonate |
| 9 | 7.3 | 11 | 1.96 | 26.8% | 1rqnA | Phosphonoacetaldehyde hydrolase complexed with magnesium |
| 10 | 6.7 | 13 | 2.44 | 22.1% | 2b0cA | The crystal structure of the putative phosphatase from Escherichia coli |
The high score values between Neu5Ac phosphatase and the other proteins (Table 1), proving that the folding of the different proteins match.
The Z-score measures the statistical significance of a match in terms of standard Gaussian statistics. It is based on the quality of the match
between the query and target structures and assumes a Gaussian distribution of quality scores would be obtained from a large enough databases
of protein structures. The higher the Z-score, the higher is the statistical significance of the match is the number of matched secondary
structure elements, examples; helices and strands between the two structures.
Sequence Similarity
|
Hydrolase: domain 1 of 1, from 18 to 224: score 96.2, E = 1e-25 *->ikavvFDkDGTLtdgkeppiaeaiveaaaelgl.........lplee
++av+FD+D+TL+d+ + + ++ + e+ ++l + + +++ ++ +
query 18 VRAVFFDLDNTLIDT-AGASRRGMLEVIKLLQSkyhykeeaeIICDK 63
vekllgrgl.g.erilleggltaell...................d.evl
v l +++ ++ ++ t ++ + +++++ ++++ ++ ++
query 64 VQVKLSKECfHpYSTCITDVRTSHWEeaiqetkggadnrklaeecYfLWK 113
glial.dklypgarealkaLkrrGikvailTggdr.naeallealgla.l
++ ++ l +++++ l +L++ +++ +lT+gdr++++++ ea+++ ++
query 114 STRLQhMILADDVKAMLTELRKE-VRLLLLTNGDRqTQREKIEACACQsY 162
fdviidsdevggvgpivvgKPkpeifllalerlgvkpeevgpevlmVGDg
fd+i++++e + KP+p if + ++ lgv+p ++ +mVGD+
query 163 FDAIVIGGEQK------EEKPAPSIFYHCCDLLGVQPGDC----VMVGDT 202
vnDapalaa.AGv.gvamgngg<-*
+ +++ + +AG+++++++n +
query 203 LETDIQGGLnAGLkATVWINKS 224
|
Figure 10. The alignments of the top-scoring domains of 2gfh protein (query) using Pfam 21.0 (Janelia Farm). (http://pfam.janelia.org)
A search of using Pfam (Figure 10) matched the query sequence in this case Neu5Ac-9-P phosphatase with hydrolase. The E value of 1e-25 gives
significant results proving that it is not by chance nor random that the match made was a hydrolase.
Surface Properties
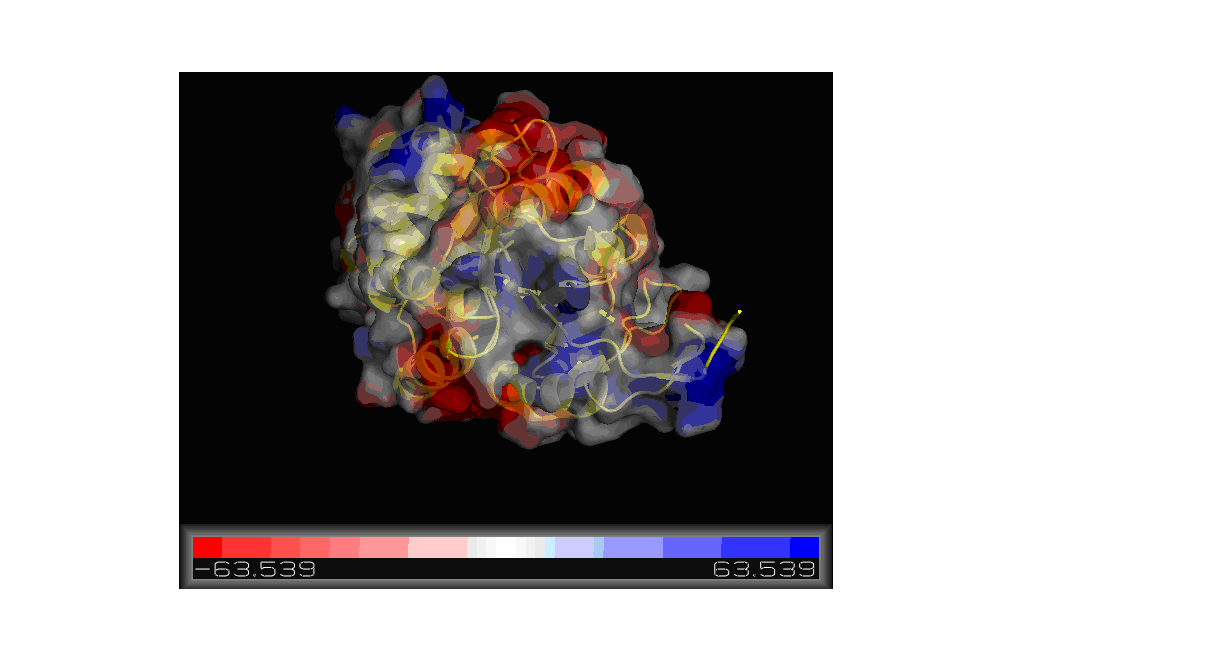
Figure 11. Molecular surface of 2gfh colored by electrostatic potential shown using Pymol.
