COASY results
Structure of Coenzyme A Synthase
Conezyme A Synthase is structurally composed of seven strands, eleven helices and thirteen beta turns (EMBL EBI, 2005) (see Figure 2). Analysis of structurally related proteins (Holm & Sander, 1993) showed a trend for transferase (RCSB, 2007) class proteins with Rossmann class folds (Rossmann, 1973). The Rossmann topologies of fold’s are an alpha-beta class fold that forms a three or more layer beta strand sandwich alternating with alpha helices (beta-alpha-beta-alpha-beta). PFAM classification placed these into either the Cytidylyltransferase or Dephospho-CoA kinase familes, matching the two domains of Coenzyme A Synthase. Those classified under the Dephospho-CoA kinase type were commonly of the P-loop containing nucleotide triphosphate hydrolases (Sanger Institute, 2005) homology whilst those classified as Cytidylyltransferase were of the Tyrosol-Transfer RNA Synthetase (Sanger Institute, 2005). As the majority of the structurally related proteins identified contained the DPCK domain solely, and the motif for a P-loop (Table 1) was identified in the Conzyme A Synthase sequence (Bairoch, Bucher, & Hofmann, 1997) it is suggested that Coenzyme A Synthase is also of the P-loop containing nucleotide triphosphate hydrolases homology of folds.
Figure 2
The secondary structure of Mus musculus with indicated ligand interaction sites (EMBL EBI, 2005).
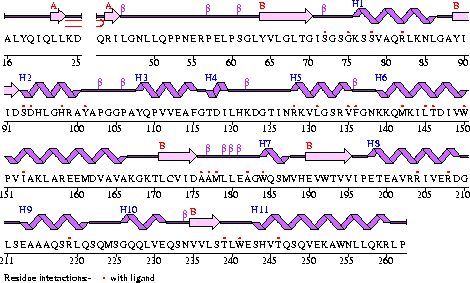
Table 1
| Pattern-ID: | GLYCOSAMINOGLYCAN PS00002 PDOC00002 |
| Pattern-DE: | Glycosaminoglycan attachment site |
| Pattern: | SG.G |
| Position: | 84 SGSG |
| Pattern-ID: | PKC_PHOSPHO_SITE PS00005 PDOC00005 |
| Pattern-DE: | Protein kinase C phosphorylation site |
| Pattern: | [ST].[RK] |
| Position: | 3 SDK |
| Position: | 52 SFR |
| Position: | 86 SGK |
| Pattern-ID: | CK2_PHOSPHO_SITE PS00006 PDOC00006 |
| Pattern-DE: | Casein kinase II phosphorylation site |
| Pattern: | [ST].{2}[DE] |
| Position: | 39 SHNE |
| Position: | 251 TLWE |
| Position: | 260 SQVE |
| Pattern-ID: | MYRISTYL PS00008 PDOC00008 |
| Pattern-DE: | N-myristoylation site |
| Pattern: | G[^EDRKHPFYW].{2}[STAGCN][^P] |
| Position: | 82 GISGSG |
| Position: | 222 GLSEAA |
| Pattern-ID: | ATP_GTP_A PS00017 PDOC00017 |
| Pattern-DE: | ATP/GTP-binding site motif A (P-loop) |
| Pattern: | [AG].{4}GK[ST] |
| Position: | 82 GISGSGKS |
| Pattern-ID: | UPF0038 PS01294 PDOC00996 |
| Pattern-DE: | Uncharacterized protein family UPF0038 signature |
| Pattern: | G.[LI].R.{2}L.{4}F.{8}[LIV].{5}P.[LIV] |
| Position: | 136 GTINRKVLGSRVFGNKKQMKILTDIVWPVI |
Figure tablessss
Structural alignment of structurally related proteins to Mus. musculus Coenzyme A Synthase
Localisation Expression of Coenzyme A Synthase
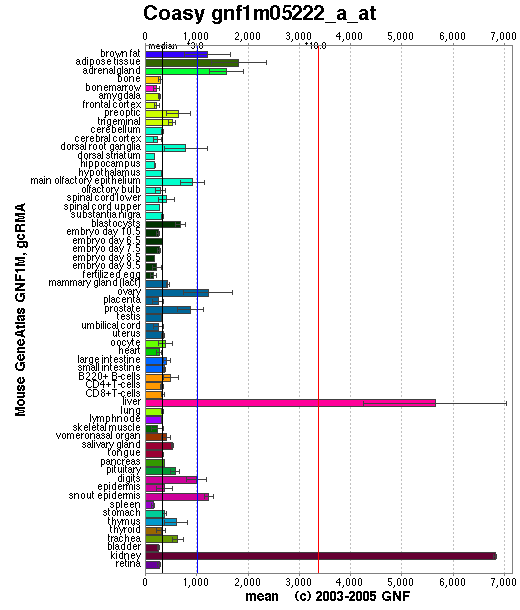
Baseline expression of CoAsy was present in all tissues. Expression was slightly elevated in adipose tissue, and highly elevated in the liver and kidney (See Figure 3). Baseline expression of the other enzymes involved in the CoAsy pathway was also present in all tissues. However, expression of these enzymes was otherwise unlike that of CoA, as they were not particularly elevated in the liver and kidney (data not shown).
Figure 3
CoA Synthase Expression. Reproduced from Genomics Institute of Novartis Research Foundation (2007).
Sequence Conservation of Coenzyme A Synthase
Structural Elements and Functional Binding Sites of Coenzyme A Synthase
Abstract | Introduction | Results | Discussion | Conclusion | Method | References