Using the PDB file name 2gfh, a model was constructed using Pymol showing the electrostatic potential of the molecular surface. As shown in
Figure 11, the red color portions are negatively charged while the blue would be positively charged region. The charge ranges from -63.539 to
63.539.
Figure 12. (A) Molecular structure of 2gfh showing the possible binding sites with the different colors represent classes of amino
acids. (B) Results from Profunc show that 2gfh comprises of 2 ligands: phostphate ion (PO4) and ethylene glycol (EDO).
Profunc helps to identify the likely biochemical function of a protein from its 3 dimensional (3D) structure. It uses fold matching, residue
conservation, surface cleft analysis, and functional 3D templates, to identify both the protein’s likely active site and
possible homologues in the PDB. The search provided information on the possible binding sites and important identification of potential ligands
like PO4 and EDO. Based on comprehension and research, EDO (Figure 14) could most likely be a chemical compound widely used to
crystallize protein from its native form and used as automotive antifreeze. Finding of the PO4 ligand (Figure 13) was important as
it would most likely be an active site. As Neu5Ac-9-P phosphatase is a hydrolase, the PO4 could well be involved in the mechanism
and function of the protein.
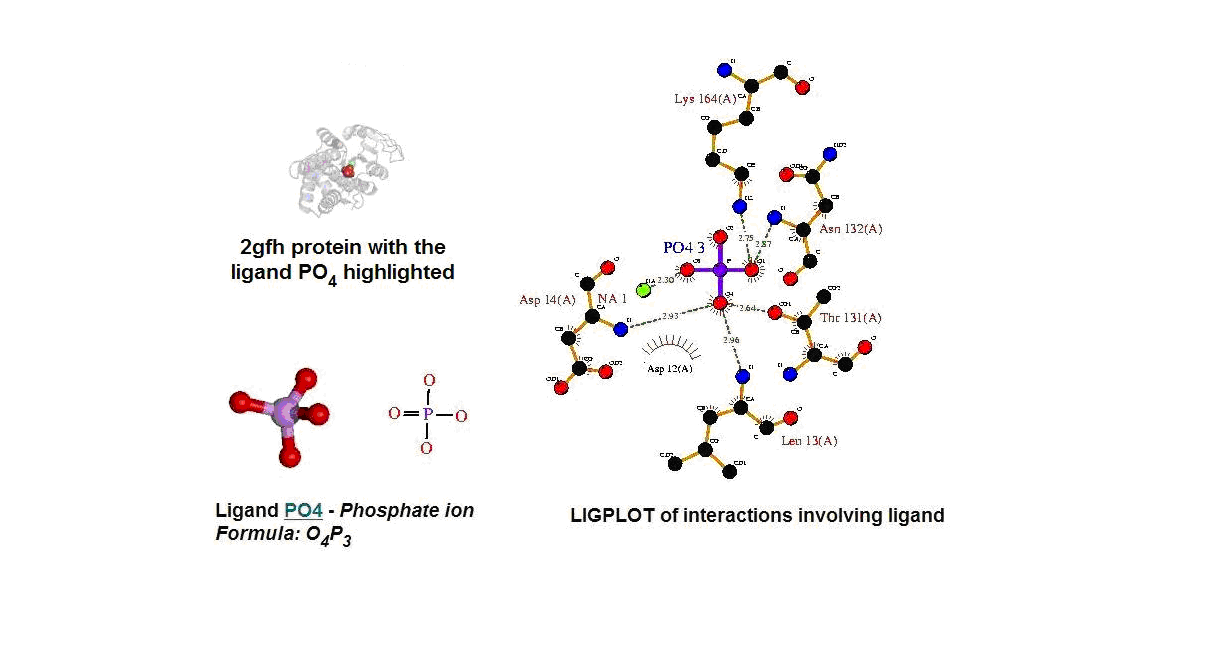 Figure 13. (A) Molecular structure of 2gfh with the ligand PO4. (B) Molecular and chemical structure of PO4. (C) Ligand interaction involving PO4.
Figure 13. (A) Molecular structure of 2gfh with the ligand PO4. (B) Molecular and chemical structure of PO4. (C) Ligand interaction involving PO4.
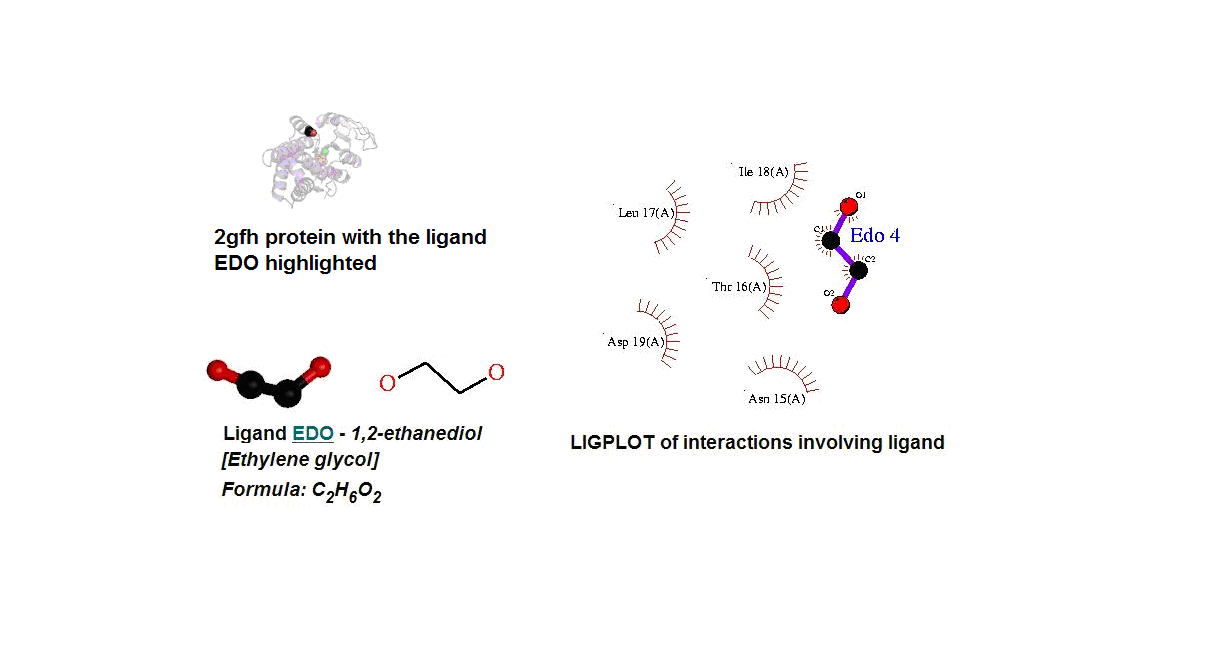 Figure 14. (A) Molecular structure of 2gfh with the ligand EDO. (B) Molecular and chemical structure of EDO. (C) Ligand interaction involving EDO.
Figure 14. (A) Molecular structure of 2gfh with the ligand EDO. (B) Molecular and chemical structure of EDO. (C) Ligand interaction involving EDO.
Table 2. Number 3 shows siginificant scores implying possible convserved residues in N-acetylneuraminic acid phosphatase.
| No | Score | Number of residues |
Cleft | Average accessibility | Average conservation | Residues |
| 1 | 3.770 | 3 | 3 | 2 | 0.437 | Ser212(A), Gly213(A), Arg214(A) |
| 2 | 3.579 | 3 | 3 | - | 0.913 | Ala201(A), Gly202(A), Leu203(A) |
| 3 | 3.483 | 3 | 3 | - | 0.816 | Leu177(A), Gly178(A), Val179(A) |
| 4 | 3.000 | 3 | 3 | 2 | 1.000 | Asn15(A), Thr16(A), Leu17(A) |
| 5 | 0.646 | 4 | 4 | - | 0.646 | Cys145(A), Ala146(A), Cys147(A), Gln148(A) |
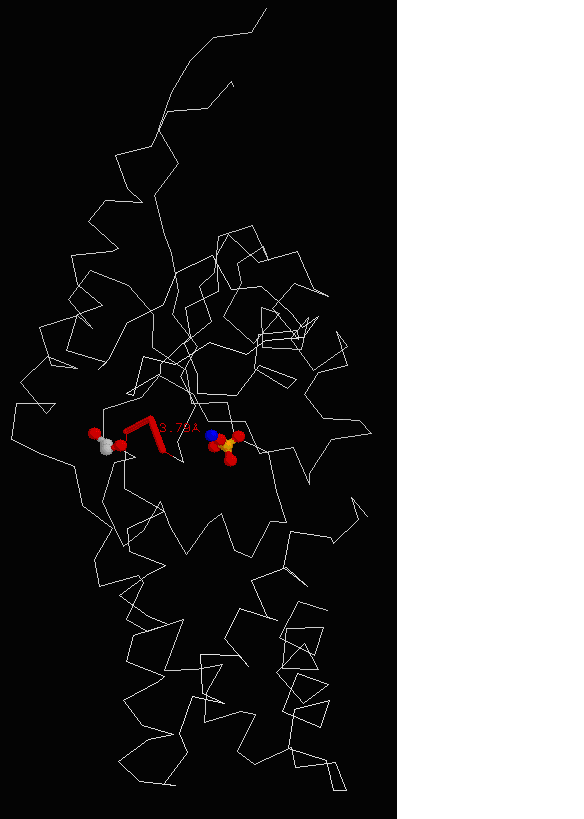
Figure 15. Molecular structure of Neu5Ac-9-P was determined using RasMol, showing the conserved region of asparagine, threonine and
leucine with EDO molecule in grey and PO4 in yellow.
Profunc also provided information of the conserved residues in Neu5Ac-9-P phosphatase. By using nest analysis whereby, nests are structural
motifs that are often found in functionally important regions of protein structures and given a score value. When a score is above 2.0, it
implies that the nest is a functionally significant one. The results were tabulated showing the nest’s start and end residues
residues making up the nest. Residue conservation was given to each nest residue. The score ranges from 0.0 to 1.0 which signifies that the
residue is not at all conserved or perfectly conserved respectively. It is determined from a multiple sequence alignment of the
protein’s sequence against BLAST hits from UniProt sequence database. Results (Figure 15) show 2 highly conserved
region asparagine, threonine and leucine as the residue conservation score was 1.0.
Functional analysis
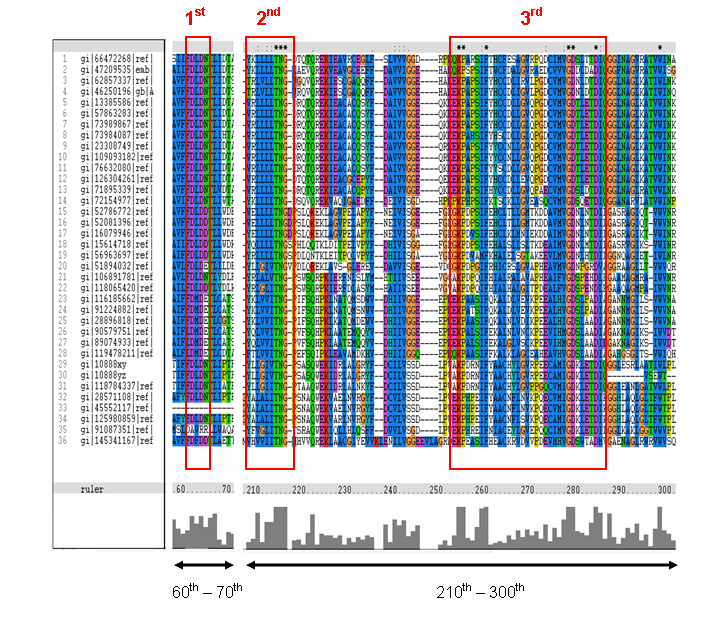
The MSA (Figure 16) for the query sequence and the other 35 sequences shows several conserved motifs. The 1st conserved motif
consists of almost invariant region of aspartic acid (D), only the 33rd protein (gi: |45552117|)
showing gap. The 2nd motif shows conserved and invariant of leucine (L), threonine (T), asparagine (N) and glycine (G). The
3rd motif shows 2 invariant amino acid residues of lysine (K), proline (P), valine (V), glycine (G), aspartic acid (D) and
isoleucine (I). This correlates with the study done by Maliekal et al and strongly suggested that the query protein is a phosphatase.
Figure 16. MSA of the query protein Neu5Ac phosphatase with 35 others proteins. Only the 60th – 70th and the
210th -300th amino acid sequence were shown to illustrate the conserved and invariant regions. The 3 boxed-up sequences
were either conserved or invariant regions.